Abstract
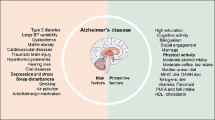
Alzheimer’s Disease (AD) is a neurodegenerative disorder characterized by the impairment of cognitive function and loss of memory. Previous studies indicate an essential role of immune response in AD, but the detailed mechanisms remain unclear. In this study, we obtained 1664 credible risk variants (CRVs) based on the most significant SNP detected by International Genomics of Alzheimer’s Project, from which 99 genes (CRVs-related genes) were identified. Function analysis revealed that these genes were mainly involved in immune response and amyloid-β and its precursor metabolisms, indicating a potential role of immune response in regulating neurobiological processes in the etiology of neurodegenerative disease. Pathway crosstalk analysis revealed the complicated connections between immune-related pathways. Further, we found that the CRVs-related genes showed temporal-specific expression in the thalamus in adolescence developmental period. Cell type-specific expression analysis found that CRVs-related genes might be specifically expressed in brain cells such as astrocytes and oligodendrocytes. Protein–protein interaction network analysis identified the highly interconnected 'hub' genes, all of which were susceptible loci of AD. These results indicated that the CRVs may exert a potential influence in AD by regulating immune response, thalamus development, astrocytes activities, and amyloid-β binding. Our results provided hints for further experimental verification of AD pathophysiology.






Similar content being viewed by others
Data availability
The data used in the current study are available from the corresponding author on reasonable request.
References
Ager RR, Davis JL, Agazaryan A, Benavente F, Poon WW, LaFerla FM, Blurton-Jones M (2015) Human neural stem cells improve cognition and promote synaptic growth in two complementary transgenic models of Alzheimer’s disease and neuronal loss. Hippocampus 25:813–826. https://doi.org/10.1002/hipo.22405
Aggleton JP, Pralus A, Nelson AJ, Hornberger M (2016) Thalamic pathology and memory loss in early Alzheimer’s disease: moving the focus from the medial temporal lobe to Papez circuit. Brain 139:1877–1890. https://doi.org/10.1093/brain/aww083
Agostinho P, Cunha RA, Oliveira C (2010) Neuroinflammation, oxidative stress and the pathogenesis of Alzheimer’s disease. Curr Pharm Des 16:2766–2778. https://doi.org/10.2174/138161210793176572
Aikawa T et al (2019) ABCA7 haplodeficiency disturbs microglial immune responses in the mouse brain. Proc Natl Acad Sci USA 116:23790–23796. https://doi.org/10.1073/pnas.1908529116
Akiyama H et al (2000) Inflammation and Alzheimer’s disease. Neurobiol Aging 21:383–421. https://doi.org/10.1016/s0197-4580(00)00124-x
Alcolea D et al (2015) Amyloid precursor protein metabolism and inflammation markers in preclinical Alzheimer disease. Neurology 85:626–633. https://doi.org/10.1212/wnl.0000000000001859
Bateman RJ et al (2012) Clinical and biomarker changes in dominantly inherited Alzheimer’s disease. N Engl J Med 367:795–804. https://doi.org/10.1056/NEJMoa1202753
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Royal Stat Soc Series B 57:289–300
Blasko I, Marx F, Steiner E, Hartmann T, Grubeck-Loebenstein B (1999) TNFalpha plus IFNgamma induce the production of Alzheimer beta-amyloid peptides and decrease the secretion of APPs FASEB journal : official publication of the Federation of American Societies for. Exp Biol 13:63–68. https://doi.org/10.1096/fasebj.13.1.63
