Abstract
Mammalian orthoreoviruses (MRVs) are non-enveloped double-stranded RNA viruses with a broad host range. MRVs are prevalent worldwide, and in Japan, they have been isolated from various hosts, including humans, dogs, cats, wild boars, and pigs, and they have also been found in sewage. However, Japanese porcine MRVs have not been genetically characterized. While investigating porcine enteric viruses including MRV, five MRVs were isolated from the feces of Japanese pigs using MA104 cell culture. Genetic analysis of the S1 gene revealed that the Japanese porcine MRV isolates could be classified as MRV-2 and MRV-3. Whole genome analysis showed that Japanese porcine MRVs exhibited genetic diversity, although they shared sequence similarity with porcine MRV sequences in the DDBJ/EMBL/GenBank database. Several potential intragenetic reassortment events were detected among MRV strains from pigs, sewage, and humans in Japan, suggesting zoonotic transmission. Furthermore, homologous recombination events were identified in the M1 and S1 genes of Japanese porcine MRV. These findings imply that different strains of Japanese porcine MRV share a porcine MRV genomic backbone and have evolved through intragenetic reassortment and homologous recombination events.


Similar content being viewed by others
References
Day JM (2009) The diversity of the orthoreoviruses: molecular taxonomy and phylogenetic divides. Infect Genet Evol 9:390–400. https://doi.org/10.1016/j.meegid.2009.01.011
Guglielmi KM, Johnson EM, Stehle T, Dermody TS (2006) Attachment and cell entry of mammalian orthoreovirus. Curr Top Microbiol Immunol 309:1–38. https://doi.org/10.1007/3-540-30773-7_1
Tai JH, Williams JV, Edwards KM, Wright PF, Crowe JE Jr, Dermody TS (2005) Prevalence of reovirus-specific antibodies in young children in Nashville, Tennessee. J Infect Dis 191:1221–1224. https://doi.org/10.1086/428911
Yamamoto SP, Motooka D, Egawa K, Kaida A, Hirai Y, Kubo H, Motomura K, Nakamura S, Iritani N (2020) Novel human reovirus isolated from children and its long-term circulation with reassortments. Sci Rep 10(1):963. https://doi.org/10.1038/s41598-020-58003-9
Johansson PJ, Sveger T, Ahlfors K, Ekstrand J, Svensson L (1996) Reovirus type 1 associated with meningitis. Scand J Infect Dis 28:117–120. https://doi.org/10.3109/00365549609049060
Hermann L, Embree J, Hazelton P, Wells B, Coombs RT (2004) Reovirus type 2 isolated from cerebrospinal fluid. Pediatr Infect Dis J 23:373–375. https://doi.org/10.1097/00006454-200404000-00026
Tyler KL, Barton ES, Ibach ML, Robinson C, Campbell JA, O’Donnell SM, Valyi-Nagy T, Clarke P, Wetzel JD, Dermody T (2004) Isolation and molecular characterization of a novel type 3 reovirus from a child with meningitis. J Infect Dis 189:1664–1675. https://doi.org/10.1086/383129
Chua KB, Crameri G, Hyatt A, Yu M, Tompang MR, Rosli J, McEachern J, Crameri S, Kumarasamy V, Eaton BT, Wang LF (2007) A previously unknown reovirus of bat origin is associated with an acute respiratory disease in humans. Proc Natl Acad Sci USA 104:11424–11429. https://doi.org/10.1073/pnas.0701372104
Cheng P, Lau CS, Lai A, Ho E, Leung P, Chan F, Wong A, Lim W (2009) A novel reovirus isolated from a patient with acute respiratory disease. J Clin Virol 45:79–80. https://doi.org/10.1016/j.jcv.2009.03.001
Chua KB, Voon K, Yu M, Keniscope C, Abdul Rasid K, Wang LF (2011) Investigation of a potential zoonotic transmission of orthoreovirus associated with acute influenza-like illness in an adult patient. PLoS ONE 6(10):e25434. https://doi.org/10.1371/journal.pone.0025434
Kwon HJ, Kim HH, Kim HJ, Park JG, Son KY, Jung J, Lee WS, Cho KO, Park SJ, Kang MI (2012) Detection and molecular characterization of porcine type 3 orthoreoviruses circulating in South Korea. Vet Microbiol 157:456–463. https://doi.org/10.1016/j.vetmic.2011.12.032
Thimmasandra Narayanappa A, Sooryanarain H, Deventhiran J, Cao D, Ammayappan Venkatachalam B, Kambiranda D, LeRoith T, Heffron CL, Lindstrom N, Hall K, Jobst P, Sexton C, Meng XJ, Elankumaran S (2015) A novel pathogenic mammalian orthoreovirus from diarrheic pigs and swine blood meal in the United States. MBio. https://doi.org/10.1128/mBio.00593-15,e00593-e00515
