Abstract
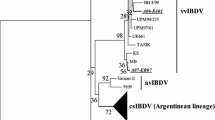
Since 1987, infectious bursal disease virus (IBDV) has circulated and evolved in Vietnam, but little is known about the genotypes present. IBDV samples were collected in 1987, 2001-2006, 2008, 2011, 2015-2019, and 2021 in 18 provinces. We conducted phylogenotyping analysis based on an alignment of 143 VP2-HVR (hypervariable region) sequences from 64 Vietnamese isolates (26 previous and 38 additional sequences and two vaccines, and alignment of 82 VP1 B-marker sequences, including one vaccine and four Vietnamese field strains. The analysis identified three A-genotypes, A1, A3, and A7, and two B-genotypes, B1 and B3, among the Vietnamese IBDV isolates. The lowest average evolutionary distance (8.6%) was seen between the A1 and A3 genotypes, and the highest (21.7%) was between A5 and A7, while there was a distance of 14% between B1 and B3 and 17% between B3 and B2. Unique signature residues were observed for genotypes A2, A3, A5, A6, and A8, which could be used for genotypic discrimination. A timeline statistical summary revealed that the A3-genotype predominated (79.8% presence) in Vietnam from 1987 to 2021 and that it remained the dominant IBDV genotype over the last five years (2016–2021). The current study contributes to a better understanding of the circulating genotypes and evolution of IBDV in Vietnam and worldwide.



Similar content being viewed by others
Availability of data and materials
Genomic and sequence data presented in this paper are publicly available via GenBank and provided in supplemental files.
References
Cosgrove AS (1962) An apparently new disease of chickens—avian nephrosis. Avian Dis 6(3):385–389. https://www.jstor.org/stable/1587909
Lukert PD, Saif YM (2003) Infectious bursal disease. In: Saif YM, Barnes HJ, Glisson JR, Fadly AM, McDougald LR, Swayne DE (eds) Diseases of poultry, 11th edn. Iowa State Press, Ames, pp 161–179
Zhang W, Wang X, Gao Y, Qi X (2022) The over-40-years-epidemic of infectious bursal disease virus in China. Viruses 14(10):2253. https://doi.org/10.3390/v14102253
Pikuła A, Śmietanka K, Perez LJ (2020) Emergence and expansion of novel pathogenic reassortant strains of infectious bursal disease virus causing acute outbreaks of the disease in Europe. Transbound Emerg Dis 67:1739–1174. https://doi.org/10.1111/tbed.13510
Luque D, Saugar I, Rejas MT, Carrascosa JL, Rodríguez JF, Castón JR (2009) Infectious Bursal disease virus: ribonucleoprotein complexes of a double-stranded RNA virus. J Mol Biol 386(3):891–901. https://doi.org/10.1016/j.jmb.2008.11.029
Ye C, Wang Y, Zhang E, Han X, Yu Z, Liu H (2018) VP1 and VP3 are required and sufficient for translation initiation of uncapped infectious bursal disease virus genomic double-stranded RNA. J Virol 92(2):e0134517. https://doi.org/10.1128/JVI.01345-17
Qin Y, Zheng SJ (2017) Infectious bursal disease virus-host interactions: multifunctional viral proteins that perform multiple and differing jobs. Int J Mol Sci 18(1):161. https://doi.org/10.3390/ijms18010161
Van den Berg TP, Morales D, Eterradossi N, Rivallan G, Toquin D, Raue R, Zierenberg K, Zhang MF, Zhu YP, Wang CQ, Zheng HJ, Wang X, Chen GC, Lim BL, Müller H (2004) Assessment of genetic antigenic and pathotypic criteria for the characterization of IBDV strains. Avian Pathol 1:470–476. https://doi.org/10.1080/03079450400003650
