Abstract
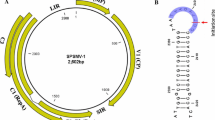
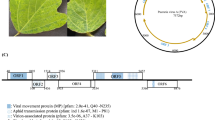
The complete nucleotide sequence of a sweet potato virus, first identified two decades ago as virus “C-6”, was determined in this study. Sequence similarity and phylogenetic analysis clearly place it as a member of a distinct species within the genus Carlavirus, family Betaflexiviridae. Its genome structure was typical for that of other carlaviruses except that the ORF for the cysteine-rich protein was replaced by an ORF encoding a predicted protein with no similarity to any known protein.

Similar content being viewed by others
References
Fuentes S (1994) Preliminary identification of a sweet potato virus (C-6). Fitopatologia 29:38 (Abstract)
Fuentes S, Heider B, Tasso RC, Romero E, Zum Felde T, Kreuze JF (2012) Complete genome sequence of a potyvirus infecting yam beans (Pachyrhizus spp.) in Peru. Arch Virol 157(4):773–776. doi:10.1007/s00705-011-1214-6
Fuentes S, Salazar L (1989) Identification of sweet potato (Ipomoea batatas (L.) Lam.) viruses. Fitoptatologia 24:43 (Abstract)
Gramstat A, Prüfer D, Rohde W (1994) The nucleic acid-binding zinc finger protein of potato virus M is translated by internal initiation as well as by ribosomal frameshifting involving a shifty stop codon and a novel mechanism of P-site slippage. Nucleic Acids Res 22(19):3911–3917. doi:10.1093/nar/22.19.3911
King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (2012) Virus taxonomy: classification and nomenclature of viruses. Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier Academic Press, San Diego
Loebenstein G, Thottappilly G, Fuentes S, Cohen J (2009) Virus and phytoplasma diseases. In: Loebenstein G, Thottappilly G (eds) The Sweetpotato, Part I. Springer Sciences Business Media BV, Dordrecht, pp 105–134. doi:10.1007/978-1-4020-9475-0_8
Martelli GP, Adams MJ, Kreuze JF, Dolja VV (2007) Family Flexiviridae: a case study in virion and genome plasticity. Ann Rev Phytopathol 45:73–100. doi:10.1146/annurev.phyto.45.062806.094401
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
Turner R, Foster GD (1997) Deletion analysis of a translational enhancer upstream from the coat protein open reading frame of potato virus S. Arch Virol 142(1):167–175. doi:10.1007/s007050050067
Untiveros M, Fuentes S, Salazar LF (2007) Synergistic interaction of sweet potato chlorotic stunt virus (Crinivirus) with carla-, cucumo-, ipomo-, and potyviruses infecting sweet potato. Plant Dis 91(6):669–676
Zerbino DR, Birney E (2008) Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18(5):821–829. doi:10.1101/gr.074492.107
Acknowledgements
We gratefully acknowledge the financial contribution from ICGEB-TWAS-UNESCO/IBSP Joint Programme on Capacity Building in Basic Molecular Biology and SLU 2011 funding for international collaboration. This work was also partially funded by the Bill & Melinda Gates Foundation through the Sweet potato Action for Security and Health in Africa project. The findings and conclusions contained within are those of the authors and do not necessarily reflect positions or policies of the Bill & Melinda Gates Foundation. We thank J. Arellano for technical assistance in maintaining the virus isolate in the greenhouse.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
De Souza, J., Fuentes, S., Savenkov, E.I. et al. The complete nucleotide sequence of sweet potato C6 virus: a carlavirus lacking a cysteine-rich protein. Arch Virol 158, 1393–1396 (2013). https://doi.org/10.1007/s00705-013-1614-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1614-x