Abstract
Anelloviruses are small negative-sense single-stranded DNA viruses with genomes ranging in size from 1.6 to 3.9 kb. The family Anelloviridae comprised 14 genera before the present changes. However, in the last five years, a large number of diverse anelloviruses have been identified in various organisms. Here, we undertake a global analysis of mammalian anelloviruses whose full genome sequences have been determined and have an intact open reading frame 1 (ORF1). We established new criteria for the classification of anelloviruses, and, based on our analyses, we establish new genera and species to accommodate the unclassified anelloviruses. We also note that based on the updated species demarcation criteria, some previously assigned species (n = 10) merge with other species. Given the rate at which virus sequence data are accumulating, and with the identification of diverse anelloviruses, we acknowledge that the taxonomy will have to be dynamic and continuously evolve to accommodate new members.
Similar content being viewed by others
Anelloviruses are circular negative-sense DNA viruses. They have genomes ranging in size from 1.6 to 3.9 kb. Anelloviruses that infect mammals have one large and two or three smaller open reading frames (ORFs) [5, 6, 71]. Gyroviruses (genus Gyrovirus) were assigned to the family Anelloviridae in 2017 [57]. Gyroviruses that have been identified primarily in avian species have at least three large ORFs.
Mammalian anelloviruses have been identified in a broad range of animals of the families Aotidae, Callitrichidae, Canidae, Cercopithecidae, Cricetidae, Didelphidae, Equidae, Felidae, Hominidae, Indriidae, Leporidae, Molossidae, Muridae, Mustelidae, Otariidae, Phocidae, Phyllostomidae, Procyonidae, Suidae, Tupaiidae, Ursidae, and Viverridae [1,2,3,4, 7, 8, 12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27, 29, 32,33,34, 39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56, 61,62,63,64,65, 72,73,74,75,76] (Supplementary Table S1). They have also been found in blood-feeding invertebrates of the families Culicidae [38] and Ixodidae [70] (likely derived from the blood meal from their host, but they do not infect these invertebrates), and in faecal samples of predators such as the South Polar skua (family Stercorariidae) [17] (Supplementary Table S1).
The family Anelloviridae was composed of 14 genera (Table 1). Except for viruses in the genus Gyrovirus, all anelloviruses had previously been classified based on pairwise identity values derived from a global alignment of nucleotide sequences of the ORF1 coding region. Previously a 65% sequence identity threshold was established for species demarcation, whereas a 44% sequence identity threshold was used for genus demarcation classification [5, 6]. The use of global alignment-derived pairwise identity values can result in the deflation of the sequence identity values due to fixed gaps. This becomes problematic, especially when diverse sequences are included in such alignments. Thus, here, we use true pairwise sequence identity values to determine their distribution in order to establish species demarcation thresholds. We analysed the annotated ORF1 coding (complete ORF1) regions of complete anellovirus genome sequences (n = 749) available in the GenBank database (downloaded 10 July 2020). We determined the pairwise identity values for all the ORF1 nucleotide sequences using SDT v1.2 [37]. The plot of the distribution of the pairwise identity values (Fig. 1) revealed a trough at ~69%.
Distribution of pairwise identity values of the ORF1 nucleotide sequences of anellovirus genomes available in the GenBank database (n = 749), determined using SDT v1.2 [37]
Using 69% as a species demarcation threshold, of the 75 currently established species, viruses in only 10 species did not fit this criterion, and thus these species were abolished and the corresponding viruses were reassigned to already established species (see Table 2 for abolished species and the reassignment of viruses). The 69% species demarcation threshold has also been used for the classification of the viruses in the genus Gyrovirus, which now has nine new species (see Kraberger et al. [30]).
