Abstract
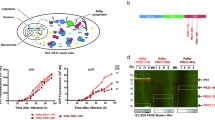
The subunits PA, PB1, and PB2 of influenza A virus RNA polymerase are essential for efficient viral RNA synthesis and virus replication because of their role in recruiting multiple nuclear proteins. ANP32A is an acidic leucine-rich nuclear phosphoprotein 32 (ANP32) family member and a crucial cellular protein that determines the species specificity of the influenza virus RNA polymerase activity. However, how ANP32A modulates polymerase activity remains largely unknown. In this study, we showed that viral RNA synthesis was increased in A549 cells overexpressing ANP32A and decreased after treatment with ANP32A RNAi. This decrease in RNA synthesis was reversed by rescued ANP32A expression. The results of docking modeling, co-immunoprecipitation, and yeast two-hybrid assays showed that PB2 was the only subunit of the three that interacted with ANP32A. The C-terminal portion of ANP32A and the middle domains (residues 307-534) of PB2 were required for PB2-ANP32A interaction. Glu189 and Glu196 in ANP32A and Gly450 and Gln447 in PB2 were essential for interaction between ANP32A and PB2. These residues were located in conserved regions of the ANP32A or PB2 protein sequences. These data suggest that ANP32A is recruited to the polymerase through direct interaction with PB2 via critical amino acid residue interactions and promotes viral RNA synthesis. Our findings might provide new insights into the molecular mechanisms underlying influenza virus RNA synthesis and replication in infected human cells.







Similar content being viewed by others
References
Te Velthuis AJ, Fodor E (2016) Influenza virus RNA polymerase: insights into the mechanisms of viral RNA synthesis. Nat Rev Microbiol 14:479–493
Fodor E (2013) The RNA polymerase of influenza a virus: mechanisms of viral transcription and replication. Acta Virol 57:113–122
Bortz E, Westera L, Maamary J, Steel J, Albrecht RA, Manicassamy B, Chase G, Martinez-Sobrido L, Schwemmle M, Garcia-Sastre A (2011) Host- and strain-specific regulation of influenza virus polymerase activity by interacting cellular proteins. MBio 2:e00151-11
Zhao M, Wang L, Li S (2017) Influenza A virus-host protein interactions control viral pathogenesis. Int J Mol Sci 18:1673
Nagata K, Kawaguchi A, Naito T (2008) Host factors for replication and transcription of the influenza virus genome. Rev Med Virol 18:247–260
Momose F, Naito T, Yano K, Sugimoto S, Morikawa Y, Nagata K (2002) Identification of Hsp90 as a stimulatory host factor involved in influenza virus RNA synthesis. J Biol Chem 277:45306–45314
Momose F, Basler CF, O’Neill RE, Iwamatsu A, Palese P, Nagata K (2001) Cellular splicing factor RAF-2p48/NPI-5/BAT1/UAP56 interacts with the influenza virus nucleoprotein and enhances viral RNA synthesis. J Virol 75:1899–1908
Wang Q, Li Q, Liu R, Zheng M, Wen J Zhao G (2016) Host cell interactome of PA protein of H5N1 influenza A virus in chicken cells. J Proteom 136:48–54
Naito T, Kiyasu Y, Sugiyama K, Kimura A, Nakano R, Matsukage A, Nagata K (2007) An influenza virus replicon system in yeast identified Tat-SF1 as a stimulatory host factor for viral RNA synthesis. Proc Natl Acad Sci USA 104:18235–18240
Cao M, Wei C, Zhao L, Wang J, Jia Q, Wang X, Jin Q, Deng T (2014) DnaJA1/Hsp40 is co-opted by influenza A virus to enhance its viral RNA polymerase activity. J Virol 88:14078–14089
Zheng W, Tao YJ (2013) Structure and assembly of the influenza A virus ribonucleoprotein complex. FEBS Lett 587:1206–1214
Dawson WK, Lazniewski M, Plewczynski D (2017) RNA structure interactions and ribonucleoprotein processes of the influenza A virus. Brief Funct Genom 17:402–414
Compans RW, Content J, Duesberg PH (1972) Structure of the ribonucleoprotein of influenza virus. J Virol 10:795–800
