Abstract
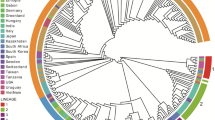
Canine circovirus (canineCV) has been found to be associated with vasculitis, hemorrhage, hemorrhagic enteritis, and diarrhea of canines. CanineCV, like other circoviruses, may also be associated with lymphoid depletion and immunosuppression. This circovirus has been detected worldwide in different countries and species. Recombination and mutation events in the canineCV genome have been described, indicating that the virus is continuing to evolve. However, the origin, codon usage patterns, and host adaptation of canineCV remain to be studied. Here, the coding sequences of 93 canineCV sequences available in the GenBank database were used for analysis. The results showed that canineCV sequences could be classified into five genotypes, as confirmed by phylogenetic and principal component analysis (PCA). Maximum clade credibility (MCC) and maximum-likelihood (ML) trees suggested that canineCV originated from bat circovirus. G/T and A/C nucleotide biases were observed in ORF1 and ORF2, respectively, and a low codon usage bias (CUB) was found in canineCV using an effective number of codon (ENC) analysis. Correlation analysis, ENC plot analysis and neutrality plot analysis indicated that the codon usage pattern was mainly shaped by natural selection. Codon adaptation index (CAI) analysis, relative codon deoptimization index (RCDI) analysis, and similarity index (SiD) analysis revealed a better adaption to Vulpes vulpes than to Canis familiaris. Furthermore, a cross-species transmission hypothesis that canineCV may have evolved from bats (origin analysis) and subsequently adapted to wolves, arctic foxes, dogs, and red foxes, was proposed. This study contributes to our understanding of the factors related to canineCV evolution and host adaption.








Similar content being viewed by others
References
Kapoor A, Dubovi EJ, Henriquez-Rivera JA, Lipkin WI (2012) Complete genome sequence of the first canine circovirus. J Virol 86:7018. https://doi.org/10.1128/JVI.00791-12
Li L, McGraw S, Zhu K, Leutenegger CM, Marks SL, Kubiski S, Gaffney P, Dela Cruz FN, Wang C, Delwart E, Pesavento PA (2013) Circovirus in tissues of dogs with vasculitis and hemorrhage. Emerg Infect Dis 19:534–541. https://doi.org/10.3201/eid1904.121390
Decaro N, Martella V, Desario C, Lanave G, Circella E, Cavalli A, Elia G, Camero M, Buonavoglia C (2014) Genomic characterization of a circovirus associated with fatal hemorrhagic enteritis in dog, Italy. PLoS ONE 9:e105909. https://doi.org/10.1371/journal.pone.0105909
Hsu HS, Lin TH, Wu HY, Lin LS, Chung CS, Chiou MT, Lin CN (2016) High detection rate of dog circovirus in diarrheal dogs. BMC Vet Res 12:116. https://doi.org/10.1186/s12917-016-0722-8
Anderson A, Hartmann K, Leutenegger CM, Proksch AL, Mueller RS, Unterer S (2017) Role of canine circovirus in dogs with acute haemorrhagic diarrhoea. Vet Rec 180:542. https://doi.org/10.1136/vr.103926
Thaiwong T, Wise AG, Maes RK, Mullaney T, Kiupel M (2016) Canine circovirus 1 (CaCV-1) and canine parvovirus 2 (CPV-2): recurrent dual infections in a papillon breeding colony. Vet Pathol 53:1204–1209. https://doi.org/10.1177/0300985816646430
Dowgier G, Lorusso E, Decaro N, Desario C, Mari V, Lucente MS, Lanave G, Buonavoglia C, Elia G (2017) A molecular survey for selected viral enteropathogens revealed a limited role of Canine circovirus in the development of canine acute gastroenteritis. Vet Microbiol 204:54–58. https://doi.org/10.1016/j.vetmic.2017.04.007