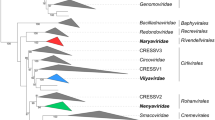
To establish genus demarcation criteria, since the pairwise identity plot does not give a clear demarcation threshold, we opted for a phylogeny-based approach using the ORF1 amino acid sequences. A dataset of the ORF1 amino acid sequences of a representative member of each species was assembled, and this was then aligned using MAFFT [28]. The alignment was trimmed with TrimAL [11] using the gappyout option, and the alignment was used to infer a maximum-likelihood tree with IQTree [35] with the LG+F+G4 substitution model. The TrimAL alignment contained 361 amino acid sites in the final alignment. The resulting maximum-likelihood phylogenetic tree was rooted at the midpoint and edited in iTOL v4 [31]. Based on the phylogeny (Fig. 2), we established a total of 16 new genera (Table 1).
Maximum-likelihood phylogenetic tree of the ORF1-encoded amino acid sequences of representative member of each species in the family Anelloviridae. The sequences of gyroviruses are not included, as their VP1 is not homologous to ORF1. Numbers at the nodes represent aLRT branch support values. Branches with less than 60% support have been collapsed with TreeGraph2 [60]
We used the Greek alphabet for naming new genera. In the spirit of using ancient alphabets, we adopt the Phoenician alphabet for an additional six new genera in series without using ‘bet’ (to avoid confusion with "beta"). In the future, the following letters of the Phoenician alphabet can be used for new genus names with minimal conflict with current names: yod, lamed, mem, samek, ayin, pe, sade, qop, res, sin, taw.
Furthermore, based on the criteria discussed above, we have established 80 new species (see Table 3). A comprehensive list of the new classification of anelloviruses is provided in Supplementary Table S1. This includes all sequences with sufficient information for their classification available in GenBank on 20 May 2021. The new species (n = 9) associated with the genus Gyrovirus are discussed in detail in Kraberger et al. [30].
We recommend that true pairwise identity determination tools be used for determining anellovirus species assignments. We also recommend the following guideline for determining new species of mammalian viruses within the family Anelloviridae, which aligns with previous recommendations for full-genome pairwise identity-based classification for single-stranded DNA satellite molecules in the family Alphasatellitidae [9] and viruses in the families Circoviridae [57], Geminiviridae [10, 36, 66, 67], Genomoviridae [68], and Smacoviridae [69].
-
1.
If the complete ORF1 coding region nucleotide sequence of a new anellovirus shares >69% pairwise identity with that of any member assigned to a currently classified anellovirus species, the virus belongs to that particular species.
-
a.
In the event that the complete ORF1 coding region nucleotide sequence of a new anellovirus has >69% pairwise identity to those of members of more than one anellovirus species, the virus should be considered a member of the species with whose members it shares the highest percentage ORF1 coding region nucleotide sequence pairwise identity.
-
b.
In the event that the complete ORF1 coding region nucleotide sequence of a new anellovirus has >69% pairwise identity to that of one or more members assigned to a particular anellovirus species, even if it shares <69% identity with those of the majority of the members assigned to that particular anellovirus species, the virus should nevertheless be considered a member of that particular species.
-
a.
-
2.
If the complete ORF1 coding region nucleotide sequence of a new anellovirus has <69% pairwise identity to those of all members of currently classified anellovirus species, the virus should be considered a member of a new species.
We anticipate a few more changes to the taxonomy of the family Anelloviridae, especially in the light of diverse new sequences being deposited from various studies and large viral metagenomic projects using high-throughput sequencing approaches. We also would like to highlight that metagenomic sequence-derived genomes can be classified [59].
We would also like to inform the anellovirus research community that the International Committee on Taxonomy of Viruses (ICTV) has ratified the adoption of standardized binomial virus species names, which can be either in Latinized or free-form format [58]. In establishing the nine new species in the genus Gyrovirus, we adopted a binomial “Genus + freeform epithet” species nomenclature (see Kraberger et al. [30]). We plan to adopt the binomial species nomenclature for all species in the family Anelloviridae by the year 2023. Thus, we encourage the community to engage with the ICTV Anelloviridae Study Group to determine the binomial names for current and new species in the family Anelloviridae.