Coloma R, Valpuesta JM, Arranz R, Carrascosa JL, Ortin J, Martin-Benito J (2009) The structure of a biologically active influenza virus ribonucleoprotein complex. PLoS Pathog 5:e1000491
Pflug A, Lukarska M, Resa-Infante P, Reich S, Cusack S (2017) Structural insights into RNA synthesis by the influenza virus transcription-replication machine. Virus Res 234:103–117
Eisfeld AJ, Neumann G, Kawaoka Y (2015) At the centre: influenza A virus ribonucleoproteins. Nat Rev Microbiol 13:28–41
Engelhardt OG, Fodor E (2006) Functional association between viral and cellular transcription during influenza virus infection. Rev Med Virol 16:329–345
Guilligay D, Tarendeau F, Resa-Infante P, Coloma R, Crepin T, Sehr P, Lewis J, Ruigrok RW, Ortin J, Hart DJ, Cusack S (2008) The structural basis for cap binding by influenza virus polymerase subunit PB2. Nat Struct Mol Biol 15:500–506
Dias A, Bouvier D, Crepin T, McCarthy AA, Hart DJ, Baudin F, Cusack S, Ruigrok RW (2009) The cap-snatching endonuclease of influenza virus polymerase resides in the PA subunit. Nature 458:914–918
Crepin T, Dias A, Palencia A, Swale C, Cusack S, Ruigrok RW (2010) Mutational and metal binding analysis of the endonuclease domain of the influenza virus polymerase PA subunit. J Virol 84:9096–9104
Reilly PT, Yu Y, Hamiche A, Wang L (2014) Cracking the ANP32 whips: important functions, unequal requirement, and hints at disease implications. BioEssays 36:1062–1071
Tsujio I, Zaidi T, Xu J, Kotula L, Grundke-Iqbal I, Iqbal K (2005) Inhibitors of protein phosphatase-2A from human brain structures, immunocytological localization and activities towards dephosphorylation of the Alzheimer type hyperphosphorylated tau. FEBS Lett 579:363–372
Zamora-Caballero S, Siauciunaite-Gaubard L, Bravo J (2015) High-resolution crystal structure of the leucine-rich repeat domain of the human tumour suppressor PP32A (ANP32A). Acta Crystallogr Sect F Struct Biol Commun 71:684–687
Kadota S, Nagata K (2011) pp32, an INHAT component, is a transcription machinery recruiter for maximal induction of IFN-stimulated genes. J Cell Sci 124:892–899
Sugiyama K, Kawaguchi A, Okuwaki M, Nagata K (2015) pp32 and APRIL are host cell-derived regulators of influenza virus RNA synthesis from cRNA. eLife 4:e08939
Long JS, Giotis ES, Moncorge O, Frise R, Mistry B, James J, Morisson M, Iqbal M, Vignal A, Skinner MA, Barclay WS (2016) Species difference in ANP32A underlies influenza A virus polymerase host restriction. Nature 529:101–104
Mehle A (2016) The Avian influenza virus polymerase brings ANP32A home to roost. Cell Host Microbe 19:137–138
Domingues P, Hale BG (2017) Functional insights into ANP32A-dependent influenza a virus polymerase host restriction. Cell Rep 20:2538–2546
Zhou Z, Cao M, Guo Y, Zhao L, Wang J, Jia X, Li J, Wang C, Gabriel G, Xue Q, Yi Y, Cui S, Jin Q, Wang J, Deng T (2014) Fragile X mental retardation protein stimulates ribonucleoprotein assembly of influenza A virus. Nat Commun 5:3259
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43:W174–181
Huyton T, Wolberger C (2007) The crystal structure of the tumor suppressor protein pp32 (Anp32a): structural insights into Anp32 family of proteins. Protein Sci 16:1308–1315
Tuncbag N, Gursoy A, Nussinov R, Keskin O (2011) Predicting protein-protein interactions on a proteome scale by matching evolutionary and structural similarities at interfaces using PRISM. Nat Protoc 6:1341–1354
Hatada E, Hasegawa M, Mukaigawa J, Shimizu K, Fukuda R (1989) Control of influenza virus gene expression: quantitative analysis of each viral RNA species in infected cells. J Biochem 105:537–546
Meyerson NR, Zhou L, Guo YR, Zhao C, Tao YJ, Krug RM, Sawyer SL (2017) Nuclear TRIM25 specifically targets influenza virus ribonucleoproteins to block the onset of RNA chain elongation. Cell Host Microbe 22(627–638):e627
