Abstract
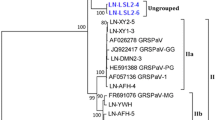
Partial genomic sequences of two divergent grapevine leafroll-associated virus 3 (GLRaV-3) variants, NZ1-B and NZ2, from New Zealand were determined and analysed (11,827 nt and 7,612 nt, respectively). At the nucleotide level, both variants are more than 20 % different from the previously published GLRaV-3 sequences, from phylogenetic groups 1 to 5. Phylogenetic analysis indicated that NZ1-B is a variant of the previously identified divergent NZ-1, while NZ2 is a novel sequence with only 76 % nucleotide sequence identity to GLRaV-3 variants NZ-1, GH11, and GH30. Therefore, NZ2 is a new variant of GLRaV-3. Amino acid sequence analysis of the NZ1-B and NZ2 coat proteins indicated significant substitutions that are predicted to alter the coat protein structure, which potentially leads to the observed reduced immunological reactivity of both variants to the Bioreba anti-GLRaV-3 conjugated monoclonal antibody.


Similar content being viewed by others
References
Baskaran N, Kandpal RP, Bhargava AK, Glynn MW, Bale A, Weissman SM (1996) Uniform amplification of a mixture of deoxyribonucleic acids with varying GC content. Genome Res 6:633–638
Bester R, Maree HJ, Burger JT (2012) Complete nucleotide sequence of a new strain of Grapevine leafroll-associated virus 3 in South Africa. Arch Virol 157:1815–1819
Betts MJ, Russell RB (2003) Amino acid properties and consequences of subsitutions. In: Barnes MR, Gray IC (eds) Bioinformatics for Geneticists. Wiley, New York, pp 289–316
Chooi KM, Pearson MN, Cohen D, Pong JCH (2009) Sequence variation in Grapevine leafroll-associated virus 3 (GLRaV-3) New Zealand isolates. In: Proceedings of 16th Meeting of the International Council for the Study of virus and virus-like diseases of the grapevine, Dijon, France, pp 290–291
Chooi KM, Cohen D, Pearson MN (2012) Generic and sequence-variant specific molecular assays for the detection of the highly variable Grapevine leafroll-associated virus 3. J Virol Methods 189:20–29
Chooi KM, Pearson MN, Cohen D, MacDiarmid RM (2012) Development of generic and variant-specific molecular assays for the detection of the highly variable Grapevine leafroll-associated virus 3. In: Proceedings of 17th Meeting of the International Council for the Study of virus and virus-like diseases of the grapevine, California, USA, pp 142–143
Cohen D, Chooi KM, Bell VA, Blouin AG, Pearson MN, MacDiarmid RM (2012) Detection of new strains of GLRaV-3 in New Zealand using ELISA and RT-PCR. In: Proceedings of 17th Meeting of the International Council for the Study of virus and virus-like diseases of the grapevine, California, USA, pp 118–119
Dolja VV, Kreuze JF, Valkonen JPT (2006) Comparative and functional genomics of closteroviruses. Virus Res 117:38–51
Drummond AJ, Ashton B, Buxton S, Cheung M, Cooper A, Duran C, Field M, Heled J, Kearse M, Markowitz S, Moir R, Stones-Havas S, Sturrock S, Thierer T, Wilson A (2011) Geneious v5.5
Engel EA, Girardi C, Escobar PF, Arredondo V, Domínguez C, Pérez-Acle T, Valenzuela PDT (2008) Genome analysis and detection of a Chilean isolate of Grapevine leafroll associated virus-3. Virus Genes 37:110–118
Garnier J, Osguthorpe DJ, Robson B (1978) Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. J Mol Biol 120:97–120
Gouveia P, Santos MT, Eiras-Dias JE, Nolasco G (2011) Five phylogenetic groups identified in the coat protein gene of Grapevine leafroll-associated virus 3 obtained from Portuguese grapevine varieties. Arch Virol 156:413–420