Wang Z, Shi Y, Wang Y, Zhao L, Cui X, Wen S, Liu H, Cui W, Chen H, Ge J (2020) Detection of antibodies against canine circovirus in naturally and experimentally infected canines by recombinant capsid enzyme-linked immunosorbent assay. Front Vet Sci 7:294. https://doi.org/10.3389/fvets.2020.00294
De Arcangeli S, Balboni A, Kaehler E, Urbani L, Verin R, Battilani M (2020) Genomic characterization of canine circovirus detected in red foxes (Vulpes vulpes) from Italy using a new real-time PCR assay. J Wildl Dis 56:239–242. https://doi.org/10.7589/2018-11-270
Urbani L, Tryland M, Ehrich D, Fuglei E, Battilani M, Balboni A (2020) Ancient origin and genetic segregation of canine circovirus infecting arctic foxes (Vulpes lagopus) in Svalbard and red foxes (Vulpes vulpes) in Northern Norway. Transbound Emerg Dis. https://doi.org/10.1111/tbed.13783.10.1111/tbed.13783
Zaccaria G, Malatesta D, Scipioni G, Di Felice E, Campolo M, Casaccia C, Savini G, Di Sabatino D, Lorusso A (2016) Circovirus in domestic and wild carnivores: an important opportunistic agent? Virology 490:69–74. https://doi.org/10.1016/j.virol.2016.01.007
Niu L, Wang Z, Zhao L, Wang Y, Cui X, Shi Y, Chen H, Ge J (2020) Detection and molecular characterization of canine circovirus circulating in northeastern China during 2014–2016. Arch Virol 165:137–143. https://doi.org/10.1007/s00705-019-04433-4
Piewbang C, Jo WK, Puff C, van der Vries E, Kesdangsakonwut S, Rungsipipat A, Kruppa J, Jung K, Baumgärtner W, Techangamsuwan S, Ludlow M, Osterhaus A (2018) Novel canine circovirus strains from Thailand: evidence for genetic recombination. Sci Rep 8:7524. https://doi.org/10.1038/s41598-018-25936-1
Sun W, Zhang H, Zheng M, Cao H, Lu H, Zhao G, Xie C, Cao L, Wei X, Bi J, Yi C, Yin G, Jin N (2019) The detection of canine circovirus in Guangxi, China. Virus Res 259:85–89. https://doi.org/10.1016/j.virusres.2018.10.021
Kotsias F, Bucafusco D, Nuñez DA, Lago Borisovsky LA, Rodriguez M, Bratanich AC (2019) Genomic characterization of canine circovirus associated with fatal disease in dogs in South America. PLoS ONE 14:e0218735. https://doi.org/10.1371/journal.pone.0218735
Palinski R, Piñeyro P, Shang P, Yuan F, Guo R, Fang Y, Byers E, Hause BM (2016) A novel porcine circovirus distantly related to known circoviruses is associated with porcine dermatitis and nephropathy syndrome and reproductive failure. J Virol 91:e01879-e1916. https://doi.org/10.1128/JVI.01879-16
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol 1:1 vev003. https://doi.org/10.1093/ve/vev003
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol 4:16. https://doi.org/10.1093/ve/vey016
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818. https://doi.org/10.1093/bioinformatics/14.9.817
Li G, He W, Zhu H, Bi Y, Wang R, Xing G, Zhang C, Zhou J, Yuen KY, Gao GF, Su S (2018) Origin, genetic diversity, and evolutionary dynamics of novel porcine circovirus 3. Adv Sci (Weinh) 5:1800275. https://doi.org/10.1002/advs.201800275
Huynh TM, Nguyen BH, Nguyen VG, Dang HA, Mai TN, Tran TH, Ngo MH, Le VT, Vu TN, Ta TK, Vo VH, Kim HK, Park BK (2014) Phylogenetic and phylogeographic analysis of porcine circovirus type 2 among pig farms in Vietnam. Transbound Emerg Dis 61:e25–e34. https://doi.org/10.1111/tbed.12066
