Abstract
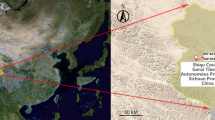
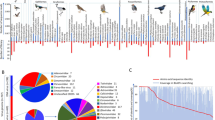
Rodent populations are known to be reservoirs of viruses with the potential to infect humans. However, a large number of such viruses remain undiscovered. In this study, we investigated the shedding of unknown viruses in long-tailed ground squirrel (Spermophilus undulatus) feces by high-throughput sequencing. A novel and highly divergent virus related to members of the genus Hepacivirus was identified in ground squirrel liver. This virus, tentatively named RHV-GS2015, was found to have a genome organization that is typical of hepaciviruses, including a long open reading frame encoding a polyprotein of 2763 aa. Sequence alignment of RHV-GS2015 with the most closely related hepaciviruses yielded p-distances of the NS3 and NS5B regions of 0.546 and 0.476, respectively, supporting the conclusion that RHV-GS2015 is a member of a new hepacivirus species, which we propose to be named “Hepacivirus P”. Phylogenetic analysis of the NS3 and NS5B regions indicated that RHV-GS2015 shares common ancestry with other rodent hepaciviruses (species Hepacivirus E, and species Hepacivirus F), Norway rat hepacivirus 1 (species Hepacivirus G), and Norway rat hepacivirus 2 (species Hepacivirus H). A phylogenetic tree including the seven previously identified rodent hepaciviruses revealed extreme genetic heterogeneity among these viruses. RHV-GS2015 was detected in 7 out of 12 ground squirrel pools and was present in liver, lung, and spleen tissues. Furthermore, livers showed extremely high viral loads of RHV-GS2015, ranging from 2.5 × 106 to 2.0 × 108 copies/g. It is reasonable to assume that this novel virus is hepatotropic, like hepatitis C virus. The discovery of RHV-GS2015 extends our knowledge of the genetic diversity and host range of hepaciviruses, helping to elucidate their origins and evolution.




Similar content being viewed by others
Reference
Drexler JF, Corman VM, Müller MA, Lukashev AN, Gmyl A et al (2013) Evidence for Novel Hepaciviruses in Rodents. PLoS Pathogens 9(6):e1003438. https://doi.org/10.1371/journal.ppat.1003438
Hartlage AS, Cullen JM, Kapoor A (2016) The strange, expanding world of animal hepaciviruses. Ann Rev Virol 3(1):53–75. https://doi.org/10.1146/annurev-virology-100114-055104
Thézé J, Lowes S, Parker J, Pybus OG (2015) Evolutionary and phylogenetic analysis of the hepaciviruses and pegiviruses. Genome Biol Evolut 7(11):2996–3008. https://doi.org/10.1093/gbe/evv202
Bukh J, Apgar CL, Yanagi M (1999) Toward a surrogate model for hepatitis C virus: an infectious molecular clone of the GB virus-B hepatitis agent. Virology. https://doi.org/10.1006/viro.1999.9941
Pybus OG, Theze J (2016) Hepacivirus cross-species transmission and the origins of the hepatitis C virus. Curr Opin Virol 16:1–7. https://doi.org/10.1016/j.coviro.2015.10.002
Scheel TKSP (2015) Kapoor A (2015) Surveying the global virome: Identification and characterization of HCV-related animal hepaciviruses. Antiviral research 115:83–93. https://doi.org/10.1016/j.antiviral.2014.12.014
Quan PL, Firth C, Conte JM, Williams SH, Zambrana-Torrelio CM et al (2013) Bats are a major natural reservoir for hepaciviruses and pegiviruses. Proc Natl Acad Sci 110(20):8194–8199. https://doi.org/10.1073/pnas.1303037110