Le Nouën C, Toquin D, Müller H, Raue R, Kean KM, Langlois P, Cherbonnel M, Eterradossi N (2012) Different domains of the RNA polymerase of infectious bursal disease virus contribute to virulence. PLoS ONE 7:e28064. https://doi.org/10.1371/journal.pone.0028064
Escaffre O, Le Nouën C, Amelot M, Ambroggio X, Ogden KM, Guionie O, Toquin D, Müller H, Islam MR, Eterradossi N (2013) Both genome segments contribute to the pathogenicity of very virulent infectious bursal disease virus. J Virol 87(5):2767–2780. https://doi.org/10.1128/JVI.02360-12
Jackwood DJ, Sommer-Wagner S (2007) Genetic characteristics of infectious bursal disease viruses from four continents. Virology 365(2):369–375. https://doi.org/10.1016/j.virol.2007.03.046
Islam MR, Nooruzzaman M, Rahman T, Mumu TT, Rahman MM, Chowdhury EH, Eterradossi N, Müller H (2021) A unified genotypic classification of infectious bursal disease virus based on both genome segments. Avian Pathol 50(2):190–206. https://doi.org/10.1080/03079457.2021.1873245
Graziosi G, Catelli E, Fanelli A, Lupini C (2022) Infectious bursal disease virus in free-living wild birds: a systematic review and meta-analysis of its sero-viroprevalence on a global scale. Transbound Emerg Dis 69(5):2800–2815. https://doi.org/10.1111/tbed.14433
Tomás G, Marandino A, Courtillon C, Amelot M, Keita A, Pikula A, Hernández M, Hernández D, Vagnozzi A, Panzera Y, Domańska-Blicharz K, Eterradossi N, Pérez R, Soubies SM (2019) Antigenicity, pathogenicity and immunosuppressive effect caused by a South American isolate of infectious bursal disease virus belonging to the “distinct” genetic lineage. Avian Pathol 48(3):245–254. https://doi.org/10.1080/03079457.2019.1572867
Lupini C, Giovanardi D, Pesente P, Bonci M, Felice V, Rossi G, Morandini E, Cecchinato M, Catelli E (2016) A molecular epidemiology study based on VP2 gene sequences reveals that a new genotype of infectious bursal disease virus is dominantly prevalent in Italy. Avian Pathol 45:458–464. https://doi.org/10.1080/03079457.2016.1165792
Michel LO, Jackwood DJ (2017) Classification of infectious bursal disease virus into genogroups. Arch Virol 162:3661–3670. https://doi.org/10.1007/s00705-017-3500-4
Wang Y, Fan L, Jiang N, Gao L, Li K, Gao Y, Liu C, Cui H, Pan Q, Zhang Y, Wang X, Qi X (2021) An improved scheme for infectious bursal disease virus genotype classification based on both genome-segments A and B. J Integr Agric 20:1372–1381. https://doi.org/10.1016/S2095-3119(20)63424-4
Pikuła A, Lisowska A (2022) Genetics and pathogenicity of natural reassortant of infectious bursal disease virus emerging in latvia. Pathogens 11(10):1081. https://doi.org/10.3390/pathogens11101081
Jackwood DJ, Schat KA, Michel LO, de Wit S (2018) A proposed nomenclature for infectious bursal disease virus isolates. Avian Pathol 47(6):576–584. https://doi.org/10.1080/03079457.2018.1506092
Nooruzzaman M, Hossain I, Rahman MM, Uddin AJ, Mustari A, Parvin R, Chowdhury EH, Islam MR (2022) Comparative pathogenicity of infectious bursal disease viruses of three different genotypes. Microb Pathog 169:105641. https://doi.org/10.1016/j.micpath.2022.105641
Wang Y, Jiang N, Fan L, Niu X, Zhang W, Huang M, Gao L, Li K, Gao Y, Liu C, Cui H, Liu A, Pan Q, Zhang Y, Wang X, Qi X (2021) Identification and pathogenicity evaluation of a novel reassortant infectious bursal disease virus (Genotype A2dB3). Viruses 13(9):1682. https://doi.org/10.3390/v13091682