Lelli D, Beato MS, Cavicchio L, Lavazza A, Chiapponi C, Leopardi S, Baioni L, De Benedictis P, Moreno A (2016) First identification of mammalian orthoreovirus type 3 in diarrheic pigs in Europe. Virol J 13:139. https://doi.org/10.1186/s12985-016-0593-4
Qin P, Li H, Wang JW, Wang B, Xie RH, Xu H, Zhao LY, Li L, Pan Y, Song Y, Huang YW (2017) Genetic and pathogenic characterization of a novel reassortant mammalian Orthoreovirus 3 (MRV3) from a diarrheic piglet and seroepidemiological survey of MRV3 in diarrheic pigs from east China. Vet Microbiol 208:126–136. https://doi.org/10.1016/j.vetmic.2017.07.021
Luo Y, Fei L, Yue H, Li S, Ma H, Tang C (2020) Prevalence and genomic characteristics of a novel reassortment mammalian orthoreovirus type 2 in diarrhea piglets in Sichuan, China. Infect Genet Evol 85:104420. https://doi.org/10.1016/j.meegid.2020.104420,104420
Wang L, Li Y, Walsh T, Shen Z, Li Y, Deb Nath N, Lee J, Zheng B, Tao Y, Paden CR, Queen K, Zhang S, Tong S, Ma W (2021) Isolation and characterization of novel reassortant mammalian orthoreovirus from pigs in the United States. Emerg Microbes Infect 10:1137–1147. https://doi.org/10.1080/22221751.2021.1933608
Dermody TS, Parker JSL, Sherry B (2013) Orthoreoviruses. In: David MK, Peter MH (eds) Fields virology. Lippincott Williams & Wilkins, Philadelphia, pp 1304–1346
Hirahara T, Yasuhara H, Matsui O, Kodama K, Nakai M, Sasaki N (1988) Characteristics of reovirus type 1 from the respiratory tract of pigs in Japan. Nihon Juigaku Zasshi. 50:353–361. https://doi.org/10.1292/jvms1939.50.353
Fukutomi T, Sanekata T, Akashi H (1996) Isolation of reovirus type 2 from diarrheal feces of pigs. J Vet Med Sci 58:555–557. https://doi.org/10.1292/jvms.58.555
Zhang W, Kataoka M, Doan YH, Oi T, Furuya T, Oba M, Mizutani T, Oka T, Li TC, Nagai M (2021) Isolation and characterization of mammalian orthoreovirus type 3 from a fecal sample from a wild boar in Japan. Arch Virol 166:1671–1680. https://doi.org/10.1007/s00705-021-05053-7
Leary TP, Erker JC, Chalmers ML, Cruz AT, Wetzel JD, Desai SM, Mushahwar IK, Dermody TS (2002) Detection of mammalian reovirus RNA by using reverse transcription-PCR: sequence diversity within the lambda3-encoding L1 gene. J Clin Microbiol 40:1368–1375. https://doi.org/10.1128/JCM.40.4.1368-1375.2002
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol 1(1):vev003. https://doi.org/10.1093/ve/vev003
Lole KS, Bollinger RC, Paranjape RS, Gadkari D, Kulkarni SS, Novak NG, Ingersoll R, Sheppard HW, Ray SC (1999) Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J Virol 73:152–160. https://doi.org/10.1128/JVI.73.1.152-160.1999
Farkas SL, Marton S, Dandar E, Kugler R, Gal B, Jakab F, Balint A, Kecskemeti S, Banyai K (2016) Lineage diversification, homo- and heterologous reassortment and recombination shape the evolution of chicken orthoreoviruses. Sci Rep 10(6):36960. https://doi.org/10.1038/srep36960
Noh JY, Lee DH, Lim TH, Lee JH, Day JM, Song CS (2018) Isolation and genomic characterization of a novel avian orthoreovirus strain in Korea, 2014. Arch Virol 163:1307–1316. https://doi.org/10.1007/s00705-017-3667-8
Ye D, Ji Z, Shi H, Chen J, Shi D, Cao L, Liu J, Li M, Dong H, Jing Z, Wang X, Liu Q, Fan Q, Cong G, Zhang J, Han Y, Zhou J, Gu J, Zhang X, Feng L (2020) Molecular characterization of an emerging reassortant mammalian orthoreovirus in China. Arch Virol 165:2367–2372. https://doi.org/10.1007/s00705-020-04712-5
Kitamura K, Takagi H, Oka T, Kataoka M, Ueki Y, Sakagami A (2021) Intertypic reassortment of mammalian orthoreovirus identified in wastewater in Japan. Sci Rep 11(1):12583. https://doi.org/10.1038/s41598-021-92019-z
Yang XL, Tan B, Wang B, Li W, Wang N, Luo CM, Wang MN, Zhang W, Li B, Peng C, Ge XY, Zhang LB, Shi ZL (2015) Isolation and identification of bat viruses closely related to human, porcine and mink orthoreoviruses. J Gen Virol 96:3525–3531. https://doi.org/10.1099/jgv.0.000314
Cavicchio L, Tassoni L, Zamperin G, Campalto M, Carrino M, Leopardi S, Benedictis P, Beato MS (2020) Unexpected genetic diversity of two novel swine MRVs in Italy. Viruses 12(5):574. https://doi.org/10.3390/v12050574