Hassan S, Mahalingam V, Kumar V (2009) Synonymous codon usage analysis of thirty two mycobacteriophage genomes. Adv Bioinform 2009:316936. https://doi.org/10.1155/2009/316936
Sharp PM, Li WH (1986) Codon usage in regulatory genes in Escherichia coli does not reflect selection for ‘rare’ codons. Nucleic Acids Res 14:7737–7749. https://doi.org/10.1093/nar/14.19.7737
Clarke RT (1985) Theory and applications of correspondence analysis. J Anim Ecol 54:1031. https://doi.org/10.2307/4399
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132. https://doi.org/10.1016/0022-2836(82)90515-0
Uddin A (2020) Compositional features and codon usage pattern of genes associated with anxiety in human. Mol Neurobiol 57:4911–4920. https://doi.org/10.1007/s12035-020-02068-0
Deb B, Uddin A, Chakraborty S (2021) Genome-wide analysis of codon usage pattern in herpesviruses and its relation to evolution. Virus Res 292:198248. https://doi.org/10.1016/j.virusres.2020.198248
Wright F (1990) The ‘effective number of codons’ used in a gene. Gene 87:23–29. https://doi.org/10.1016/0378-1119(90)90491-9
Zhao Y, Zheng H, Xu A, Yan D, Jiang Z, Qi Q, Sun J (2016) Analysis of codon usage bias of envelope glycoprotein genes in nuclear polyhedrosis virus (NPV) and its relation to evolution. BMC Genomics 17:677. https://doi.org/10.1186/s12864-016-3021-7
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657. https://doi.org/10.1073/pnas.85.8.2653
Deb B, Uddin A, Chakraborty S (2020) Codon usage pattern and its influencing factors in different genomes of hepadnaviruses. Adv Virol 165:557–570. https://doi.org/10.1007/s00705-020-04533-6
Puigbò P, Bravo IG, Garcia-Vallve S (2008) CAIcal: a combined set of tools to assess codon usage adaptation. Biol Direct 3:38. https://doi.org/10.1186/1745-6150-3-38
Nakamura Y, Gojobori T, Ikemura T (2000) Codon usage tabulated from international DNA sequence databases: status for the year 2000. Nucleic Acids Res 28:292. https://doi.org/10.1093/nar/28.1.292
Mueller S, Papamichail D, Coleman JR, Skiena S, Wimmer E (2006) Reduction of the rate of poliovirus protein synthesis through large-scale codon deoptimization causes attenuation of viral virulence by lowering specific infectivity. J Virol 80:9687–9696. https://doi.org/10.1128/JVI.00738-06
Zhou JH, Zhang J, Sun DJ, Ma Q, Chen HT, Ma LN, Ding YZ, Liu YS (2013) The distribution of synonymous codon choice in the translation initiation region of dengue virus. PLoS ONE 8:e77239. https://doi.org/10.1371/journal.pone.0077239
Marais G, Mouchiroud D, Duret L (2001) Does recombination improve selection on codon usage? Lessons from nematode and fly complete genomes. Proc Natl Acad Sci USA 98:5688–5692. https://doi.org/10.1073/pnas.091427698
Behura SK, Severson DW (2013) Codon usage bias: causative factors, quantification methods and genome-wide patterns: with emphasis on insect genomes. Biol Rev Camb Philos Soc 88:49–61. https://doi.org/10.1111/j.1469-185X.2012.00242.x
Chen Y, Chen YF (2014) Extensive homologous recombination in classical swine fever virus: a re-evaluation of homologous recombination events in the strain AF407339. Saudi J Biol Sci 21:311–316. https://doi.org/10.1016/j.sjbs.2013.12.004
Liu X, Wu C, Chen AY (2010) Codon usage bias and recombination events for neuraminidase and hemagglutinin genes in Chinese isolates of influenza A virus subtype H9N2. Arch Virol 155:685–693. https://doi.org/10.1007/s00705-010-0631-2