Jiang N, Wang Y, Zhang W, Niu X, Gao Y, Gao L, Li K, Cui H, Liu A, Pan Q, Liu C, Zhang Y, Wang X, Qi X (2021) Naturally occurring mutated infectious bursal disease virus of genotype A8B1 associated with bursa damage in China. Virus Res 302:198498. https://doi.org/10.1016/j.virusres.2021.198498
Jiang N, Wang Y, Zhang W, Niu X, Huang M, Gao Y, Liu A, Gao L, Li K, Pan Q, Liu C, Zhang Y, Cui H, Wang X, Qi X (2021) Genotyping and molecular characterization of infectious bursal disease virus identified in important poultry-raising areas of China during 2019 and 2020. Front Vet Sci 8:759861. https://doi.org/10.3389/fvets.2021.759861
Cao YC, Yeung WS, Law M, Bi YZ, Leung FC, Lim BL (1998) Molecular characterization of seven Chinese isolates of infectious bursal disease virus: classical, very virulent, and variant strains. Avian Dis 42(2):340–351. https://www.jstor.org/stable/1592484
Sapats SI, Ignjatovic J (2000) Antigenic and sequence heterogeneity of infectious bursal disease virus strains isolated in Australia. Arch Virol 145:773–785. https://doi.org/10.1007/s007050050670
Silva FM, Vidigal PM, Myrrha LW, Fietto JL, Silva A Jr, Almeida MR (2013) Tracking the molecular epidemiology of Brazilian Infectious bursal disease virus (IBDV) isolates. Infect Genet Evol 13:18–26. https://doi.org/10.1016/j.meegid.2012.09.005
Le XTK, Doan HTT, Do RT, Le TH (2019) Molecular characterization of field isolates of infectious bursal disease virus from three decades, 1987–2018, reveals a distinct genotypic subgroup in Vietnam. Arch Virol 164(8):2137–2145. https://doi.org/10.1007/s00705-019-04287-w
Hou B, Wang CY, Luo ZB, Shao GQ (2022) Commercial vaccines used in China do not protect against a novel infectious bursal disease virus variant isolated in Fujian. Vet Rec 191(10):e1840. https://doi.org/10.1002/vetr.1840
To H, Yamaguchi T, Nguyen NT, Nguyen OT, Nguyen SV, Agus S, Kim HJ, Fukushi H, Hirai K (1999) Sequence comparison of the VP2 variable region of infectious bursal disease virus isolates from Vietnam. J Vet Med Sci 61(4):429–432. https://doi.org/10.1292/jvms.61.429
Aliyu HB, Hair-Bejo M, Omar AR, Ideris A (2021) Genetic diversity of recent infectious bursal disease viruses isolated from vaccinated poultry flocks in Malaysia. Front Vet Sci 8:643976. https://doi.org/10.3389/fvets.2021.643976
Franciosini MP, Davidson IA (2022) Walk through Gumboro disease. Poultry 1:229–243. https://doi.org/10.3390/poultry1040020
Wu T, Wang Y, Li H, Fan L, Jiang N, Gao L, Li K, Gao Y, Liu C, Cui H, Pan Q, Zhang Y, Wang X, Qi X (2020) Naturally occurring homologous recombination between novel variant infectious bursal disease virus and intermediate vaccine strain. Vet Microbiol 245:108700. https://doi.org/10.1016/j.vetmic.2020.108700
OIE (2018) Infectious bursal disease (Gumboro disease). In: Manual of Diagnostic Tests and Vaccines for Terrestrial Animals 2018. World Organisation for Animal Health. Terrestrial Manual Online Access: https://www.woah.org/en/what-we-do/standards/codes-and-manuals/terrestrial-manual-online-access/3.03.12_IBD.pdf. Accessed 22 Dec 2022
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Junier T, Zdobnov EM (2010) The Newick utilities: high-throughput phylogenetic tree processing in the UNIX shell. Bioinformatics 26(13):1669–1670. https://doi.org/10.1093/bioinformatics/btq243
Rambaut A (2018) FigTree, version 1.4.4. http://tree.bio.ed.ac.uk/software/figtree/.