References
Agueda-Pinto A, Kraberger S, Lund MC, Gortazar C, McFadden G, Varsani A, Esteves PJ (2020) Coinfections of novel polyomavirus, anelloviruses and a recombinant strain of myxoma virus-MYXV-Tol identified in Iberian Hares. Viruses 12:340
Amatya R, Deem SL, Porton IJ, Wang D, Lim ES (2017) Complete genome sequence of torque teno indri virus 1, a novel anellovirus in blood from a free-living lemur. Genome Announc 5:e00698-17
Biagini P, Gallian P, Attoui H, Touinssi M, Cantaloube JF, de Micco P, de Lamballerie X (2001) Genetic analysis of full-length genomes and subgenomic sequences of TT virus-like mini virus human isolates. J Gen Virol 82:379–383
Biagini P, Uch R, Belhouchet M, Attoui H, Cantaloube JF, Brisbarre N, de Micco P (2007) Circular genomes related to anelloviruses identified in human and animal samples by using a combined rolling-circle amplification/sequence-independent single primer amplification approach. J Gen Virol 88:2696–2701
Biagini P (2009) Classification of TTV and related viruses (anelloviruses). In: Villiers E-M, Hausen H (eds) TT Viruses. Springer, Berlin Heidelberg, pp 21–33
Biagini P, Bendinelli M, Hino S, Kakkola L, Mankertz A, Niel C, Okamoto H, Raidal S, Teo CG, Todd D (2012) Family anelloviridae. In: King AMQ, Adams EB, Carstens EB, EJ L (eds) Virus taxonomy: ninth report of the international committee on taxonomy of viruses. Academic press, London, pp 331–341
Bodewes R, Rubio Garcia A, Wiersma LC, Getu S, Beukers M, Schapendonk CM, van Run PR, van de Bildt MW, Poen MJ, Osinga N, Sanchez Contreras GJ, Kuiken T, Smits SL, Osterhaus AD (2013) Novel B19-like parvovirus in the brain of a harbor seal. PLoS ONE 8:e79259
Bodewes R, Contreras GJS, Garcia AR, Hapsari R, van de Bildt MWG, Kuiken T, Osterhaus A (2015) Identification of DNA sequences that imply a novel gammaherpesvirus in seals. J Gen Virol 96:1109–1114
Briddon RW, Martin DP, Roumagnac P, Navas-Castillo J, Fiallo-Olive E, Moriones E, Lett JM, Zerbini FM, Varsani A (2018) Alphasatellitidae: a new family with two subfamilies for the classification of geminivirus- and nanovirus-associated alphasatellites. Arch Virol 163:2587–2600
Brown JK, Zerbini FM, Navas-Castillo J, Moriones E, Ramos-Sobrinho R, Silva JC, Fiallo-Olive E, Briddon RW, Hernandez-Zepeda C, Idris A, Malathi VG, Martin DP, Rivera-Bustamante R, Ueda S, Varsani A (2015) Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch Virol 160:1593–1619
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973
Cibulski SP, Teixeira TF, de Sales Lima FE, do Santos HF, Franco AC, Roehe PM (2014) A Novel anelloviridae species detected in tadarida brasiliensis bats: first sequence of a chiropteran anellovirus. Genome Announc 2:e01028-14
Cornelissen-Keijsers V, Jimenez-Melsio A, Sonnemans D, Cortey M, Segales J, van den Born E, Kekarainen T (2012) Discovery of a novel Torque teno sus virus species: genetic characterization, epidemiological assessment and disease association. J Gen Virol 93:2682–2691
Crane A, Goebel ME, Kraberger S, Stone AC, Varsani A (2018) Novel anelloviruses identified in buccal swabs of Antarctic fur seals. Virus Genes 54:719–723