Firth C, Bhat M, Firth MA, Williams SH, Frye MJ et al (2014) Detection of zoonotic pathogens and characterization of novel viruses carried by commensal Rattus norvegicus in New York City. mBiol 5(5):e01933–e01914. https://doi.org/10.1128/mbio.01933-14
Kapoor A, Simmonds P, Scheel TK, Hjelle B, Cullen JM et al (2013)identification of rodent homologs of hepatitis C virus and pegiviruses. mBiol 4(2):e00216–e00213. https://doi.org/10.1128/mbio.00216-13
El-Attar LM, Mitchell JA, Brooks Brownlie H, Priestnall SL, Brownlie J (2015) Detection of non-primate hepaciviruses in UK dogs. Virology 484:93–102. https://doi.org/10.1016/j.virol.2015.05.005
Lauck M, Sibley SD, Lara J, Purdy MA, Khudyakov Y et al (2013) A novel hepacivirus with an unusually long and intrinsically disordered NS5A protein in a wild Old World primate. J Virol 87(16):8971–8981. https://doi.org/10.1128/JVI.00888-13
Vercauteren K, de Jong YP, Meuleman P (2015) Animal models for the study of HCV. Curr Opin Virol 3:67–74. https://doi.org/10.1016/j.coviro.2015.04.009
Pfaender S, Brown RJ, Pietschmann T, Steinmann E (2014) Natural reservoirs for homologs of hepatitis C virus. Emerg Microb Infect 3(3):e21. https://doi.org/10.1038/emi.2014.19
Walter S, Rasche A, Moreira-Soto A et al (2017) Differential infection patterns and recent evolutionary origins of equine hepaciviruses in donkeys. J Virol 91(1):e01711–e01716. https://doi.org/10.1128/JVI.01711-16
Pfaender S, Walter S, Grabski E et al (2017) Immune protection against reinfection with nonprimate hepacivirus. Proc Natl Acad Sci USA 114(12):E2430–E2439. https://doi.org/10.1073/pnas.1619380114
Ramsay JD, Evanoff R, Wilkinson TE Jr, Divers TJ, Knowles DP et al (2015) Experimental transmission of equine hepacivirus in horses as a model for hepatitis C virus. HEPATOLOGY 61(5):1533–1546. https://doi.org/10.1002/hep.27689
Smith DB, Becher P, Bukh J, Gould EA, Meyers G et al (2016) Proposed update to the taxonomy of the genera Hepacivirus and Pegivirus within the Flaviviridae family. J Gen Virol 97(11):2894–2907. https://doi.org/10.1099/jgv.0.000612
Scheel TK, Simmonds P, Kapoor A (2015) Surveying the global virome: identification and characterization of HCV-related animal hepaciviruses. Antiviral Res 115:83–93. https://doi.org/10.1016/j.antiviral.2014.12.014
Li LL, Liu N, Humphries EM, Yu JM, Li S et al (2016) Aetiology of diarrhoeal disease and evaluation of viral-bacterial coinfection in children under 5 years old in China: a matched case–control study. Clin Microbiol Infect 22(4):381 e389–381 e416. https://doi.org/10.1016/j.cmi.2015.12.018
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evolut 1(1):1–5. https://doi.org/10.1093/ve/vev003
Rho J, Choi S, Seong YR, Choi J, Im DS (2001) The Arginine-1493 residue in QRRGRTGR1493G Motif IV of the hepatitis C Virus NS3 helicase domain is essential for NS3 protein methylation by the protein arginine methyltransferase 1. J Virol 75(17):8031–8044
Xu Z, Choi J, Yen TS, Lu W, Strohecker A et al (2001) Synthesis of a novel hepatitis C virus protein by ribosomal frameshift. EMBO J 20(14):3840–3848. https://doi.org/10.1093/emboj/20.14.3840
Chen SL, Morgan TR (2006) The natural history of hepatitis C virus (HCV) infection. Int J Med Sci 3(2):47–52
Reau N, Vekeman F, Wu E, Bao Y, Gonzalez YS (2017) Prevalence and economic burden of extrahepatic manifestations of hepatitis C virus are underestimated but can be improved with therapy. Hepatol Commun 1(5):439–452. https://doi.org/10.1002/hep4.1049