Brown MD, Green P, Skinner MA (1992) VP2 sequences of recent European “very virulent” isolates of infectious bursal disease virus are closely related to each other but are distinct from those of “classical” strains. J Gen Virol 75(3):675–680. https://doi.org/10.1099/0022-1317-75-3-675
Lazarus D, Pasmanik-Chor M, Gutter B, Gallili G, Barbakov M, Krispel S, Pitcovski J (2008) Attenuation of very virulent infectious bursal disease virus and comparison of full sequences of virulent and attenuated strains. Avian Pathol 37(2):151–159. https://doi.org/10.1080/03079450801910206
Lian J, Wang Z, Xu Z, Pang Y, Leng M, Tang S, Zhang X, Qin J, Chen F, Lin W (2022) Pathogenicity and molecular characterization of infectious bursal disease virus in China. Poult Sci 101(1):101502. https://doi.org/10.1016/j.psj.2021.101502
He X, Xiong Z, Yang L, Guan D, Yang X, Wei P (2014) Molecular epidemiology studies on partial sequences of both genome segments reveal that reassortant infectious bursal disease viruses were dominantly prevalent in southern China during 2000–2012. Arch Virol 159(12):3279–3292. https://doi.org/10.1007/s00705-014-2195-z
Chen Z, Lian J, Liang Z, Leng M, Lin W, Chen F (2022) Characterization and pathogenicity of infectious bursal disease virus in Southern China. Poult Sci 101(10):102018. https://doi.org/10.1016/j.psj.2022.102018
Thai TN, Jang I, Kim HA, Kim HS, Kwon YK, Kim HR (2021) Characterization of antigenic variant infectious bursal disease virus strains identified in South Korea. Avian Pathol 50(2):174–181. https://doi.org/10.1080/03079457.2020.1869698
Patel AK, Pandey VC, Pal JK (2016) Evidence of genetic drift and reassortment in infectious bursal disease virus and emergence of outbreaks in poultry farms in India. Virusdisease 27:161–169. https://doi.org/10.1007/s13337-016-0306-z
Ali Khan RS, Habib M, Ali W, Salah Ud Din Shah M, Ashraf A, Ali Tahir Z, Helal ZH, Khan MI, Mahboob S, A-Al-Ghanim K, Al-Misned F (2019) Phylogenetic analysis of infectious bursal disease viruses according to newly proposed model of classification into geno-groups. J Infect Public Health 12(3):410–418. https://doi.org/10.1016/j.jiph.2018.12.012
Hussain A, Wu T, Li H, Fan L, Li K, Gao L, Wang Y, Gao Y, Liu C, Cui H, Pan Q, Zhang Y, Aslam A, Muti-Ur-Rehman K, Munir M, Butt SL, Wang X, Qi X (2019) Pathogenic characterization and full length genome sequence of a reassortant infectious bursal disease virus newly isolated in Pakistan. Virol Sin 34(1):102–105. https://doi.org/10.1007/s12250-019-00082-8
Mató T, Medveczki A, Kiss I (2022) Research Note: “Hidden” infectious bursal disease virus infections in Central Europe. Poult Sci 101(8):101958. https://doi.org/10.1016/j.psj.2022.101958
Abed M, Soubies S, Courtillon C, Briand FX, Allée C, Amelot M, De Boisseson C, Lucas P, Blanchard Y, Belahouel A, Kara R, Essalhi A, Temim S, Khelef D, Eterradossi N (2018) Infectious bursal disease virus in Algeria: detection of highly pathogenic reassortant viruses. Infect Genet Evol 60:48–57. https://doi.org/10.1016/j.meegid.2018.01.029
Drissi Touzani C, Fellahi S, Fassi Fihri O, Gaboun F, Khayi S, Mentag R, Lico C, Baschieri S, El Houadfi M, Ducatez M (2020) Complete genome analysis and time scale evolution of very virulent infectious bursal disease viruses isolated from recent outbreaks in Morocco. Infect Genet Evol 77:104097. https://doi.org/10.1016/j.meegid.2019.104097
Mosad SM, Eladl AH, El-Tholoth M, Ali HS, Hamed MF (2020) Molecular characterization and pathogenicity of very virulent infectious bursal disease virus isolated from naturally infected turkey poults in Egypt. Trop Anim Health Prod 52(6):3819–3831. https://doi.org/10.1007/s11250-020-02420-5
Arowolo OA, George UE, Luka PD, Maurice NA, Atuman YJ, Shallmizhili JJ, Shittu I, Oluwayelu DO (2021) Infectious bursal disease in Nigeria: continuous circulation of reassortant viruses. Trop Anim Health Prod 53(2):271. https://doi.org/10.1007/s11250-021-02719-x
Stoute ST, Jackwood DJ, Sommer-Wagner SE, Crossley BM, Woolcock PR, Charlton BR (2013) Pathogenicity associated with coinfection with very virulent infectious bursal disease and Infectious bursal disease virus strains endemic in the United States. J Vet Diagn Invest 25(3):352–358. https://doi.org/10.1177/1040638713483538
