Abstract
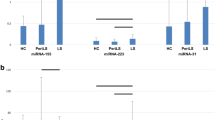
In this study, miRNA profiling of cells infected with lumpy skin disease virus (LSDV) was conducted for the first time. When compared to mock-infected cells, LSDV-infected primary lamb testicle (LT) cells showed dysregulation of 64, 85, and 85 miRNAs at 12 hours postinfection (hpi), 48 hpi, and 72 hpi, respectively. While some of these miRNAs were found to be dysregulated at a particular time point following LSDV infection, others were dysregulated at all three time points. Analysis of the differentially expressed miRNA-mRNA interaction networks, Gene Ontology analysis of the predicted targets, and KEGG analysis of highly enriched pathways revealed several cellular factors/pathways involved in protein/ion/enzyme binding, cell differentiation, movement of subcellular components, calcium reabsorption, aldosterone synthesis and secretion, and melanogenesis. Some selected upregulated (oar-mir-379-5p, oar-let-7d, Chr10-18769, Chr2_5162 and oar-miR-493-5p) and downregulated (ChrX-33741, Chr3_8257 and Chr26_32680) miRNAs were further confirmed by quantitative real-time PCR. These findings contribute to our understanding of virus replication, virus-host interactions, and disease pathogenesis, and the differentially expressed miRNAs and their cellular targets may serve as biomarkers as well as novel targets for therapeutic intervention against LSDV.








Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Babar IA, Slack FJ, Weidhaas JB (2008) miRNA modulation of the cellular stress response. Future Oncol 4:289–298
Bavelloni A, Ramazzotti G, Poli A, Piazzi M, Focaccia E, Blalock W, Faenza I (2017) MiRNA-210: a current overview. Anticancer Res 37:6511–6521
Bernier A, Sagan SM (2018) The diverse roles of microRNAs at the host–virus interface. Viruses 10:440
Brudvik KW, Taskén K (2012) Modulation of T cell immune functions by the prostaglandin E2–cAMP pathway in chronic inflammatory states. Br J Pharmacol 166:411–419
Buggele WA, Horvath CM (2013) MicroRNA profiling of Sendai virus-infected A549 cells identifies miR-203 as an interferon-inducible regulator of IFIT1/ISG56. J Virol 87:9260–9270
Condrat CE, Thompson DC, Barbu MG, Bugnar OL, Boboc A, Cretoiu D, Suciu N, Cretoiu SM, Voinea SC (2020) miRNAs as biomarkers in disease: latest findings regarding their role in diagnosis and prognosis. Cells 9:276
DAHD (2023) Lumpy Skin Disease. Department of Animal Husbandary, Fisheries and Dairying, Govt of India
Diggins NL, Hancock MH (2023) Viral miRNA regulation of host gene expression. Semin Cell Dev Biol 146:2–19
Edelsten M (2014) Threat to European cattle from lumpy skin disease. Vet Rec 175:330
Evripioti AA, Ortega-Prieto AM, Skelton JK, Bazot Q, Dorner M (2019) Phosphodiesterase-induced cAMP degradation restricts hepatitis B virus infection. Philos Trans R Soc B 374:20180292
FAO (2023) Lumpy skin disease symposium. How science can support evidence-based disease management and control. Organized in collaboration with the European Commission for the Control of Foot-and-Mouth Disease (EuFMD) and the Emergency Prevention System for Animal Health (EMPRES-AH) of FAO from 14–16 March 2023. Hybrid: Virtual and in FAO headquarter Rome.