Gouveia P, Nolasco G (2012) The p19.7 RNA silencing suppressor from Grapevine leafroll-associated virus 3 shows different levels of activity across phylogenetic groups. Virus Genes 45:333–339
Jarugula S, Gowda S, Dawson WO, Naidu RA (2010) 3′-coterminal subgenomic RNAs and putative cis-acting elements of Grapevine leafroll-associated virus 3 reveals ‘unique’ features of gene expression strategy in the genus Ampelovirus. Virol J 7:180
Jooste AEC, Maree HJ, Bellstedt DU, Goszczynski DE, Pietersen G, Burger JT (2010) Three genetic Grapevine leafroll-associated virus 3 variants identified from South African vineyards show high variability in their 5′UTR. Arch Virol 155:1997–2006
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Ling KS, Zhu HY, Gonsalves D (2004) Complete nucleotide sequence and genome organization of Grapevine leafroll-associated virus 3, type member of the genus Ampelovirus. J Gen Virol 85:2099–2102
Maree HJ, Freeborough MJ, Burger JT (2008) Complete nucleotide sequence of a South African isolate of Grapevine leafroll-associated virus 3 reveals a 5′UTR of 737 nucleotides. Arch Virol 153:755–757
Martelli GP, Agranovsky AA, Bar-Joseph M, Boscia D, Candresse T, Coutts RHA, Dolja VV, Falk BW, Gonsalves D, Jelkmann W, Karasev AV, Minafra A, Namba S, Vetten HJ, Wisler GC, Yoshikawa N (2002) The family Closteroviridae revised. Arch Virol 147:2039–2044
Martin DP, Lemey P, Lott M, Moulton V, Posada D, Lefeuvre P (2010) RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics 26:2462–2463
Orecchia M, Nölke G, Saldarelli P, Dell’Orco M, Uhde-Holzem K, Sack M, Martelli G, Fischer R, Schillberg S (2008) Generation and characterization of a recombinant antibody fragment that binds to the coat protein of Grapevine leafroll-associated virus 3. Arch Virol 153:1075–1084
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Seah YM, Sharma AM, Zhang S, Almeida RPP, Duffy S (2012) A divergent variant of Grapevine leafroll-associated virus-3 is present in California. Virol J 9:235
Sharma AM, Wang J, Duffy S, Zhang S, Wong MK, Rashed A, Cooper ML, Daane KM, Almeida RPP (2011) Occurrence of grapevine leafroll-associated virus complex in Napa Valley. PLoS ONE 6:e26227
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Turturo C, Saldarelli P, Yafeng D, Digiaro M, Minafra A, Savino V, Martelli GP (2005) Genetic variability and population structure of Grapevine leafroll-associated virus 3 isolates. J Gen Virol 86:217–224
Wang J, Sharma AM, Duffy S, Almeida RPP (2011) Genetic diversity in the 3′ terminal 4.7-kb region of Grapevine leafroll-associated virus 3. Phytopathology 101:445–450
Zhou Z, Turturo C, Potere O, Saldarelli P, Boscia D, Martelli GP (2003) Production and characterization of monoclonal antibodies specific for GLRaV-3 and epitope mapping of the coat protein. In: Proceedings of 14th Meeting of the International Council for the Study of viruses and virus-like diseases of the grapevine (ICVG), Locorotondo, Italy, p 203
Zhou Z, Turturo C, Potere O, Saldarelli P, Boscia D, Martelli GP (2003) Production and characterization of monoclonal antibodies specific for GLRaV-3 and epitope mapping of the coat protein. J Plant Pathol 85:316
Acknowledgments
This research was supported by Corbans Viticulture Ltd, the University of Auckland, New Zealand Winegrowers, Plant and Food Research, and the Tertiary Education Commission.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chooi, K.M., Cohen, D. & Pearson, M.N. Molecular characterisation of two divergent variants of grapevine leafroll-associated virus 3 in New Zealand. Arch Virol 158, 1597–1602 (2013). https://doi.org/10.1007/s00705-013-1631-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1631-9