Hernández M, Tomás G, Marandino A, Iraola G, Maya L, Mattion N, Hernández D, Villegas P, Banda A, Panzera Y, Pérez R (2015) Genetic characterization of South American infectious bursal disease virus reveals the existence of a distinct worldwide-spread genetic lineage. Avian Pathol 44(3):212–221. https://doi.org/10.1080/03079457.2015.1025696
Stuart JC (1989) Acute infectious bursal disease in poultry. Vet Rec 125:281
Chettle NJ, Stuart JC, Wyeth PJ (1989) Outbreaks of virulent infectious bursal disease in East Anglia. Vet Rec 125:271–272. https://doi.org/10.1136/vr.125.10.271
Ignjatovic J, Sapats S (2002) Confirmation of the existence of two distinct genetic groups of infectious bursal disease virus in Australia. Aust Vet J 80(11):689–694. https://doi.org/10.1111/j.1751-0813.2002.tb11299.x
Mwenda R, Changula K, Hang’ombe BM, Chidumayo N, Mangani AS, Kaira T, Takada A, Mweene AS, Simulundu E (2018) Characterization of field infectious bursal disease viruses in Zambia: evidence of co-circulation of multiple genotypes with predominance of very virulent strains. Avian Pathol 47(3):300–313. https://doi.org/10.1080/03079457.2018.1449941
Feng X, Zhu N, Cui Y, Hou L, Zhou J, Qiu Y, Yang X, Liu C, Wang D, Guo J, Sun T, Shi Y, Han N, Mo M, Liu J (2022) Characterization and pathogenicity of a naturally reassortant and recombinant infectious bursal disease virus in China. Transbound Emerg Dis 69(4):e746–e758. https://doi.org/10.1111/tbed.14347
Diel DG, da Silva LH, Liu H, Wang Z, Miller PJ, Afonso CL (2012) Genetic diversity of avian paramyxovirus type 1: proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infect Genet Evol 12(8):1770–1779. https://doi.org/10.1016/j.meegid.2012.07.012
Dimitrov KM, Abolnik C, Afonso CL, Albina E, Bahl J, Berg M et al (2019) Updated unified phylogenetic classification system and revised nomenclature for Newcastle disease virus. Infect Genet Evol 74:103917. https://doi.org/10.1016/j.meegid.2019.103917
Valastro V, Holmes EC, Britton P, Fusaro A, Jackwood MW, Cattoli G, Monne I (2016) S1 gene-based phylogeny of infectious bronchitis virus: an attempt to harmonize virus classification. Infect Genet Evol 39:349–364. https://doi.org/10.1016/j.meegid.2016.02.015
World Health Organization/World Organisation for Animal Health/Food and Agriculture Organization (WHO/OIE/FAO) (2014) H5N1 Evolution Working Group. Revised and updated nomenclature for highly pathogenic avian influenza A (H5N1) viruses. Influenza Other Respir Viruses 8(3):384–388. https://doi.org/10.1111/irv.12230.
Fan L, Wu T, Wang Y, Hussain A, Jiang N, Gao L, Li K, Gao Y, Liu C, Cui H, Pan Q, Zhang Y, Wang X, Qi X (2020) Novel variants of infectious bursal disease virus can severely damage the bursa of fabricius of immunized chickens. Vet Microbiol 240:108507. https://doi.org/10.1016/j.vetmic.2019.108507
Bao K, Qi X, Li Y, Gong M, Wang X, Zhu P (2022) Cryo-EM structures of infectious bursal disease viruses with different virulences provide insights into their assembly and invasion. Sci Bull (Beijing) 67(6):646–654. https://doi.org/10.1016/j.scib.2021.12.009
Acknowledgments
This research was funded by the Vietnam National Foundation for Science and Technology Development (NAFOSTED) under grant number 106.02-2020.03 (PI: Xuyen Thi Kim Le). We express our thanks to colleagues for their kind provision of the field materials used in this study.
Funding
This research was funded by the Vietnam National Foundation for Science and Technology Development (NAFOSTED) under grant number 106.02-2020.03.
Author information
Authors and Affiliations
Contributions
XTKL supervised the laboratory work and analyzed the datasets for the manuscript draft. RTD, HTTD, KTN, and LTKP collected, processed, and performed experiments and sequence analysis. THL finalized the datasets, tables, and figures and wrote, revised, completed, and approved the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: William G Dundon .
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Le, X.T.K., Do, R.T., Doan, H.T.T. et al. Phylogenotyping of infectious bursal disease virus in Vietnam according to the newly unified genotypic classification scheme. Arch Virol 168, 201 (2023). https://doi.org/10.1007/s00705-023-05830-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05830-6