Abstract
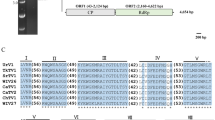
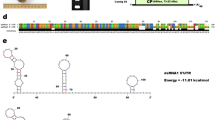
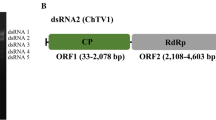
Mycoviruses are widely distributed across the kingdom Fungi, including ascomycetous yeast strains of the class Saccharomycetes. Geotrichum candidum is an important fungal pathogen belonging to Saccharomycetes and has a diverse host range. Here, we report the characterization of four new classical totiviruses from two distinct Geotrichum candidum strains from Pakistan. The four identified viruses were tentatively named “Geotrichum candidum totivirus 1, 2, 3a, and 3b” (GcTV1-3b). The complete dsRNA genomes of the identified totiviruses are 4621, 4592, 4576, and 4576 bp in length, respectively. All totivirus genomes have two open reading frames, encoding a capsid protein (CP) and an RNA-dependent RNA polymerase (RdRP), respectively. The downstream RdRP domain is assumed to be expressed as a CP-RdRP fusion product via -1 frameshifting mediated by a heptameric slippery site. Sequence comparisons and phylogenetic analysis showed that each of the discovered viruses belongs to a new species of the genus Totivirus in the family Totiviridae, with GcTV1 and GcTV3 (a and b strains) clustering in one subgroup and GcTV2 in another subgroup.


Similar content being viewed by others
References
Atkins JF, Loughran G, Bhatt PR, Firth AE, Baranov PV (2016) Ribosomal frameshifting and transcriptional slippage: from genetic steganography and cryptography to adventitious use. Nucleic Acids Res 44:7007–7078
Chen S, Cao L, Huang Q, Qian Y, Zhou X (2016) The complete genome sequence of a novel maize-associated totivirus. Arch Virol 161:487–490
Dinman JD, Icho T, Wickner RB (1991) A -1 ribosomal frameshift in a double-stranded RNA virus of yeast forms a gag-pol fusion protein. Proc Natl Acad Sci USA 88:174–178
Eusebio-Cope A, Suzuki N (2015) Mycoreovirus genome rearrangements associated with RNA silencing deficiency. Nucleic Acids Res 43:3802–3813
Fujimura T, Esteban R (2011) Cap-snatching mechanism in yeast L-A double-stranded RNA virus. Proc Natl Acad Sci USA 108:17667–17671
Garcia-Pedrajas MD, Canizares MC, Sarmiento-Villamil JL, Jacquat AG, Dambolena JS (2019) Mycoviruses in biological control: From basic research to field implementation. Phytopathology 109:1828–1839
Ghabrial SA, Caston JR, Jiang D, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 479–480:356–368
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Hillman BI, Cohen AB (2021) Totiviruses (Totiviridae). In: Bamford D, Zuckerman M (eds) Encyclopedia of virology, 4th edn. Elsevier, Oxford. https://doi.org/10.1016/B1978-012374410-012374414.012300518-012374415
Isawa H, Kuwata R, Hoshino K, Tsuda Y, Sakai K, Watanabe S, Nishimura M, Satho T, Kataoka M, Nagata N, Hasegawa H, Bando H, Yano K, Sasaki T, Kobayashi M, Mizutani T, Sawabe K (2011) Identification and molecular characterization of a new nonsegmented double-stranded RNA virus isolated from Culex mosquitoes in Japan. Virus Res 155:147–155
Khan HA, Sato Y, Kondo H, Jamal A, Bhatti MF, Suzuki N (2021) A second capsidless hadakavirus strain with 10 positive-sense single-stranded RNA genomic segments from Fusarium nygamai. Arch Virol 166:2711–2722
Khan HA, Shamsi W, Jamal A, Javaied M, Sadiq M, Fatma T, Ahmed A, Arshad M, Waseem M, Babar S, Dogar MM, Virk N, Janjua HA, Kondo H, Suzuki N, Bhatti MF (2021) Assessment of mycoviral diversity in Pakistani fungal isolates revealed infection by 11 novel viruses of a single strain of Fusarium mangiferae isolate SP1. J Gen Virol. https://doi.org/10.1099/jgv.1090.001690
Khan HA, Sato Y, Kondo H, Jamal A, Bhatti MF, Suzuki N (2022) A novel victorivirus from the phytopathogenic fungus Neofusicoccoum parvum. Arch Virol 167:923–929. https://doi.org/10.1007/s00705-00021-05304-00707