Blennow K, de Leon MJ, Zetterberg H (2006) Alzheimer’s disease. Lancet 368:387–403. https://doi.org/10.1016/s0140-6736(06)69113-7
Bolos M, Perea JR, Avila J (2017) Alzheimer’s disease as an inflammatory disease Biomolecular concepts 8:37–43. https://doi.org/10.1515/bmc-2016-0029
Bostanciklioglu M (2019) An update on the interactions between Alzheimer’s disease, autophagy and inflammation. Gene 705:157–166. https://doi.org/10.1016/j.gene.2019.04.040
Boyle AP et al (2012) Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 22:1790–1797. https://doi.org/10.1101/gr.137323.112
Cai Z, Wan CQ, Liu Z (2017) Astrocyte and Alzheimer’s disease. J Neurol 264:2068–2074. https://doi.org/10.1007/s00415-017-8593-x
Carter CJ (2010) Alzheimer’s disease: a pathogenetic autoimmune disorder caused by herpes simplex in a gene-dependent manner. Int J Alzheimer’s Dis. https://doi.org/10.4061/2010/140539
Chan SL, Kim WS, Kwok JB, Hill AF, Cappai R, Rye KA, Garner B (2008) ATP-binding cassette transporter A7 regulates processing of amyloid precursor protein in vitro. J Neurochem 106:793–804. https://doi.org/10.1111/j.1471-4159.2008.05433.x
Chen J, Bardes EE, Aronow BJ, Jegga AG (2009) ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res 37:W305-311. https://doi.org/10.1093/nar/gkp427
Chen CH, Lin CL, Kao CH (2016) Irritable bowel syndrome is associated with an increased risk of dementia: a nationwide population-based study. PLoS ONE 11:e0144589. https://doi.org/10.1371/journal.pone.0144589
De Roeck A et al (2017) Deleterious ABCA7 mutations and transcript rescue mechanisms in early onset Alzheimer’s disease. Acta Neuropathol 134:475–487. https://doi.org/10.1007/s00401-017-1714-x
Du J, Li M, Yuan Z, Guo M, Song J, Xie X, Chen Y (2016) A decision analysis model for KEGG pathway analysis. BMC Bioinformatics 17:1–12. https://doi.org/10.1186/s12859-016-1285-1
Dunn AR, O’Connell KMS, Kaczorowski CC (2019) Gene-by-environment interactions in Alzheimer’s disease and Parkinson’s disease. Neurosci Biobehav Rev 103:73–80. https://doi.org/10.1016/j.neubiorev.2019.06.018
Field HJ, Vere Hodge RA (2013) Recent developments in anti-herpesvirus drugs. Br Med Bull 106:213–249. https://doi.org/10.1093/bmb/ldt011
Grossberg GT et al (2013) The safety, tolerability, and efficacy of once-daily memantine (28 mg): a multinational, randomized, double-blind, placebo-controlled trial in patients with moderate-to-severe Alzheimer’s disease taking cholinesterase inhibitors. CNS Drugs 27:469–478. https://doi.org/10.1007/s40263-013-0077-7
Harris SA, Harris EA (2018) Molecular mechanisms for herpes simplex virus type 1 pathogenesis in Alzheimer’s Disease. Front Aging Neurosci 10:48. https://doi.org/10.3389/fnagi.2018.00048
Heneka MT et al (2015) Neuroinflammation in Alzheimer’s disease Lancet Neurol 14:388–405
Heppner FL, Ransohoff RM, Becher B (2015) Immune attack: the role of inflammation in Alzheimer disease Nature reviews. Neuroscience 16:358–372. https://doi.org/10.1038/nrn3880
Howard R et al (2012) Donepezil and memantine for moderate-to-severe Alzheimer’s disease. N Engl J Med 366:893–903. https://doi.org/10.1056/NEJMoa1106668
Hu Y et al (2017a) Rs4878104 contributes to Alzheimer’s disease risk and regulates DAPK1 gene expression. Neurol Sci 38:1255–1262. https://doi.org/10.1007/s10072-017-2959-9
Itzhaki RF, Dobson CB, Wozniak MA (2004) Herpes simplex virus type 1 and Alzheimer's disease. Ann Neurol 55:299–300; author reply 300–291 doi:https://doi.org/10.1002/ana.10852
