Summary.
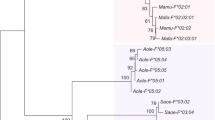
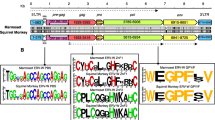
More than 50 copies of HERV-E family have been estimated to exist in the human genome. Here, we examined the expression pattern and their relationships of the HERV-E in Japanese monkey tissues by RT-PCR and sequence analysis. The env gene of HERV-E family was expressed in monkey tissues (testis, prostate, kidney, thymus, intestine and stomach) except for cerebellum, pancreas and ovary, exhibiting that they may have transcriptional potential. Phylogenetic analysis of the HERV-E env family from Japanese monkey tissues and our previous data could be divided into two distinctive groups (I and II). They were integrated into the genomes of anthropoids and have evolved at the rate of 0.3% nucleotide differences per MYr through evolutionary divergence in primate evolution. Divergence times of the two groups were estimated as 11.6 MYr for group I and 41.6 MYr for group II. Those HERV-E sequences were extensively proliferated in the genome of humans and great apes. These data will contribute to further studies on the transcriptional potential of the HERV-E family in the Japanese monkey genome and to biomedical knowledge related to human diseases.
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Yi, JM., Takenaka, O. & Kim, HS. Molecular cloning and phylogeny of HERV-E family that is expressed in Japanese monkey (Macaca fuscata) tissues. Arch Virol 150, 869–882 (2005). https://doi.org/10.1007/s00705-004-0480-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-004-0480-y