Khan HA, Telengech P, Kondo H, Bhatti MF, Suzuki N (2022) Mycovirus hunting revealed the presence of diverse viruses in a single isolate of the phytopathogenic fungus Diplodia seriata from Pakistan. Front Cell Infect Microbiol 12:913619. https://doi.org/10.3389/fcimb.2022.913619
Kondo H, Hisano S, Chiba S, Maruyama K, Andika IB, Toyoda K, Fujimori F, Suzuki N (2016) Sequence and phylogenetic analyses of novel totivirus-like double-stranded RNAs from field-collected powdery mildew fungi. Virus Res 213:353–364
Kondo H, Bottela L, Suzuki N (2022) Mycovirus diversity and evolution revealed/inferred from recent studies. Annu Rev Phytopathol. https://doi.org/10.1146/annurev-phyto-021621-122122
Mor H, Steinlauf R, Barash I (1984) Virus-like particles and double-stranded RNA in Geotrichum candidum, the causal agent of citrus sour rot. Phytopathology 74:921–924
Pottier I, Gente S, Vernoux JP, Gueguen M (2008) Safety assessment of dairy microorganisms: Geotrichum candidum. Int J Food Microbiol 126:327–332
Poulos BT, Tang KF, Pantoja CR, Bonami JR, Lightner DV (2006) Purification and characterization of infectious myonecrosis virus of penaeid shrimp. J Gen Virol 87:987–996
Shiba K, Hatta C, Sasai S, Tojo M, Ohki ST, Mochizuki T (2019) A novel toti-like virus from a plant pathogenic oomycete Globisporangium splendens. Virology 537:165–171
Suzuki N, Supyani S, Maruyama K, Hillman BI (2004) Complete genome sequence of Mycoreovirus-1/Cp9B21, a member of a novel genus within the family Reoviridae, isolated from the chestnut blight fungus Cryphonectria parasitica. J Gen Viro 85:3437–3448
Wade NL, Morris SC (1982) Causes and control of cantaloupe postharvest wastage in Australia. Plant Dis 66:549–552
Wells JM (1977) Sour rot of peaches caused by Monilinia implicata and Geotrichum candidum. Phytopathology 67:404–408
Wickner RB (1996) Double-stranded RNA viruses of Saccharomyces cerevisiae. Microbiol Rev 60:250–265
Wickner RB, Ghabrial SA, Nibert ML, Patterson JL, Wang CC (2011) Family Totiviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowits EJ (eds) Virus taxonomy: ninth report of the international committee for the taxonomy of viruses. Elsevier, New York, pp 639–650
Wuest PJ, Baker KF, Conway WS (1970) Sensitivity of selected mushroom pathogens to aerated steam. Phytopathology 60:1274–1275
Acknowledgements
The authors are grateful to Ms. Sakae Hisano for technical assistance. HAK is thankful to the Higher Education Commission (HEC) of Pakistan for a fellowship under the International Research Support Initiative Program (IRSIP).
Funding
This investigation was partly supported by a HEC grant (no. PM-IPFP/ HRD/HEC/2012/2718 and NRPU-4109 awarded to MFB) and NUST students research funds, Grants-in-Aid for Scientific Research (S) and (A) and Grants-in-Aid for Scientific Research on Innovative Areas from the Japanese Ministry of Education, Culture, Sports, Science and Technology (KAKENHI 21H05035, 17H01463, 16H06436, 16H06429 and 16K21723 to NS and HK).
Author information
Authors and Affiliations
Contributions
Conceptualization: NS, MFB. Investigation: HAK, SS and HK. Writing—original draft: NS, HAK and MFB. Writing—reviewing and editing: NS, HAK, MFB, and HK. Funding acquisition: NS, HK, and MFB.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Ioly Kotta-Loizou.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, H.A., Kondo, H., Shahi, S. et al. Identification of novel totiviruses from the ascomycetous fungus Geotrichum candidum. Arch Virol 167, 2833–2838 (2022). https://doi.org/10.1007/s00705-022-05611-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05611-7