Hu Y et al (2017b) GAB2 rs2373115 variant contributes to Alzheimer’s disease risk specifically in European population. J Neurol Sci 375:18–22. https://doi.org/10.1016/j.jns.2017.01.030
Jha HC et al (2015) Gammaherpesvirus Infection of Human Neuronal Cells. MBio 6:e01844-e1815. https://doi.org/10.1128/mBio.01844-15
Jia P, Kao C-F, Kuo P-H, Zhao Z (2011) A comprehensive network and pathway analysis of candidate genes in major depressive disorder. BMC Syst Biol 5:S12
Jonsson T et al (2013) Variant of TREM2 associated with the risk of Alzheimer’s disease. N Engl J Med 368:107–116. https://doi.org/10.1056/NEJMoa1211103
Jun H et al (2018) An immune-beige adipocyte communication via nicotinic acetylcholine receptor signaling. Nat Med 24:814–822. https://doi.org/10.1038/s41591-018-0032-8
Kang HJ et al (2011) Spatio-temporal transcriptome of the human brain. Nature 478:483–489. https://doi.org/10.1038/nature10523
Karch CM, Goate AM (2015) Alzheimer’s disease risk genes and mechanisms of disease pathogenesis. Biol Psychiat 77:43–51. https://doi.org/10.1016/j.biopsych.2014.05.006
Khera R, Das N (2009) Complement Receptor 1: disease associations and therapeutic implications. Mol Immunol 46:761–772. https://doi.org/10.1016/j.molimm.2008.09.026
Kim JH (2018) Genetics of Alzheimer’s Disease. Dementia Neurocogn Disorders 17:131–136. https://doi.org/10.12779/dnd.2018.17.4.131
Koekkoek PS, Rutten GE, Biessels GJ (2014) Cognitive disorders in diabetic patients. Handbook Clin Neurol 126:145–166. https://doi.org/10.1016/b978-0-444-53480-4.00011-4
Kohl M, Wiese S, Warscheid B (2011) Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 696:291–303. https://doi.org/10.1007/978-1-60761-987-1_18
Kremer AN et al (2014) Development of a coordinated allo T cell and auto B cell response against autosomal PTK2B after allogeneic hematopoietic stem cell transplantation. Haematologica 99:365–369. https://doi.org/10.3324/haematol.2013.086652
Lambert JC et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45:1452–1458. https://doi.org/10.1038/ng.2802
Li MX, Sham PC, Cherny SS, Song YQ (2010) A knowledge-based weighting framework to boost the power of genome-wide association studies. PLoS ONE 5:e14480. https://doi.org/10.1371/journal.pone.0014480
Li MX, Gui HS, Kwan JS, Sham PC (2011) GATES: a rapid and powerful gene-based association test using extended Simes procedure. Am J Hum Genet 88:283–293. https://doi.org/10.1016/j.ajhg.2011.01.019
Li Puma DD et al (2019) Herpes simplex virus type-1 infection impairs adult hippocampal neurogenesis via amyloid-beta protein accumulation. Stem cells. https://doi.org/10.1002/stem.3072
Litvinchuk A et al (2018) Complement C3aR inactivation attenuates tau pathology and reverses an immune network deregulated in tauopathy models and Alzheimer’s Disease. Neuron 100:1337-1353.e1335. https://doi.org/10.1016/j.neuron.2018.10.031
Liu M, Fan R, Liu X, Cheng F, Wang J (2015) Pathways and Networks-Based Analysis of Candidate Genes Associated with Nicotine Addiction. PLoS ONE 10:e0127438
Liu C, Liu Y, Liang L, Cui S, Zhang Y (2019) RNA-Seq based transcriptome analysis during bovine viral diarrhoea virus (BVDV) infection. BMC Genom 20:774. https://doi.org/10.1186/s12864-019-6120-4
Lozano R et al (2012) Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380:2095–2128. https://doi.org/10.1016/S0140-6736(12)61728-0
Lu Y, Quan C, Chen H, Bo X, Zhang C (2017) 3DSNP: a database for linking human noncoding SNPs to their three-dimensional interacting genes. Nucleic Acids Res 45:D643-d649. https://doi.org/10.1093/nar/gkw1022