de Souza WM, Fumagalli MJ, de Araujo J, Sabino-Santos G Jr, Maia FGM, Romeiro MF, Modha S, Nardi MS, Queiroz LH, Durigon EL, Nunes MRT, Murcia PR, Figueiredo LTM (2018) Discovery of novel anelloviruses in small mammals expands the host range and diversity of the Anelloviridae. Virology 514:9–17
Eibach D, Hogan B, Sarpong N, Winter D, Struck NS, Adu-Sarkodie Y, Owusu-Dabo E, Schmidt-Chanasit J, May J, Cadar D (2019) Viral metagenomics revealed novel betatorquevirus species in pediatric inpatients with encephalitis/meningoencephalitis from Ghana. Sci Rep 9:2360
Fahsbender E, Burns JM, Kim S, Kraberger S, Frankfurter G, Eilers AA, Shero MR, Beltran R, Kirkham A, McCorkell R, Berngartt RK, Male MF, Ballard G, Ainley DG, Breitbart M, Varsani A (2017) Diverse and highly recombinant anelloviruses associated with Weddell seals in Antarctica. Virus Evol 3:vex017
Galmes J, Li Y, Rajoharison A, Ren L, Dollet S, Richard N, Vernet G, Javouhey E, Wang J, Telles JN, Paranhos-Baccala G (2013) Potential implication of new torque teno mini viruses in parapneumonic empyema in children. Eur Respir J 42:470–479
Hallett RL, Clewley JP, Bobet F, McKiernan PJ, Teo CG (2000) Characterization of a highly divergent TT virus genome. J Gen Virol 81:2273–2279
Heller F, Zachoval R, Koelzer A, Nitschko H, Froesner GG (2001) Isolate KAV: a new genotype of the TT-virus family. Biochem Biophys Res Commun 289:937–941
Hijikata M, Takahashi K, Mishiro S (1999) Complete circular DNA genome of a TT virus variant (isolate name SANBAN) and 44 partial ORF2 sequences implicating a great degree of diversity beyond genotypes. Virology 260:17–22
Hrazdilova K, Slaninkova E, Brozova K, Modry D, Vodicka R, Celer V (2016) New species of Torque Teno miniviruses infecting gorillas and chimpanzees. Virology 487:207–214
Inami T, Obara T, Moriyama M, Arakawa Y, Abe K (2000) Full-length nucleotide sequence of a simian TT virus isolate obtained from a chimpanzee: evidence for a new TT virus-like species. Virology 277:330–335
Jazaeri Farsani SM, Jebbink MF, Deijs M, Canuti M, van Dort KA, Bakker M, Grady BP, Prins M, van Hemert FJ, Kootstra NA, van der Hoek L (2013) Identification of a new genotype of Torque Teno Mini virus. Virol J 10:323
Kamahora T, Hino S, Miyata H (2000) Three spliced mRNAs of TT virus transcribed from a plasmid containing the entire genome in COS1 cells. J Virol 74:9980–9986
Kang YJ, Zhou MF, Huang W, Deng C, Yan G, Lu ZH (2017) Identification of a novel torque teno mini virus in cerebrospinal fluid from a child with encephalitis. Virol Sin 32:541–544
Kapusinszky B, Mulvaney U, Jasinska AJ, Deng X, Freimer N, Delwart E (2015) Local virus extinctions following a host population bottleneck. J Virol 89:8152–8161
Katoh K, Standley DM (2016) A simple method to control over-alignment in the MAFFT multiple sequence alignment program. Bioinformatics 32:1933–1942
Khalifeh A, Blumstein DT, Fontenele RS, Schmidlin K, Richet C, Kraberger S, Varsani A (2021) Diverse cressdnaviruses and an anellovirus identified in the fecal samples of yellow-bellied marmots. Virology 554:89–96
Kraberger S, Opriessnig T, Celer V, Maggi F, Okamoto H, Blomström A-L, Cadar D, Harrach B, Biagini P, Varsani A (2021) Taxonomic updates for the genus Gyrovirus (family Anelloviridae): recognition of several new members and establishment of species demarcation criteria. Arch Virol. https://doi.org/10.1007/s00705-021-05194-9