Zheng X, Wang X, Tu F, Wang Q, Fan Z, Gao G (2017) trim25 is required for the antiviral activity of zinc finger antiviral protein. J Virol 91:e00088-17
Zhu Y, Chen G, Lv F, Wang X, Ji X, Xu Y, Sun J, Wu L, Zheng YT, Gao G (2011) Zinc-finger antiviral protein inhibits HIV-1 infection by selectively targeting multiply spliced viral mRNAs for degradation. Proc Natl Acad Sci USA 108:15834–15839
Guo X, Carroll JW, Macdonald MR, Goff SP, Gao G (2004) The zinc finger antiviral protein directly binds to specific viral mRNAs through the CCCH zinc finger motifs. J Virol 78:12781–12787
Guo X, Ma J, Sun J, Gao G (2007) The zinc-finger antiviral protein recruits the RNA processing exosome to degrade the target mRNA. Proc Natl Acad Sci USA 104:151–156
Zhu Y, Wang X, Goff SP, Gao G (2012) Translational repression precedes and is required for ZAP-mediated mRNA decay. EMBO J 31:4236–4246
Thierry E, Guilligay D, Kosinski J, Bock T, Gaudon S, Round A, Pflug A, Hengrung N, El Omari K, Baudin F, Hart DJ, Beck M, Cusack S (2016) Influenza polymerase can adopt an alternative configuration involving a radical repacking of PB2 domains. Mol Cell 61:125–137
Long JX, Wang QZ, Lu JH, Liu YL, Liu XF (2005) [Cloning of full-length genes of H5N1 subtype Avian influenza virus strain A/duck/Shandong/093/2004 and analysis of the sequences]. Wei sheng wu xue bao = Acta microbiologica Sinica 45:690–696
Zhao B, Zhang X, Zhu W, Teng Z, Yu X, Gao Y, Wu D, Pei E, Yuan Z, Yang L, Wang D, Shu Y, Wu F (2014) Novel avian influenza A(H7N9) virus in tree sparrow, Shanghai, China, 2013. Emerg Infect Dis 20:850–853
El Houadfi M, Fellahi S, Nassik S, Guerin JL, Ducatez MF (2016) First outbreaks and phylogenetic analyses of avian influenza H9N2 viruses isolated from poultry flocks in Morocco. Virol J 13:140
Beerens N, Koch G, Heutink R, Harders F, Vries DPE, Ho C, Bossers A, Elbers A (2018) Novel highly pathogenic avian influenza A(H5N6) virus in the Netherlands, December 2017. Emerg Infect Dis 24:770–773
Mi Z, Liu W, Fan H, An X, Pei G, Wang W, Xu X, Ma M, Zhang Z, Cao W, Tong Y (2013) Complete genome sequence of avian influenza virus A/chicken/Jiangsu/1001/2013(H5N2), demonstrating continuous reassortance of H5N2 in China. Genome Announc 1:e00469-13
Gao Z, Hu J, Liang Y, Yang Q, Yan K, Liu D, Wang X, Gu M, Liu X, Hu S, Hu Z, Liu H, Liu W, Chen S, Peng D, Jiao XA, Liu X (2017) Generation and comprehensive analysis of host cell interactome of the PA protein of the highly pathogenic H5N1 avian influenza virus in mammalian cells. Front Microbiol 8:739
Baker SF, Ledwith MP, Mehle A (2018) Differential splicing of ANP32A in birds alters its ability to stimulate RNA synthesis by restricted influenza polymerase. Cell Rep 24(2581–2588):e2584
Reich S, Guilligay D, Pflug A, Malet H, Berger I, Crepin T, Hart D, Lunardi T, Nanao M, Ruigrok RW, Cusack S (2014) Structural insight into cap-snatching and RNA synthesis by influenza polymerase. Nature 516:361–366
Funding
This study was funded by the Faculty Development Grants of Hubei University of Medicine (No. 2016QDJZR03), the Natural Science Foundation of Hubei Province (No. 2018CFB185), the Scientific and Technological Project of Shiyan City (No. 17Y32), and Project of Hubei Provincial Department of Education.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they had no conflict of interest.
Additional information
Handling Editor: Ayato Takada.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, X., Liu, Z., Wang, J. et al. The interaction of cellular protein ANP32A with influenza A virus polymerase component PB2 promotes vRNA synthesis. Arch Virol 164, 787–798 (2019). https://doi.org/10.1007/s00705-018-04139-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-018-04139-z