Abstract
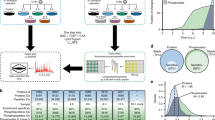
Herpesviruses are predicted to express more than 80 proteins during their infection cycle. The proteins synthesized by the immediate early genes and early genes target signaling pathways in host cells that are essential for the successful initiation of a productive infection and for latency. In this study, proteomic and phosphoproteomic tools showed the occurrence of changes in Madin-Darby bovine kidney cells at the early stage of the infection by bovine herpesvirus 1 (BoHV-1). Proteins that had already been described in the early stage of infection for other herpesviruses but not for BoHV-1 were found. For example, stathmin phosphorylation at the initial stage of infection is described for the first time. In addition, two proteins that had not been described yet in the early stages of herpesvirus infections in general were ribonuclease/angiogenin inhibitor and Rab GDP dissociation inhibitor beta. The biological processes involved in these cellular responses were repair and replication of DNA, splicing, microtubule dynamics, and inflammatory responses. These results reveal pathways that might be used as targets for designing antiviral molecules against BoHV-1 infection.
Highlights
-
BoHV-1 infection at early stages influenced various biological processes.
-
BoHV-1 infection at early stages showed proteins reported for other virus and stages.
-
Proteins not yet reported for BoHV-1 infection in the early stage were ribonuclease/angiogenin inhibitor and Rab GDP dissociation inhibitor beta






Similar content being viewed by others
Abbreviations
- 2-DE:
-
two-dimensional gel electrophoresis
- ACN:
-
acetonitrile
- ACTB:
-
beta-actin
- AHSG:
-
alpha-2-HS glycoprotein
- Apo-AI:
-
apolipoprotein A-I
- BoHV-1:
-
bovine herpesvirus 1
- BRDC:
-
bovine respiratory disease complex
- C1/C2:
-
heterogeneous nuclear ribonucleoprotein C
- CHAPS:
-
3-3-cholamidopropyl dimethylammonio-1-propanesulfonate
- CyHV-2:
-
cyprinid herpesvirus 2
- DTT:
-
dithiothreitol
- DUT:
-
deoxyuridine triphosphate
- dUTPase:
-
deoxyuridine triphosphatase
- EBNA2:
-
Epstein-Barr virus nuclear antigen 2
- EG:
-
early gene
- FA:
-
formic acid
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
- Grp78:
-
78 kDa glucose-regulated protein
- HHV-1:
-
human herpesvirus 1
- HNRNPK:
-
heterogeneous nuclear ribonucleoprotein K
- hpi:
-
hours postinfection
- HSP27:
-
heat shock protein beta-1
- HSP70:
-
heat shock protein 70
- HVEM:
-
HHV-1 entry mediator
- IEG:
-
immediate-early gene
- IPG:
-
immobilized pH gradient
- LG:
-
late gene
- MDBK:
-
Madin-Darby bovine kidney
- MEM:
-
minimum essential medium
- MOI:
-
multiplicity of infection
- MS:
-
mass spectrometry
- NPM:
-
nucleophosmin
- NPM1:
-
nucleophosmin-1
- OSF1:
-
osteoclast stimulating factor
- PCNA:
-
proliferating cell nuclear antigen
- PNP:
-
purine nucleoside phosphorylase
- PTGES3:
-
prostaglandin E synthase 3
- qPCR:
-
real-time quantitative PCR
- RNH1:
-
ribonuclease/angiogenin inhibitor 1
- RPLP0:
-
ribosomal protein large P0
- RPS18:
-
ribosomal protein S18
- SR:
-
serine/arginine-rich
- STMN1:
-
stathmin 1
- TBB:
-
tubulin beta
- TFA:
-
trifluoroacetic acid
- VHS:
-
virion host shut-off
- vRNP:
-
viral ribonucleoproteins
References
Raaperi K, Orro T, Viltrop A (2014) Epidemiology and control of bovine herpesvirus 1 infection in Europe. Vet J 201:249–256. https://doi.org/10.1016/j.tvjl.2014.05.040
Schwyzer M, Ackermann M (1996) Molecular virology of ruminant herpesviruses. Vet Microbiol 53:17–29. https://doi.org/10.1016/S0378-1135(96)01231-X
Robinson KE, Meers J, Gravel JL et al (2008) The essential and non-essential genes of Bovine herpesvirus 1. J Gen Virol 89:2851–2863. https://doi.org/10.1099/vir.0.2008/002501-0
Roizman B, Knipe DM, Whitley RJ (2007) Fields virology, 5th ed. Philadelphia