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259
Li L, Giannitti F, Low J, Keyes C, Ullmann LS, Deng X, Aleman M, Pesavento PA, Pusterla N, Delwart E (2015) Exploring the virome of diseased horses. J Gen Virol 96:2721–2733
Lu L, Robertson G, Ashworth J, Pham Hong A, Shi T, Ivens A, Thwaites G, Baker S, Woolhouse M (2020) Epidemiology and phylogenetic analysis of viral respiratory infections in Vietnam. Front Microbiol 11:833
Luo K, He H, Liu Z, Liu D, Xiao H, Jiang X, Liang W, Zhang L (2002) Novel variants related to TT virus distributed widely in China. J Med Virol 67:118–126
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534
Muhire B, Martin DP, Brown JK, Navas-Castillo J, Moriones E, Zerbini FM, Rivera-Bustamante R, Malathi VG, Briddon RW, Varsani A (2013) A genome-wide pairwise-identity-based proposal for the classification of viruses in the genus Mastrevirus (family Geminiviridae). Arch Virol 158:1411–1424
Muhire BM, Varsani A, Martin DP (2014) SDT: a virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE 9:e108277
Ng TF, Willner DL, Lim YW, Schmieder R, Chau B, Nilsson C, Anthony S, Ruan Y, Rohwer F, Breitbart M (2011) Broad surveys of DNA viral diversity obtained through viral metagenomics of mosquitoes. PLoS ONE 6:e20579
Ng TFF, Suedmeyer WK, Wheeler E, Gulland F, Breitbart M (2009) Novel anellovirus discovered from a mortality event of captive California sea lions. J Gen Virol 90:1256–1261
Ng TFF, Wheeler E, Greig D, Waltzek TB, Gulland F, Breitbart M (2011) Metagenomic identification of a novel anellovirus in Pacific harbor seal (Phoca vitulina richardsii) lung samples and its detection in samples from multiple years. J Gen Virol 92:1318–1323
Ng TFF, Dill JA, Camus AC, Delwart E, Van Meir EG (2017) Two new species of betatorqueviruses identified in a human melanoma that metastasized to the brain. Oncotarget 8:105800–105808
Niel C, Diniz-Mendes L, Devalle S (2005) Rolling-circle amplification of Torque teno virus (TTV) complete genomes from human and swine sera and identification of a novel swine TTV genogroup. J Gen Virol 86:1343–1347
Ninomiya M, Nishizawa T, Takahashi M, Lorenzo FR, Shimosegawa T, Okamoto H (2007) Identification and genomic characterization of a novel human torque teno virus of 3.2 kb. J Gen Virol 88:1939–1944
Ninomiya M, Takahashi M, Shimosegawa T, Okamoto H (2007) Analysis of the entire genomes of fifteen torque teno midi virus variants classifiable into a third group of genus Anellovirus. Arch Virol 152:1961–1975
Ninomiya M, Takahashi M, Hoshino Y, Ichiyama K, Simmonds P, Okamoto H (2009) Analysis of the entire genomes of torque teno midi virus variants in chimpanzees: infrequent cross-species infection between humans and chimpanzees. J Gen Virol 90:347–358
Nishiyama S, Dutia BM, Stewart JP, Meredith AL, Shaw DJ, Simmonds P, Sharp CP (2014) Identification of novel anelloviruses with broad diversity in UK rodents. J Gen Virol 95:1544–1553
Nishiyama S, Dutia BM, Sharp CP (2015) Complete genome sequences of novel anelloviruses from laboratory rats. Genome Announc 3:e01262-14
Nishizawa T, Sugimoto Y, Takeda T, Kodera Y, Hatano Y, Takahashi M, Okamoto H (2018) Identification and whole genome characterization of novel anelloviruses in masked palm civets (Paguma larvata): segregation into four distinct clades. Virus Res 256:183–191