Harima H, Sasaki M, Kajihara M, Gonzalez G, Simulundu E, Bwalya EC, Qiu Y, Okuya K, Isono M, Orba Y, Takada A, Hang’ombe BM, Mweene AS, Sawa H (2020) Characterization of mammalian orthoreoviruses isolated from feces of pigs in Zambia. J Gen Virol 101:1027–1036. https://doi.org/10.1099/jgv.0.001476
Zhang C, Liu L, Wang P, Liu S, Lin W, Hu F, Wu W, Chen W, Cui S (2011) A potentially novel reovirus isolated from swine in northeastern China in 2007. Virus Genes 43:342–349. https://doi.org/10.1007/s11262-011-0642-4
Zhang W, Kataoka M, Doan HY, Wu FT, Haga K, Takeda N, Muramatsu M, Li TC (2020) Isolation and characterization of mammalian orthoreoviruses using a cell line resistant to Sapelovirus infection. Transbound Emerg Dis 67:2849–2859. https://doi.org/10.1111/tbed.13655
Castilla D, Escobar V, Ynga S, Llanco L, Manchego A, Lázaro C, Navarro D, Santos N, Rojas M (2021) Enteric viral infections among Domesticated South American Camelids: First detection of mammalian orthoreovirus in camelids. Animals (Basel) 11(5):1455. https://doi.org/10.3390/ani11051455
Matthijnssens J, Ciarlet M, Heiman E, Arijs I, Delbeke T, McDonald SM, Palombo EA, Iturriza-Gomara M, Maes P, Patton JT, Rahman M, Van Ranst M (2008) Full genome-based classification of rotaviruses reveals a common origin between human Wa-Like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J Virol 82:3204–3219. https://doi.org/10.1128/JVI.02257-07
Matthijnssens J, Ciarlet M, McDonald SM, Attoui H, Banyai K, Brister JR, Buesa J, Esona MD, Estes MK, Gentsch JR, Iturriza-Gomara M, Johne R, Kirkwood CD, Martella V, Mertens PP, Nakagomi O, Parreno V, Rahman M, Ruggeri FM, Saif LJ, Santos N, Steyer A, Taniguchi K, Patton JT, Desselberger U, Van Ranst M (2011) Uniformity of rotavirus strain nomenclature proposed by the Rotavirus Classification Working Group (RCWG). Arch Virol 156:1397–1413. https://doi.org/10.1007/s00705-011-1006-z
Kokubu T, Takahashi T, Takamura K, Yasuda H, Hiramatsu K, Nakai M (1993) Isolation of reovirus type 3 from dogs with diarrhea. J Vet Med Sci 55:453–454. https://doi.org/10.1292/jvms.55.453
Meng XJ, Lindsay DS, Sriranganathan N (2009) Wild boars as sources for infectious diseases in livestock and humans. Philos Trans R Soc Lond B Biol Sci 364:2697–2707. https://doi.org/10.1098/rstb.2009.0086
Ohdachi S, Ishibashi Y, Iwasa MA, Saitoh T (eds) (2009) The wild mammals of Japan. Shoukadoh Book Sellers, Kyoto, p 544
Postel A, Nishi T, Kameyama KI, Meyer D, Suckstorff O, Fukai K, Becher P (2019) Reemergence of classical swine fever. Japan, Emerg Infect Dis 25:1228–1231. https://doi.org/10.3201/eid2506.181578
Yamazaki Y, Adachi F, Sawamura A (2016) Multiple origins and admixture of recently expanding Japanese wild boar (Sus scrofa leucomystax) populations in Toyama Prefecture of Japan. Zool Sci 33:38–43. https://doi.org/10.2108/zs150092
Hoxie I (2020) Dennehy JJ (2020) Intragenic recombination influences rotavirus diversity and evolution. Virus Evol 6(1):vez059. https://doi.org/10.1093/ve/vez059
Cao D, Barro M, Hoshino Y (2008) Porcine rotavirus bearing an aberrant gene stemming from an intergenic recombination of the NSP2 and NSP5 genes is defective and interfering. J Virol 82:6073–6077. https://doi.org/10.1128/JVI.00121-08
Oki H, Masuda T, Hayashi-Miyamoto M, Kawai M, Ito M, Madarame H, Fukase Y, Takemae H, Sakaguchi S, Furuya T, Mizutani T, Oba M, Nagai M (2022) Genomic diversity and intragenic recombination of species C rotaviruses. J Gen Virol. https://doi.org/10.1099/jgv.0.001703
Smith SC, Gribble J, Diller JR, Wiebe MA, Thoner TW Jr, Denison MR, Ogden KM (2021) Reovirus RNA recombination is sequence directed and generates internally deleted defective genome segments during passage. J Virol 95(8):e02181-e2220. https://doi.org/10.1128/JVI.02181-20
Acknowledgements
This work was supported by the JSPS KAKENHI (Grant numbers 18K05977 and 21K05947).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Research involving human participants and/or animals
This study did not involve any human participants and animals.