Chhatwal J, Wang X, Ayer T, Kabiri M, Chung RT et al (2016) Hepatitis C disease Burden in the United States in the era of oral direct-acting antivirals. Hepatology 64(5):1442–1450. https://doi.org/10.1002/hep.28571
Kapoor A, Simmonds P, Gerold G, Qaisar N, Jain K et al (2011) Characterization of a canine homolog of hepatitis C virus. PNAS 108(28):11608–11613. https://doi.org/10.1073/pnas.1101794108
Lyons S, Kapoor A, Sharp C, Schneider BS, Wolfe ND et al (2012) Nonprimate hepaciviruses in domestic horses, United Kingdom. Emerg Infect Dis 18(12):1976–1982. https://doi.org/10.3201/eid1812.120498
Gather T, Walter S, Pfaender S, Todt D, Feige K et al (2016) Acute and chronic infections with nonprimate hepacivirus in young horses. Vet Res 47(1):97. https://doi.org/10.1186/s13567-016-0381-6
Deng YGS, Wang S, Hao G, Rasmussen TB (2018) The detection and phylogenetic analysis of bovine hepacivirus in China. Biomed Res Int. https://doi.org/10.1155/2018/6216853
Lu GJK, Ping X, Huang J, Luo A, Wu P, Li S (2018) Novel bovine hepacivirus in dairy cattle, China. Emerg Microb Infect. https://doi.org/10.1038/s41426-018-0055-8
Baechlein C, Fischer N, Grundhoff A, Alawi M, Indenbirken D et al (2015) Identification of a novel hepacivirus in domestic cattle from Germany. J Virol 89(14):7007–7015. https://doi.org/10.1128/JVI.00534-15
Tanaka T, Otoguro T, Yamashita A et al (2018) Roles of the 5’ untranslated region of nonprimate hepacivirus in translation initiation and viral replication. J Virol 92(7):e01997-17. https://doi.org/10.1128/jvi.01997-17
Lyons S, Kapoor A, Sharp C, Schneider BS, Wolfe ND et al (2012) Nonprimate hepaciviruses in domestic horses, United Kingdom. Emerg Infect Dis 18(12):1976–1982. https://doi.org/10.3201/eid1812.120498
Hartlage AS, Cullen JM, Kapoor A (2016) The Strange, expanding world of animal hepaciviruses. Ann Rev Virol 3(1):53–75. https://doi.org/10.1146/annurev-virology-100114-055104
Yanagi M, Purcell RH, Emerson SU, Bukh J (1997) Transcripts from a single full-length cDNA clone of hepatitis C virus are infectious when directly transfected into the liver of a chimpanzee. Proc Natl Acad Sci 94(16):8738–8743
Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P (2005) Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 309(5740):1577–1581
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Funding
This work was supported by grants from the National Key R&D Program of China (2016YFC1201904), the National Science and Technology Major Project of China (No. 2018ZX10305409-004-002) and the Science and Technology Basic Work Program (2013FY113500), from the Ministry of Science and Technology of China.
Ethical statement
The study protocol was approved by the Ethics Committee of China CDC and adhered to Chinese ethics laws and regulations.
Additional information
Handling Editor: Tim Skern.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Depositories
The GenBank [/EMBL/DDBJ] accession number for the complete genome nucleotide sequence of the novel hepacivirus found in this study is MG211815.
Li-li Li and Meng-meng Liu contributed equally to the manuscript.
Rights and permissions
About this article
Cite this article
Li, Ll., Liu, Mm., Shen, S. et al. Detection and characterization of a novel hepacivirus in long-tailed ground squirrels (Spermophilus undulatus) in China. Arch Virol 164, 2401–2410 (2019). https://doi.org/10.1007/s00705-019-04303-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04303-z