Knipe DM, Cliffe A (2008) Chromatin control of herpes simplex virus lytic and latent infection. Nat Rev Microbiol 6:211–221. https://doi.org/10.1038/nrmicro1794
Cotter CR, Kim WK, Nguyen ML et al (2011) The virion host shutoff protein of herpes simplex virus 1 blocks the replication-independent activation of NF- B in dendritic cells in the absence of type I interferon signaling. J Virol 85:12662–12672. https://doi.org/10.1128/JVI.05557-11
Liang Y, Quenelle D, Vogel JL et al (2013) Reactivation from latency A novel selective LSD1/KDM1A inhibitor epigenetically blocks. MBio 4:1–9. https://doi.org/10.1128/mBio.00558-12.Editor
Chen IB, Sciabica KS, Sandri-goldin RM (2002) ICP27 interacts with the RNA export factor Aly/REF to direct herpes simplex virus Type 1 intronless mRNAs to the TAP export pathway ICP27 interacts with the RNA export factor Aly/REF to direct herpes simplex virus Type 1 intronless mRNAs to the TAP Ex. J Virol 76:12877–12889. https://doi.org/10.1128/JVI.76.24.12877
Sciabica KS, Dai QJ, Sandri-Goldin RM (2003) ICP27 interacts with SRPK1 to mediate HSV splicing inhibition by altering SR protein phosphorylation. EMBO J 22:1608–1619. https://doi.org/10.1093/emboj/cdg166
Walsh D, Mohr I (2011) Viral subversion of the host protein synthesis machinery. Nat Rev Microbiol 9:860–875. https://doi.org/10.1038/nrmicro2655
Barzilai A, Zivony-Elbom I, Sarid R et al (2006) The herpes simplex virus Type 1 vhs-UL41 gene secures viral replication by temporarily evading apoptotic cellular response to infection: Vhs-UL41 activity might require interactions with elements of cellular mRNA degradation machinery. J Virol 80:505–513. https://doi.org/10.1128/JVI.80.1.505
Levings RL, Roth JA (2013) Immunity to bovine herpesvirus 1: I. Viral lifecycle and innate immunity. Anim Health Res Rev 14:88–102. https://doi.org/10.1017/S1466252313000042
Levings RL, Roth JA (2013) Immunity to bovine herpesvirus 1: II. Adaptive immunity and vaccinology. Anim Health Res Rev 14:103–123. https://doi.org/10.1017/S1466252313000054
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Rabilloud T, Lelong C (2011) Two-dimensional gel electrophoresis in proteomics: a tutorial. J Proteom 74:1829–1841. https://doi.org/10.1016/j.jprot.2011.05.040
Agrawal GK, Thelen JJ (2009) A high-resolution two dimensional Gel- and Pro-Q DPS-based proteomics workflow for phosphoprotein identification and quantitative profiling. Methods Mol, Biol
Neuhoff V, Arold N, Taube D, Ehrhardt W (1988) Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis 9:255–262. https://doi.org/10.1002/elps.1150090603
Shevchenko A, Tomas H, Havliš J et al (2007) In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc 1:2856–2860. https://doi.org/10.1038/nprot.2006.468
Boutet E, Lieberherr D, Tognolli M et al (2016) Uniprotkb/swiss-prot, the manually annotated section of the uniprot knowledgebase: how to use the entry view. In: Methods in molecular biology
Szklarczyk D, Franceschini A, Wyder S et al (2015) STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:D447–D452. https://doi.org/10.1093/nar/gku1003
Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25:25–29. https://doi.org/10.1038/75556
Schoen K, Plendl J, Gabler C, Kaessmeyer S (2015) Identification of stably expressed reference genes for RT-qPCR data normalization in defined localizations of cyclic bovine ovaries. J Vet Med Ser C Anat Histol Embryol 44:200–211. https://doi.org/10.1111/ahe.12128
Rekawiecki R, Rutkowska J, Kotwica J (2012) Identification of optimal housekeeping genes for examination of gene expression in bovine corpus luteum. Reprod Biol 12:362–367. https://doi.org/10.1016/j.repbio.2012.10.010
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Skiba M, Mettenleiter TC, Karger A (2008) Quantitative whole-cell proteome analysis of pseudorabies virus-infected cells. J Virol 82:9689–9699. https://doi.org/10.1128/JVI.00995-08