Friedländer MR, Mackowiak SD, Li N, Chen W, Rajewsky N (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 40:37–52
Frohman MA (2015) The phospholipase D superfamily as therapeutic targets. Trends Pharmacol Sci 36:137–144
Girigoswami K, Girigoswami A (2021) A Review on the Role of Nanosensors in Detecting Cellular miRNA Expression in Colorectal Cancer. Endocr Metab Immune Disord Drug Targets 21:12–26
Heidari M, Zhang H, Sunkara L (2022) MDV-induced differential microRNA expression in the primary lymphoid organ of thymus. Microb Pathog 170:105688
Huang W (2017) MicroRNAs: biomarkers, diagnostics, and therapeutics. Bioinformatics in MicroRNA research:57-67
Kumar N, Chander Y, Kumar R, Khandelwal N, Riyesh T, Chaudhary K, Shanmugasundaram K, Kumar S, Kumar A, Gupta MK (2021) Isolation and characterization of lumpy skin disease virus from cattle in India. PLoS ONE 16:e0241022
Kumar N, Tripathi BN (2022) A serious skin virus epidemic sweeping through the Indian subcontinent is a threat to the livelihood of farmers. Virulence 13:1943–1944
Kumar N, Barua S, Kumar R, Khandelwal N, Kumar A, Verma A, Singh L, Godara B, Chander Y, Kumar G, Riyesh T, Sharma DK, Pathak A, Kumar S, Dedar RK, Mehta V, Gaur M, Bhardwaj B, Vyas V, Chaudhary S, Yadav V, Bhati A, Kaul R, Bashir A, Andrabi A, Yousuf RW, Koul A, Kachhawaha S, Gurav A, Gautam S, Tiwari HA, Munjal VK, Gupta MK, Kumar R, Gulati BR, Misri J, Kumar A, Mohanty AK, Nandi S, Singh KP, Pal Y, Dutt T, Tripathi BN (2023) Evaluation of the safety, immunogenicity and efficacy of a new live-attenuated lumpy skin disease vaccine in India. Virulence:2190647
Li J, Li X, Kong X, Luo Q, Zhang J, Fang L (2014) MiRNA-26b inhibits cellular proliferation by targeting CDK8 in breast cancer. Int J Clin Exp Med 7:558–565
Li J, Chen Z, Zhao J, Fang L, Fang R, Xiao J, Chen X, Zhou A, Zhang Y, Ren L (2015) Difference in microRNA expression and editing profile of lung tissues from different pig breeds related to immune responses to HP-PRRSV. Sci Rep 5:9549
Li Z, Xiong Y (2017) Cytoplasmic E3 ubiquitin ligase CUL9 controls cell proliferation, senescence, apoptosis and genome integrity through p53. Oncogene 36:5212–5218
Liao Z, Zheng R, Shao G (2023) Mechanisms and application strategies of miRNA‑146a regulating inflammation and fibrosis at molecular and cellular levels (Review). International journal of molecular medicine 51
Liu B, Li J, Cairns MJ (2014) Identifying miRNAs, targets and functions. Brief Bioinform 15:1–19
Lojkic I, Simic I, Kresic N, Bedekovic T (2018) Complete Genome Sequence of a Lumpy Skin Disease Virus Strain Isolated from the Skin of a Vaccinated Animal. Genome announcements 6
Ma H, Lippolis JD, Casas E (2022) Expression Profiles and Interaction of MicroRNA and Transcripts in Response to Bovine Leukemia Virus Exposure. Frontiers in Veterinary Science 9
Martinez-Espinoza I, Banos-Lara MDR, Guerrero-Plata A (2021) The Importance of miRNA Identification During Respiratory Viral Infections. Journal of cellular immunology 3:207–214
Molla W, de Jong MCM, Gari G, Frankena K (2017) Economic impact of lumpy skin disease and cost effectiveness of vaccination for the control of outbreaks in Ethiopia. Prev Vet Med 147:100–107
OIE (2019) Lympy skin disease. OIE information Database: https://www.oieint/wahis_2/public/wahidphp/Reviewreport/Review?page_refer=MapFullEventReport&reportid=32387
Qi X, Wang T, Xue Q, Li Z, Yang B, Wang J (2018) MicroRNA expression profiling of goat peripheral blood mononuclear cells in response to peste des petits ruminants virus infection. Vet Res 49:1–10
Riolo G, Cantara S, Marzocchi C, Ricci C (2020) miRNA targets: from prediction tools to experimental validation. Methods and protocols 4:1
Roberts AP, Lewis AP, Jopling CL (2011) The role of microRNAs in viral infection. Prog Mol Biol Transl Sci 102:101–139