Okamoto H, Nishizawa T, Tawara A, Peng Y, Takahashi M, Kishimoto J, Tanaka T, Miyakawa Y, Mayumi M (2000) Species-specific TT viruses in humans and nonhuman primates and their phylogenetic relatedness. Virology 277:368–378
Okamoto H, Nishizawa T, Takahashi M, Asabe S, Tsuda F, Yoshikawa A (2001) Heterogeneous distribution of TT virus of distinct genotypes in multiple tissues from infected humans. Virology 288:358–368
Okamoto H, Nishizawa T, Takahashi M, Tawara A, Peng Y, Kishimoto J, Wang Y (2001) Genomic and evolutionary characterization of TT virus (TTV) in tupaias and comparison with species-specific TTVs in humans and non-human primates. J Gen Virol 82:2041–2050
Okamoto H, Takahashi M, Nishizawa T, Tawara A, Fukai K, Muramatsu U, Naito Y, Yoshikawa A (2002) Genomic characterization of TT viruses (TTVs) in pigs, cats and dogs and their relatedness with species-specific TTVs in primates and tupaias. J Gen Virol 83:1291–1297
Pan S, Yu T, Wang Y, Lu R, Wang H, Xie Y, Feng X (2018) Identification of a Torque Teno Mini Virus (TTMV) in Hodgkin’s Lymphoma Patients. Front Microbiol 9:1680
Parras-Molto M, Suarez-Rodriguez P, Eguia A, Aguirre-Urizar JM, Lopez-Bueno A (2014) Genome sequence of two novel species of torque teno minivirus from the human oral cavity. Genome Announc 2:e00868-14
Peng YH, Nishizawa T, Takahashi M, Ishikawa T, Yoshikawa A, Okamoto H (2002) Analysis of the entire genomes of thirteen TT virus variants classifiable into the fourth and fifth genetic groups, isolated from viremic infants. Arch Virol 147:21–41
Qiu J, Kakkola L, Cheng F, Ye C, Soderlund-Venermo M, Hedman K, Pintel DJ (2005) Human circovirus TT virus genotype 6 expresses six proteins following transfection of a full-length clone. J Virol 79:6505–6510
Rosario K, Breitbart M, Harrach B, Segales J, Delwart E, Biagini P, Varsani A (2017) Revisiting the taxonomy of the family Circoviridae: establishment of the genus Cyclovirus and removal of the genus Gyrovirus. Arch Virol 162:1447–1463
Siddell SG, Walker PJ, Lefkowitz EJ, Mushegian AR, Dutilh BE, Harrach B, Harrison RL, Junglen S, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Nibert ML, Rubino L, Sabanadzovic S, Simmonds P, Varsani A, Zerbini FM, Davison AJ (2020) Binomial nomenclature for virus species: a consultation. Arch Virol 165:519–525
Simmonds P, Adams MJ, Benko M, Breitbart M, Brister JR, Carstens EB, Davison AJ, Delwart E, Gorbalenya AE, Harrach B, Hull R, King AM, Koonin EV, Krupovic M, Kuhn JH, Lefkowitz EJ, Nibert ML, Orton R, Roossinck MJ, Sabanadzovic S, Sullivan MB, Suttle CA, Tesh RB, van der Vlugt RA, Varsani A, Zerbini FM (2017) Consensus statement: virus taxonomy in the age of metagenomics. Nat Rev Microbiol 15:161–168
Stover BC, Muller KF (2010) TreeGraph 2: combining and visualizing evidence from different phylogenetic analyses. BMC Bioinformatics 11:7
Takahashi K, Hijikata M, Samokhvalov EI, Mishiro S (2000) Full or near full length nucleotide sequences of TT virus variants (Types SANBAN and YONBAN) and the TT virus-like mini virus. Intervirology 43:119–123
Takahashi K, Iwasa Y, Hijikata M, Mishiro S (2000) Identification of a new human DNA virus (TTV-like mini virus, TLMV) intermediately related to TT virus and chicken anemia virus. Arch Virol 145:979–993