Guo L, Yang Y, Liu L et al (2015) A proteomic study of the differential protein expression in MDBK cells after bovine herpesvirus type 1 infection (BHV-1) strain treatment. Int J Clin Exp Med 8:4204–4211
Stokol T, Yeo WM, Burnett D et al (2015) Equid herpesvirus type 1 activates platelets. PLoS One 10:1–20. https://doi.org/10.1371/journal.pone.0122640
Quenelle DC, Hartman TL, Buckheit RW et al (2014) Anti-HSV activity of serpin antithrombin III. Int Trends Immun 2:87–92. https://doi.org/10.1016/j.biotechadv.2011.08.021.Secreted
Mathew SS, Della Selva MP, Burch AD (2009) Modification and reorganization of the cytoprotective cellular chaperone Hsp27 during herpes simplex virus type 1 infection. J Virol 83:9304–9312. https://doi.org/10.1128/JVI.01826-08
Tandon R, Mocarski ES (2012) Viral and host control of cytomegalovirus maturation. Trends Microbiol
Tanioka T, Nakatani Y, Kobayashi T et al (2003) Regulation of cytosolic prostaglandin E2synthase by 90-kDa heat shock protein. Biochem Biophys Res Commun 303:1018–1023. https://doi.org/10.1016/S0006-291X(03)00470-4
Kobayashi T, Nakatani Y, Tanioka T et al (2004) Regulation of cytosolic prostaglandin E synthase by phosphorylation. Biochem J 381:59–69. https://doi.org/10.1042/BJ20040118
Antrobus R, Grant K, Gangadharan B et al (2009) Proteomic analysis of cells in the early stages of herpes simplex virus type-1 infection reveals widespread changes in the host cell proteome. Proteomics 9:3913–3927. https://doi.org/10.1002/pmic.200900207
Chen PW, Lin SJ, Tsai SC et al (2010) Regulation of microtubule dynamics through phosphorylation on stathmin by epstein-barr virus kinase BGLF4. J Biol Chem 285:10053–10063. https://doi.org/10.1074/jbc.M109.044420
Rubin CI, Atweh GF (2004) The role of stathmin in the regulation of the cell cycle. J Cell Biochem 93:242–250. https://doi.org/10.1002/jcb.20187
Pizzo E, Sarcinelli C, Sheng J et al (2013) Ribonuclease/angiogenin inhibitor 1 regulates stress-induced subcellular localization of angiogenin and controls its growth and survival activities. J Cell Sci. https://doi.org/10.1242/jcs.134551
Li S, Hu G-F (2012) Emerging role of angiogenin in stress response and cell survival under adverse conditions. J Cell Physiol 227:2822–2826. https://doi.org/10.1002/jcp.23051
Zhu Y, Das K, Wu J et al (2014) RNH1 regulation of reactive oxygen species contributes to histone deacetylase inhibitor resistance in gastric cancer cells. Oncogene 33:1527–1537. https://doi.org/10.1038/onc.2013.104
Lomonte P, Thomas J, Texier P et al (2004) Functional interaction between class II histone deacetylases and ICP0 of herpes simplex virus type 1. J Virol 78:6744–6757. https://doi.org/10.1128/JVI.78.13.6744
Gross H, Hennard C, Masouris I et al (2012) Binding of the heterogeneous ribonucleoprotein K (hnRNP K) to the epstein-barr virus nuclear antigen 2 (EBNA2) enhances viral LMP2A expression. PLoS One 7:1–18. https://doi.org/10.1371/journal.pone.0042106
Schmidt T, Striebinger H, Haas J, Bailer SM (2010) The heterogeneous nuclear ribonucleoprotein K is important for herpes simplex virus-1 propagation. FEBS Lett 584:4361–4365. https://doi.org/10.1016/j.febslet.2010.09.038
Piñol-Roma S, Dreyfuss G (1993) Cell cycle-regulated phosphorylation of the pre-mRNA-binding (heterogeneous nuclear ribonucleoprotein) C proteins. Mol Cell Biol 13:5762–5770. https://doi.org/10.1128/MCB.13.9.5762
Sanders I, Boyer M, Fraser NW (2015) Early nucleosome deposition on, and replication of, HSV DNA requires cell factor PCNA. J Neurovirol 21:358–369. https://doi.org/10.1007/s13365-015-0321-7
Yang S, Pei Y, Zhao A (2017) ITRAQ-based Proteomic Analysis of Porcine Kidney Epithelial PK15 cells Infected with Pseudorabies virus. Sci Rep 7:1–10. https://doi.org/10.1038/srep45922
Vértessy GB, Tóth J (2009) Keeping NA: UTPases. Acc Chem Res 42:97–106
