Abstract
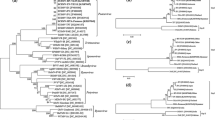
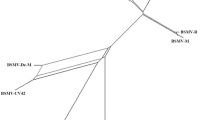
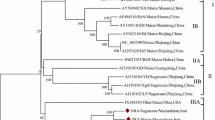
Hop mosaic virus (HpMV), a member of the genus Carlavirus, is importance to hop production worldwide. We identified variation in nucleic and amino acid sequences among 23 HpMV isolates from Australia, the USA, the Czech Republic, South Africa and Japan using a 1,455-bp fragment covering the 3′ end of the virus genome including ORFs 4, 5 and 6. Three clusters of two or more isolates were identified in phylogenies of the total nucleotide sequence and the coat protein (ORF5) amino acid sequence. Two of these clusters combined in analyses of ORF4 and ORF6 amino acid sequences. Isolates from within and outside of Australia were found in each cluster, indicating that sequence variation was not associated with geographic source. Monitoring of HpMV variants in the field and evaluation of the impact of variants on vector association, rate of spread, and hop yield and quality can now be undertaken.

Similar content being viewed by others
References
Crowle DR, Pethybridge SJ, Leggett GW, Sherriff LJ, Wilson CR (2003) Diversity of the coat protein-coding region among Ilarvirus isolates infecting hop in Australia. Plant Pathol 52:655–662
Hataya T, Arimoto R, Suda N, Uyeda I (2001) Molecular characterisation of Hop mosaic virus: its serological and molecular relationships to Hop latent virus. Arch Virol 14:1935–1948
Legg JT (1959) A mild strain of hop mosaic virus. Report of the East Malling Research Station for 1958. East Malling Research Station, Kent, pp 116–167
Mackenzie D, Salmon ES, Ware WM, Williams R (1929) The mosaic disease of the hop. Ann Appl Biol 16:359–381
Martelli GP, Adams MJ, Kreuze JF, Dolja VV (2007) Family Flexiviridae: a case study in virion and genome plasticity. Ann Rev Phytopathol 45:73–100
Munro D (1987) Viruses infecting hop, Humulus lupulus L in Australia. Aust J Agric Res 38:83–90
Pethybridge SJ, Wilson CR, Hay FS, Leggett GW, Sherriff LJ (2002) Effect of viruses on agronomic and brewing characteristics of four hop (Humulus lupulus L.) cultivars in Australia. Ann Appl Biol 140:97–105
Pethybridge SJ, Wilson CR, Leggett GW (2004) Incidence and effect of viruses on production of two newly adopted hop (Humulus lupulus) cultivars in Australia. Aust J Agric Res 55:765–770
Pethybridge SJ, Wilson CR, Sherriff LJ, Leggett GW, Munro D (2000) Virus incidence in Australian hop (Humulus lupulus L.) gardens and cultivar differences in susceptibility to infection. Aust J Agric Res 51:685–689
Pethybridge SJ, Barbara DJ, Eastwell KC, Wilson CR (2008) Virus and viroids infecting hop: significance, epidemiology and management. Plant Dis 92:324–338
Poke FS (2008) Hop mosaic virus: complete nucleotide sequence and relationship to other carlaviruses. Arch Virol 153:1615–1619
Probasco EG, Murphey JM (1996) The effects of hop viruses on brewing and agronomic characteristics in the hop variety Chinook. Master Brewers Assoc Am Tech Quart 33:160–165
Small E (1978) A numerical and nomenclatural analysis of a morphogeographic taxa of Humulus. Syst Bot 3:37–76
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thresh JM, Adams AN (1983) Hop mosaic disease. Report of the East Malling Research Station for1982. East Malling Research Station, Kent, pp 173–175
Acknowledgments
We thank Hop Products Australia Pty Ltd for access to hop gardens for isolate collection. HpMV isolates were kindly provided by Dr. K. Eastwell, E. Jooste and P. Svoboda. We also thank Drs. S. Radisek, U. Skomra, R. Beatson and D. Barbara for providing hop material for testing and Dr. S. Pethybridge and P. Cross for advice on ELISA protocols.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Poke, F.S., Crowle, D.R., Whittock, S.P. et al. Molecular variation of hop mosaic virus isolates. Arch Virol 155, 1721–1724 (2010). https://doi.org/10.1007/s00705-010-0780-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-010-0780-3