Abstract
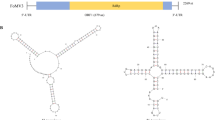
Bidens pilosa is a weed species that invades crop areas in tropical and subtropical regions. To date, only two potyviruses have been reported to infect B. pilosa. Here, we report the complete genome sequence of a tomato zonate spot tospovirus (TZSV) isolate from Bidens named TZSV-Bidens. The tripartite RNA of the TZSV-Bidens genome contains L, M, and S segments that are 8912, 4724, and 2997 nt in length, respectively. The genome contains five open reading frames (ORFs), with 92.23–95.01% amino acid sequence identity to the TZSV-YN isolate. Phylogenetic analysis based on amino acid sequences of members of the family Tospoviridae showed that TZSV-Bidens was grouped into a well-supported Eurasian cluster. The intergenic regions (IGRs) of the M and S RNAs are among the most variable regions and are far shorter than those of the TZSV-YN reference genome.


Similar content being viewed by others
Availability of data and material
Data presented in this manuscript are freely available upon request from the corresponding author.
References
Pappu HR, Jones RA, Jain RK (2009) Global status of tospovirus epidemics in diverse cropping systems: successes achieved and challenges ahead. Virus Res 141(2):219–236
Elliot RM (1990) Molecular biology of the Bunyaviridae. J Gen Virol 71:501–522
German TL, Ullman DE, Moyer JW (1992) Tospoviruses: diagnosis, molecular biology, phylogeny, and vector relationships. Annu Rev Phytopathol 30:315–348
de Haan P, Kormelink R, Resende RO, van Poelwijk F, Peters D, Goldbach R (1991) Tomato spotted wilt virus L RNA encodes a putative RNA polymerase. J Gen Virol 71:2207–2216
de Haan P, Wagemakers L, Peters D, Goldbach R (1990) The S RNA segment of tomato spotted wilt virus has an ambisense character. J Gen Virol 71:1001–1007
Kormelink R, de Haan P, Meurs C, Peters D, Goldbach R (1992) The nucleotide sequence of the M RNA segment of tomato spotted wilt virus, a bunyavirus with two ambisense RNA segments. J Gen Virol 73:2795–2804
Pappu SS, Bhat AI, Pappu HR, Deom CM, Culbreath AK (2000) Phylogenetic studies of tospoviruses (family: Bunyaviridae) based on intergenic region sequences of small and medium genomic RNAs. Arch Virol 145:1035–1045
Hallwass M, de Oliveira AS, Dianese ED, Lohuis D, Boiteux LS, Inoue-Nagata AK, Resende RO, Kormelink R (2014) The Tomato spotted wilt virus cell-to-cell movement protein (NSM) triggers a hypersensitive response in Sw-5-containing resistant tomato lines and in Nicotiana benthamiana transformed with the functional Sw-5b resistance gene copy. Mol Plant Pathol 15:871–880
Peiró A, Canizares MC, Rubio L, Lopez C, Moriones E, Aramburu J, Sánchez-Navarro J (2014) The movement protein (NSm) of Tomato spotted wilt virus is the avirulence determinant in the tomato Sw-5 gene-based resistance. Mol Plant Pathol 15:802–813
Spassova MI, Prins TW, Folkertsma RT, Klein-Lankhorst RM, Hille J, Goldbach RW (2001) The tomato gene Sw5 is a member of the coiled coil, nucleotide binding, leucine-rich repeat class of plant resistance genes and confers resistance to TSWV in tobacco. Mol Breeding 7:151–161
Zhu M, Jiang L, Bai B, Zhao W, Chen X, Li J, Liu Y, Chen Z, Wang B, Wang C et al (2017) The intracellular immune receptor Sw-5b confers broad-spectrum resistance to Tospoviruses through recognition of a conserved 21-amino acid viral effector epitope. Plant Cell 29:2214–2232
Takeda A, Sugiyama K, Nagano H, Mori M, Kaido M, Mise K, Tsuda S, Okuno T (2002) Identification of a novel RNA silencing suppressor, NSs protein of Tomato spotted wilt virus. FEBS Lett 532:75–79
Margaria P, Bosco L, Vallino M, Ciuffo M, Mautino GC, Tavella L, Turina MM (2014) The NSs protein of Tomato spotted wilt virus is required for persistent infection and transmission by Frankliniella occidentalis. J Virol 88:5788–5802
Nguyen M, Haenni AL (2003) Expression strategies of ambisense viruses. Virus Res 93(2):141–150
Baker CA, Kamenova I, Raid R, Adkins S (2007) Bidens mottle virus identified in tropical soda apple in Florida. Plant Dis 91(7):905
Sanches MM, de Marchi BR, Spadotti DM, Nozaki DN, Pavan MA, Krause-Sakate R (2014) Biological and molecular characterization of Bidens mosaic virus supports its assignment as a member of a distinct species in the genus Potyvirus. Arch Virol 159(8):2181–2183
Camelo-Garcia VM, Favara GM, Kitajima EW, Rezende JAM (2020) First report of bidens mosaic virus infecting Centella asiatica in Brazil. Plant Dis 105(2):517
Chen YK, Lee JY (2012) First report of Bidens mottle virus causing mosaic and leaf deformation in garland chrysanthemum and lettuce in Taiwan. Plant Dis 96(3):464
Jan FJ (2011) First report of Bidens mottle virus infecting calendula in Taiwan. Plant Dis 95(3):362
Liao JY, Hu CC, Chen CC, Chang CH, Deng TC (2009) Full-length sequence analysis of a distinct isolate of Bidens mottle virus infecting sunflower in Taiwan. Arch Virol 154(4):723–725
Rosales-Munar A, Alvarez-Diaz DA, Laiton-Donato K, Peláez-Carvajal D, Usme-Ciro JA (2020) Efficient method for molecular characterization of the 5’ and 3’ ends of the Dengue virus genome. Viruses 12(5):496
Jones DR (2005) Plant viruses transmitted by thrips. Eur J Plant Pathol 113:119–157
de Oliveira AS, Melo FL, Inoue-Nagata AK, Nagata T, Kitajima EW (2012) Characterization of Bean necrotic mosaic virus: a member of a novel evolutionary lineage within the genus Tospovirus. PLoS One 7(6):e38634
Acknowledgements
This work was supported by Grants from the Guizhou Province Technology R&D Program ([2019]2398), the Guizhou University Education Foundation ([2020]67), and the National Natural Science Foundation of China (Grant No. 31760503).
Funding
This work was supported by Grants from the Guizhou Province Technology R&D Program ([2019]2398), Guizhou University Education Foundation ([2020]67), and the National Natural Science Foundation of China (Grant No. 31760503).
Author information
Authors and Affiliations
Contributions
TX, LL, and MJ designed the study. TX, LL, XC, RL, XW, YL, and MJ performed the experiments. TX, LL, and MJ analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Handling Editor: Massimo Turina.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, T., Lei, L., Chen, X. et al. Identification and genome analysis of a tomato zonate spot virus isolate from Bidens pilosa. Arch Virol 167, 625–630 (2022). https://doi.org/10.1007/s00705-021-05330-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05330-5