Tisza MJ, Pastrana DV, Welch NL, Stewart B, Peretti A, Starrett GJ, Pang YS, Krishnamurthy SR, Pesavento PA, McDermott DH, Murphy PM, Whited JL, Miller B, Brenchley J, Rosshart SP, Rehermann B, Doorbar J, Ta’ala BA, Pletnikova O, Troncoso JC, Resnick SM, Bolduc B, Sullivan MB, Varsani A, Segall AM, Buck CB (2020) Discovery of several thousand highly diverse circular DNA viruses. eLife 9: e51971
Ukita M, Okamoto H, Nishizawa T, Tawara A, Takahashi M, Iizuka H, Miyakawa Y, Mayumi M (2000) The entire nucleotide sequences of two distinct TT virus (TTV) isolates (TJN01 and TJN02) remotely related to the original TTV isolates. Arch Virol 145:1543–1559
van den Brand JM, van Leeuwen M, Schapendonk CM, Simon JH, Haagmans BL, Osterhaus AD, Smits SL (2012) Metagenomic analysis of the viral flora of pine marten and European badger feces. J Virol 86:2360–2365
Varsani A, Martin DP, Navas-Castillo J, Moriones E, Hernandez-Zepeda C, Idris A, Murilo Zerbini F, Brown JK (2014) Revisiting the classification of curtoviruses based on genome-wide pairwise identity. Arch Virol 159:1873–1882
Varsani A, Navas-Castillo J, Moriones E, Hernandez-Zepeda C, Idris A, Brown JK, Murilo Zerbini F, Martin DP (2014) Establishment of three new genera in the family Geminiviridae: Becurtovirus, Eragrovirus and Turncurtovirus. Arch Virol 159:2193–2203
Varsani A, Krupovic M (2017) Sequence-based taxonomic framework for the classification of uncultured single-stranded DNA viruses of the family Genomoviridae. Virus Evol 3:vew037
Varsani A, Krupovic M (2018) Smacoviridae: a new family of animal-associated single-stranded DNA viruses. Arch Virol 163:2005–2015
Waits K, Edwards MJ, Cobb IN, Fontenele RS, Varsani A (2018) Identification of an anellovirus and genomoviruses in ixodid ticks. Virus Genes 54:155–159
Webb B, Rakibuzzaman A, Ramamoorthy S (2020) Torque teno viruses in health and disease. Virus Res 285:198013
Zhang W, Li L, Deng X, Kapusinszky B, Pesavento PA, Delwart E (2014) Faecal virome of cats in an animal shelter. J Gen Virol 95:2553–2564
Zhang W, Wang H, Wang Y, Liu Z, Li J, Guo L, Yang S, Shen Q, Zhao X, Cui L, Hua X (2016) Identification and genomic characterization of a novel species of feline anellovirus. Virol J 13:146
Zhang W, Yang S, Shan T, Hou R, Liu Z, Li W, Guo L, Wang Y, Chen P, Wang X, Feng F, Wang H, Chen C, Shen Q, Zhou C, Hua X, Cui L, Deng X, Zhang Z, Qi D, Delwart E (2017) Virome comparisons in wild-diseased and healthy captive giant pandas. Microbiome 5:90
Zhang Y, Li F, Shan TL, Deng X, Delwart E, Feng XP (2016) A novel species of torque teno mini virus (TTMV) in gingival tissue from chronic periodontitis patients. Sci Rep 6:26739
Zhu CX, Shan TL, Cui L, Luo XN, Liu ZJ, Tang SD, Liu ZW, Yuan CL, Lan DL, Zhao W, Hua XG (2011) Molecular detection and sequence analysis of feline Torque teno virus (TTV) in China. Virus Res 156:13–16
Acknowledgement
B.H. is supported by the National Research, Development and Innovation Office — NKFIH (NN128309).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Varsani, A., Opriessnig, T., Celer, V. et al. Taxonomic update for mammalian anelloviruses (family Anelloviridae). Arch Virol 166, 2943–2953 (2021). https://doi.org/10.1007/s00705-021-05192-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05192-x