Majewski J, Schwartzentruber J, Lalonde E, Montpetit A, Jabado N (2011) What can exome sequencing do for you? J Med Genet 48:580–589. https://doi.org/10.1136/jmedgenet-2011-100223
Mangold CA, Szpara ML (2019) Persistent Infection with Herpes Simplex Virus 1 and Alzheimer’s Disease-A Call to Study How Variability in Both Virus and Host may Impact Disease. Viruses. https://doi.org/10.3390/v11100966
Marciani DJ (2015) Alzheimer’s disease vaccine development: a new strategy focusing on immune modulation. J Neuroimmunol 287:54–63. https://doi.org/10.1016/j.jneuroim.2015.08.008
Masters CL, Bateman R, Blennow K, Rowe CC, Sperling RA, Cummings JL (2015) Alzheimer’s disease Nature reviews. Dis Prim 1:15056. https://doi.org/10.1038/nrdp.2015.56
McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM (1984) Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 34:939–944. https://doi.org/10.1212/wnl.34.7.939
McKhann GM et al (2011) The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7:263–269. https://doi.org/10.1016/j.jalz.2011.03.005
Michailidou K et al (2017) Association analysis identifies 65 new breast cancer risk loci. Nature 551:92–94. https://doi.org/10.1038/nature24284
Mrak RE, Griffin WS (2001) Interleukin-1, neuroinflammation, and Alzheimer’s disease. Neurobiol Aging 22:903–908. https://doi.org/10.1016/s0197-4580(01)00287-1
Naya FJ, Wu C, Richardson JA, Overbeek P, Olson EN (1999) Transcriptional activity of MEF2 during mouse embryogenesis monitored with a MEF2-dependent transgene. Development 126:2045–2052
Nicolas G et al (2016) SORL1 rare variants: a major risk factor for familial early-onset Alzheimer’s disease. Mol Psychiatry 21:831–836. https://doi.org/10.1038/mp.2015.121
Ouwens DM et al (2014) Cerebrospinal fluid levels of Alzheimer’s disease biomarkers in middle-aged patients with type 1 diabetes. Diabetologia 57:2208–2214. https://doi.org/10.1007/s00125-014-3333-6
Peter-Derex L, Yammine P, Bastuji H, Croisile B (2015) Sleep and Alzheimer’s disease. Sleep Med Rev 19:29–38. https://doi.org/10.1016/j.smrv.2014.03.007
Rabbani B, Tekin M, Mahdieh N (2014) The promise of whole-exome sequencing in medical genetics. J Hum Genet 59:5–15. https://doi.org/10.1038/jhg.2013.114
Rochowiak A, Niemir ZI (2010) [The structure and role of CR1 complement receptor in physiology] Polski merkuriusz lekarski : organ Polskiego. Towarzystwa Lekarskiego 28:79–83
Rosenthal SL, Kamboh MI (2014) Late-onset Alzheimer’s disease genes and the potentially implicated pathways. Curr Genet Med Rep 2:85–101. https://doi.org/10.1007/s40142-014-0034-x
Ryan SM, Nolan YM (2016) Neuroinflammation negatively affects adult hippocampal neurogenesis and cognition: can exercise compensate? Neurosci Biobehav Rev 61:121–131. https://doi.org/10.1016/j.neubiorev.2015.12.004
Samieri C et al (2018) Association of Cardiovascular Health Level in Older Age With Cognitive Decline and Incident Dementia. JAMA 320:657–664. https://doi.org/10.1001/jama.2018.11499
Shim SM, Cheon HS, Jo C, Koh YH, Song J, Jeon JP (2017) Elevated Epstein-Barr virus antibody level is associated with cognitive decline in the korean elderly. J Alzheimer’s Dis 55:293–301. https://doi.org/10.3233/jad-160563
Solomon A et al (2014) Advances in the prevention of Alzheimer’s disease and dementia. J Intern Med 275:229–250. https://doi.org/10.1111/joim.12178
Spielman LJ, Gibson DL, Klegeris A (2018) Unhealthy gut, unhealthy brain: the role of the intestinal microbiota in neurodegenerative diseases. Neurochemistry Int 120:149–163. https://doi.org/10.1016/j.neuint.2018.08.005