Kato A, Arii J, Koyanagi Y, Kawaguchi Y (2015) Phosphorylation of herpes simplex virus 1 dUTPase regulates viral virulence and genome integrity by compensating for low cellular dUTPase activity in the central nervous system. J Virol 89:241–248. https://doi.org/10.1128/JVI.02497-14
Sarek G, Järviluoma A, Moore HM et al (2010) Nucleophosmin phosphorylation by v-cyclin-CDK6 controls KSHV latency. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1000818
Zenner HL, Yoshimura S, Barr FA, Crump CM (2011) Analysis of Rab GTPase-activating proteins indicates that Rab1a/b and Rab43 are important for herpes simplex virus 1 secondary envelopment. J Virol 85:8012–8021. https://doi.org/10.1128/JVI.00500-11
Li S (2019) Regulation of ribosomal proteins on viral infection. Cells. https://doi.org/10.3390/cells8050508
Guo YE, Oei T, Steitz JA (2015) Herpesvirus saimiri MicroRNAs preferentially target host cell cycle regulators. J Virol. https://doi.org/10.1128/jvi.01884-15
Qiao G, Zhang M, Li Y et al (2018) Biofloc technology (BFT): an alternative aquaculture system for prevention of Cyprinid herpesvirus 2 infection in gibel carp (Carassius auratus gibelio). Fish Shellfish Immunol. https://doi.org/10.1016/j.fsi.2018.09.015
Böttcher A, Gaipl US, Fürnrohr BG et al (2006) Involvement of phosphatidylserine, αvβ3, CD14, CD36, and complement C1q in the phagocytosis of primary necrotic lymphocytes by macrophages. Arthritis Rheum 54:927–938. https://doi.org/10.1002/art.21660
Munoz LE, Frey B, Pausch F et al (2007) The role of annexin A5 in the modulation of the immune response against dying and dead cells. Curr Med Chem 14:271–277. https://doi.org/10.2174/092986707779941131
Everett RD, Murray J (2005) ND10 components relocate to sites associated with herpes simplex virus type 1 nucleoprotein complexes during virus infection. J Virol 79:5078–5089. https://doi.org/10.1128/JVI.79.8.5078
Acknowledgements
The authors thank the Brazilian Agencies Foundation for Research Support of Minas Gerais (FAPEMIG: Fellowships and Grants, PPM-00796-15), the Financier of Studies and Projects (FINEP: CT-INFRA/UFV-2004/2007/2008), the National Council for Scientific and Technological Development (CNPq: Grants: 483976/2012-1 and 455318/2014-0), and Coordination for the Improvement of Higher Education Personnel (CAPES: Fellowships) for financial support. The authors would like to thank Nucleus of Analysis of Biomolecules (NuBioMol, UFV, Viçosa-MG, Brazil) for assistance with mass spectrometric analysis, and the Institute of Biotechnology Applied to Agriculture (BIOAGRO, UFV, Viçosa-MG, Brazil) for technical support. Funding Sources in Brazil: Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) http://dx.doi.org/10.13039/501100004901. Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) http://dx.doi.org/10.13039/501100003593. Financiadora de Estudos e Projetos (FINEP) http://dx.doi.org/10.13039/501100004809. Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) http://dx.doi.org/10.13039/501100002322.
Author information
Authors and Affiliations
Contributions
MJMJ, LKJP, CEV, and KMF cultured and infected cells, performed electrophoresis assays and protein analysis. MJMJ, PSC and MRS performed real-time PCR. MCBP, ASJ, GCB, JLRF, PSC, MRS, and MRA designed and conducted the biological assays. MJMJ, MCBP, ASJ, PSC, and MRS wrote and revised the manuscript. All authors contributed and gave approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have nothing to disclose as conflicts of interest. The authors declare no competing financial interest.
Additional information
Handling Editor: Diego G. Diel.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Magalhães-Junior, M.J., Baracat-Pereira, M.C., Pereira, L.K.J. et al. Proteomic and phosphoproteomic analyses reveal several events involved in the early stages of bovine herpesvirus 1 infection. Arch Virol 165, 69–85 (2020). https://doi.org/10.1007/s00705-019-04452-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04452-1