Wu Z, Yang L, Ren X, He G, Zhang J, Yang J, Qian Z, Dong J, Sun L, Zhu Y, Du J, Yang F, Zhang S, Jin Q (2016) Deciphering the bat virome catalog to better understand the ecological diversity of bat viruses and the bat origin of emerging infectious diseases. ISME J 10:609–620. https://doi.org/10.1038/ismej.2015.138
Wong EH, Smith DK, Rabadan R, Peiris M, Poon LL (2010) Codon usage bias and the evolution of influenza viruses. BMC Evol Biol 10:253. https://doi.org/10.1186/1471-2148-10-253
Ma MR, Ha XQ, Ling H, Wang ML, Zhang FX, Zhang SD, Li G, Yan W (2011) The characteristics of the synonymous codon usage in hepatitis B virus and the effects of host on the virus in codon usage pattern. Virol J 8:544. https://doi.org/10.1186/1743-422X-8-544
Chen Y, Sun J, Tong X, Xu J, Deng H, Jiang Z, Jiang C, Duan J, Li J, Zhou P, Wang C (2014) First analysis of synonymous codon usage in porcine circovirus. Arch Virol 159:2145–2151. https://doi.org/10.1007/s00705-014-2015-5
Li G, Wang H, Wang S, Xing G, Zhang C, Zhang W, Liu J, Zhang J, Su S, Zhou J (2018) Insights into the genetic and host adaptability of emerging porcine circovirus 3. Virulence 9:1301–1313. https://doi.org/10.1080/21505594.2018.1492863
Liu XS, Zhang YG, Fang YZ, Wang YL (2012) Patterns and influencing factor of synonymous codon usage in porcine circovirus. Virol J 9:68. https://doi.org/10.1186/1743-422X-9-68
Carbone A, Zinovyev A, Kepes F (2003) Codon adaptation index as a measure of dominating codon bias. Bioinformatics 19:2005–2015. https://doi.org/10.1093/bioinformatics/btg272
Dave U, Srivathsan A, Kumar S (2019) Analysis of codon usage pattern in the viral proteins of chicken anaemia virus and its possible biological relevance. Infect Genet Evol 69:93–106. https://doi.org/10.1016/j.meegid.2019.01.002
He W, Zhao J, Xing G, Li G, Wang R, Wang Z, Zhang C, Franzo G, Su S, Zhou J (2019) Genetic analysis and evolutionary changes of Porcine circovirus 2. Mol Phylogenet Evol 139:106520. https://doi.org/10.1016/j.ympev.2019.106520
Funding
This work was supported by the SIPT Program of Northeast Agricultural University (202010224251) and the Project by State Key Laboratory of Veterinary Etiological Biology Foundation (SKLVEB2018KFKT012).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no competing interests exist.
Additional information
Handling Editor: Ana Cristina Bratanich.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
705_2021_5125_MOESM1_ESM.tif
Supplementary Fig. S1 Principal component analysis (PCA) based on the relative synonymous codon usage (RSCU). A genotype-specific PCA plot was constructed for canineCV ORFs (TIF 13427 KB)
705_2021_5125_MOESM2_ESM.tif
Supplementary Fig. S2 CAI analysis of two canineCV ORFs. ORF1 and ORF2 were analyzed separately according to genotype (a and b, respectively). RCDI analysis for two canineCV ORFs. ORF1 and ORF2 were analyzed separately according to genotype (c and d, respectively) (TIF 20227 KB)
705_2021_5125_MOESM7_ESM.doc
Supplementary Table 5 Correlation analysis among nucleotide composition, PCA, ENC, CAI , Aroma , GRAVY and axis1, axis2 in ORF1 and ORF2s (DOC 75 KB)
Rights and permissions
About this article
Cite this article
Wang, L., li, Y., Guo, Z. et al. Genetic changes and evolutionary analysis of canine circovirus. Arch Virol 166, 2235–2247 (2021). https://doi.org/10.1007/s00705-021-05125-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05125-8