Sato Y, Hayashi K, Amano Y, Takahashi M, Yonemura S, Hayashi I, Hirose H, Ohno S, Suzuki A (2014) MTCL1 crosslinks and stabilizes non-centrosomal microtubules on the Golgi membrane. Nat Commun 5:1–14
Stenfeldt C, Arzt J, Smoliga G, LaRocco M, Gutkoska J, Lawrence P (2017) Proof-of-concept study: profile of circulating microRNAs in Bovine serum harvested during acute and persistent FMDV infection. Virology journal 14:1–18
Sung T-C, Roper RL, Zhang Y, Rudge SA, Temel R, Hammond SM, Morris AJ, Moss B, Engebrecht J, Frohman MA (1997) Mutagenesis of phospholipase D defines a superfamily including a trans-Golgi viral protein required for poxvirus pathogenicity. EMBO J 16:4519–4530
Tang Q, Wu Y-Q, Chen D-S, Zhou Q, Chen H-C, Liu Z-F (2014) Bovine herpesvirus 5 encodes a unique pattern of microRNAs compared with bovine herpesvirus 1. J Gen Virol 95:671–678
Tasioudi KE, Antoniou SE, Iliadou P, Sachpatzidis A, Plevraki E, Agianniotaki EI, Fouki C, Mangana-Vougiouka O, Chondrokouki E, Dile C (2016) Emergence of Lumpy Skin Disease in Greece, 2015. Transbound Emerg Dis 63:260–265
Tian F, Luo J, Zhang H, Chang S, Song J (2012) MiRNA expression signatures induced by Marek’s disease virus infection in chickens. Genomics 99:152–159
Trobaugh DW, Klimstra WB (2017) MicroRNA Regulation of RNA Virus Replication and Pathogenesis. Trends Mol Med 23:80–93
Wang G, Cheng T, Yuan H, Zou F, Miao P, Jiao J (2023) Tracing cellular interaction of circRNA-miRNA axis with Cu metal-organic framework supported DNA cascade assembly. Biosens Bioelectron 228:115226
Wang M, Gao Z, Pan L, Zhang Y (2014) Cellular microRNAs and picornaviral infections. RNA Biol 11:808–816
Wang X, He Y, Mackowiak B, Gao B (2021) MicroRNAs as regulators, biomarkers and therapeutic targets in liver diseases. Gut 70:784–795
Woods JA (1988) Lumpy skin disease–a review. Trop Anim Health Prod 20:11–17
Xu J, Wong C-W (2013) Enrichment analysis of miRNA targets. MicroRNA Protocols. Springer, pp 91-103
Yang X, Gao W, Liu H, Li J, Chen D, Yuan F, Zhang Z, Wang H (2017) MicroRNA transcriptome analysis in chicken kidneys in response to differing virulent infectious bronchitis virus infections. Adv Virol 162:3397–3405
Zeng G, Wang Z, Huang Y, Abedin Z, Liu Y, Randhawa P (2019) Cellular and viral miRNA expression in polyomavirus BK infection. Transplant infectious disease : an official journal of the Transplantation Society 21:e13159
Zeng Y (2006) Principles of micro-RNA production and maturation. Oncogene 25:6156–6162
Zhu Z, Qi Y, Ge A, Zhu Y, Xu K, Ji H, Shi Z, Cui L, Zhou M (2014) Comprehensive characterization of serum microRNA profile in response to the emerging avian influenza A (H7N9) virus infection in humans. Viruses 6:1525–1539
Funding
This work was supported by the Indian Council of Agricultural Research, New Delhi (grant number IXX16675 to Naveen Kumar).
Author information
Authors and Affiliations
Contributions
Naveen Kumar contributed to the study conception and design. Material preparation was performed by Sakshi Pandita, Assim Verma, Yogesh Chander, and Himanshu Kamboj. Data analysis was performed by Sakshi Pandita, Naveen Kumar, Assim Verma, Yogesh Chander, Himanshu Kamboj, Ram Kumar, Bhupendra Nath Tripathi, and Sanjay Barua. The first draft of the manuscript was written by Naveen Kumar, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study was performed in accordance with the guidelines of the Institutional Biosafety Committee (IBSC) of ICAR-National Research Institute on Equines, Hisar, India. This study did not involve any experiments on animals.
Additional information
Handling Editor: Graciela Andrei.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pandita, S., Verma, A., Kamboj, H. et al. miRNA profiling of primary lamb testicle cells infected with lumpy skin disease virus. Arch Virol 168, 290 (2023). https://doi.org/10.1007/s00705-023-05917-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05917-0