Additional information
Handling Editor: Zhenhai Chen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
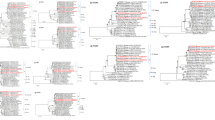
Phylogenetic analysis based on nearly complete L1 (A), L2 (B), L3 (C), M1 (D), M2 (E), M3 (F), S1 (G), S2 (H), S3 (I), and S4 (J) gene nucleotide sequences of Japanese porcine MRVs and MRV strains obtained from the DDBJ/EMBL/GenBank databases. The phylogenetic trees were constructed using the maximum-likelihood method in MEGA7 with best-fit models (the GTR+G+I model for the L1, L2, L3, M1, M2, M3, S2, and S3 phylogenetic trees and the GTR+G model for the S1 and S4 phylogenetic trees). Bootstrap values above 70 (1,000 replicates) are shown. The bars represent the corrected genetic distances. Japanese porcine MRVs and Japanese MRVs from humans, wild boar, lion, and sewage are indicated in red and blue, respectively. (PDF 408 KB)
Supplementary Fig. S2
(A-a) Recombination analysis of the L3 gene segment of Bat/WIV3/CHN vs. Deer/OV204/USA (yellow curve), Bat/WIV3/CHN vs. Bat/WIV5/CHN (blue-green curve), and Deer/OV204/USA vs. Bat/WIV5/CHN (purple curve). (A-b) Similarity plots of Bat/WIV3/CHN (blue-green curve) and Deer/OV204/USA (purple curve), and Bat/WIV5/CHN as a query sequence, with a sliding window of 200 nucleotides and a moving step size of 20 nucleotides. (B-a) Recombination analysis of the L3 gene segment of Pig/4560-3/USA vs. Pig/4560-1/USA (yellow curve), Pig/4560-3/USA vs. Pig/4560-2/USA (blue-green curve), and Pig/4560-1/USA vs. Pig/4560-2/USA (purple curve). (B-b) Similarity plots of Pig/4560-3/USA (yellow curve) and Pig/4560-2/USA (purple curve), and Pig/4560-1/USA as a query sequence, with a sliding window of 200 nucleotides and a moving step size of 20 nucleotides. (C-a) Recombination analysis of the M2 gene segment of Bat/WIV4/CHN vs. Bat/WIV3/CHN (yellow curve), Bat/WIV4/CHN vs. Bat/WIV5/CHN (blue-green curve), and Bat/WIV3/CHN vs. Bat/WIV5/CHN (purple curve). (C-b) Similarity plots of Bat/WIV3/CHN (blue-green curve) and Bat/WIV4/CHN (purple curve), and Bat/WIV5/CHN as a query sequence, with a sliding window of 200 nucleotides and a moving step size of 20 nucleotides. (D-a) Recombination analysis of the M2 gene segment of Pig/HLJ/CHN vs. Human/SI-MRV01/SVN (yellow curve), Pig/HLJ/CHN vs. Bat/RpMRV-YN2012/CHN (blue-green curve), and Human/SI-MRV01/SVN vs. Bat/RpMRV-YN2012/CHN (purple curve). (D-b) Similarity plots of Pig/HLJ/CHN (yellow curve) and Bat/RpMRV-YN2012/CHN (purple curve), and Human/SI-MRV01/SVN as a query sequence, with a sliding window of 200 nucleotides and a moving step size of 20 nucleotides (PDF 159 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fukase, Y., Minami, F., Masuda, T. et al. Genetic diversity, reassortment, and recombination of mammalian orthoreoviruses from Japanese porcine fecal samples. Arch Virol 167, 2643–2652 (2022). https://doi.org/10.1007/s00705-022-05602-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05602-8