Steinberg S et al (2015) Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat Genet 47:445–447. https://doi.org/10.1038/ng.3246
Sullivan PF, Daly MJ, O’Donovan M (2012) Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat Rev Genet 13:537–551. https://doi.org/10.1038/nrg3240
Swarup V et al (2019) Identification of evolutionarily conserved gene networks mediating neurodegenerative dementia. Nat Med 25:152–164. https://doi.org/10.1038/s41591-018-0223-3
The Alzheimer’s Association (2020) 2020 Alzheimer’s disease facts and figures. Alzheimer’s Dement 16:391–460. https://doi.org/10.1002/alz.12068
Trowsdale J, Knight JC (2013) Major histocompatibility complex genomics and human disease. Annu Rev Genomics Hum Genet 14:301–323. https://doi.org/10.1146/annurev-genom-091212-153455
van der Flier WM, Scheltens P (2005) Epidemiology and risk factors of dementia. J Neurol Neurosurg Psychiatry 76(Suppl 5):v2–7 doi:https://doi.org/10.1136/jnnp.2005.082867
Wang J, Duncan D, Shi Z (2013) Zhang B (2013) WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update. Nucleic Acids Res 41:W77-83. https://doi.org/10.1093/nar/gkt439
Wang J et al (2017) BIN1 reverses PD-L1-mediated immune escape by inactivating the c-MYC and EGFR/MAPK signaling pathways in non-small cell lung cancer. Oncogene 36:6235–6243. https://doi.org/10.1038/onc.2017.217
Wozniak MA, Itzhaki RF, Shipley SJ, Dobson CB (2007) Herpes simplex virus infection causes cellular beta-amyloid accumulation and secretase upregulation. Neurosci Lett 429:95–100. https://doi.org/10.1016/j.neulet.2007.09.077
Xu X, Wells AB, O’Brien DR, Nehorai A, Dougherty JD (2014) Cell type-specific expression analysis to identify putative cellular mechanisms for neurogenetic disorders The Journal of neuroscience : the official journal of the Society for. Neuroscience 34:1420–1431. https://doi.org/10.1523/jneurosci.4488-13.2014
Yao PL et al (2017) Androgen alleviates neurotoxicity of β-amyloid peptide (Aβ) by promoting microglial clearance of Aβ and inhibiting microglial inflammatory response to Aβ. CNS Neurosci Therap 23:855–865. https://doi.org/10.1111/cns.12757
Yegambaram M, Manivannan B, Beach TG, Halden RU (2015) Role of environmental contaminants in the etiology of Alzheimer’s disease: a review. Curr Alzheimer Res 12:116–146
Zhang B, Kirov S, Snoddy J (2005) WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res 33:W741-748. https://doi.org/10.1093/nar/gki475
Zhang B et al (2013) Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153:707–720. https://doi.org/10.1016/j.cell.2013.03.030
Zhou X et al (2018) Identification of genetic risk factors in the Chinese population implicates a role of immune system in Alzheimer’s disease pathogenesis. Proc Natl Acad Sci USA 115:1697–1706. https://doi.org/10.1073/pnas.1715554115
Acknowledgements
We are grateful to the IGAP consortium for providing summary statistics for the index SNPs.
Funding
This study was supported in part by grants from National Key Research and Development Program of China (No. 2016YFC0906300), National Natural Science Foundation of China (No. 31271411 and 91746205).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guo, P., Cao, C., Ma, Y. et al. Network-based analysis on genetic variants reveals the immunological mechanism underlying Alzheimer’s disease. J Neural Transm 128, 803–816 (2021). https://doi.org/10.1007/s00702-021-02337-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-021-02337-9