Abstract
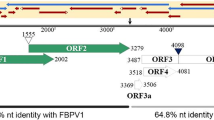
The complete genome of a novel virus, provisionally named areca palm velarivirus 1 (APV1), was identified in areca palm exhibiting leaf yellowing symptoms in Hainan province, China. The genome of APV1 consists of 16,080 nucleotides and possesses 11 open reading frames (ORFs), sharing 56.4 % nucleotide sequence identity with little cherry virus 1 (NC_001836.1). The genome organization of APV1 is highly similar to that of members of the genus Velarivirus (family Closteroviridae). Phylogenetic analysis placed APV1 together with members of the genus Velarivirus.

Similar content being viewed by others
References
Agranovsky AA, Koonin EV, Boyko VP, Maiss E, Frotschl R, Lunina NA, Atabekov JG (1994) Beet yellows closterovirus: complete genome structure and identification of a leader papain-like thiol protease. Virology 198:311–324
Alzhanova DV, Napuli AJ, Creamer R, Dolja VV (2001) Cell-to-cell movement and assembly of a plant closterovirus: roles for the capsid proteins and Hsp70 homolog. EMBO J 20:6997–7007
Alzhanova DV, Prokhnevsky AI, Peremyslov VV, Dolja VV (2007) Virion tails of Beet yellows virus: Coordinated assembly by three structural proteins. Virology 359:220–226
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16:10881–10890
Daquan LMC, Shabin Y (2001) Identification of pathogens of yellow leaf disease of arecanut in Hainan island. Chinese J Trop Crops 22:43–46
Dolja VV, Kreuze JF, Valkonen JPT (2006) Comparative and functional genomics of closteroviruses. Virus Res 117:38–51
Jelkmann W, Fechtner B, Agranovsky AA (1997) Complete genome structure and phylogenetic analysis of little cherry virus, a mealybug-transmissible closterovirus. J Gen Virol 78:2067–2071
Jelkmann W, Mikona C, Turturo C, Navarro B, Rott ME, Menzel W, Saldarelli P, Minafra A, Martelli GP (2012) Molecular characterization and taxonomy of grapevine leafroll-associated virus 7. Arch Virol 157:359–362
Karasev AV (2000) Genetic diversity and evolution of closteroviruses. Annu Rev Phytopathol 38:293–324
Kreuze JF, Perez A, Untiveros M, Quispe D, Fuentes S, Barker I, Simon R (2009) Complete viral genome sequence and discovery of novel viruses by deep sequencing of small RNAs: a generic method for diagnosis, discovery and sequencing of viruses. Virology 388:1–7
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Gen Biol 10:R25
Letunic I, Doerks T, Bork P (2015) SMART: recent updates, new developments and status in 2015. Nucleic Acids Res 43:D257–D260
Napuli AJ, Alzhanova DV, Doneanu CE, Barofsky DF, Koonin EV, Dolja VV (2003) The 64-kilodalton capsid protein homolog of Beet yellows virus is required for assembly of virion tails. J Virol 77:2377–2384
Peremyslov VV, Pan YW, Dolja VV (2004) Movement protein of a closterovirus is a type III integral transmembrane protein localized to the endoplasmic reticulum. J Virol 78:3704–3709
Satyanarayana T, Gowda S, Ayllon MA, Dawson WO (2004) Closterovirus bipolar virion: evidence for initiation of assembly by minor coat protein and its restriction to the genomic RNA 5’ region. Proc Natl Acad Sci USA 101:799–804
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Wu Q, Luo Y, Lu R, Lau N, Lai EC, Li WX, Ding SW (2010) Virus discovery by deep sequencing and assembly of virus-derived small silencing RNAs. Proc Natl Acad Sci USA 107:1606–1611
Acknowledgments
This study was supported by grants from the National Basic Research Program of China (no. 2014CB138405), the Chinese National Natural Science Foundation (no. 31272011), and the Doctoral Fund of the Ministry of Education of China (20123402110013).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, H., Qi, S., Chang, Z. et al. Complete genome sequence of a novel velarivirus infecting areca palm in China. Arch Virol 160, 2367–2370 (2015). https://doi.org/10.1007/s00705-015-2489-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2489-9