Abstract
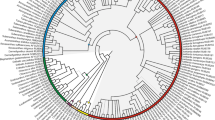
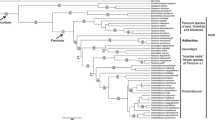
A molecular systematic study of Globba section Nudae (Zingiberaceae) using ITS and matK sequences identifies three major clades, Globba subsection Nudae, G. subsection Mediocalcaratae and a new subsection, Globba subsection Pelecantherae, which is described here. The two species belonging in this subsection, Globba pelecanthera and Globba securifer, which are both new, are described. Rectangular anther appendages are reported in Globba for the first time. Evidence of hybridisation is given. The morphological characters of the flowers, which are likely to be important in pollination, are discussed.









Similar content being viewed by others
References
Beaulieu JM, O’Meara BC, Donoghue MJ (2013) Identifying hidden rate changes in the evolution of a binary morphological character: the evolution of plant habit in campanulid angiosperms. Syst Biol 62:725–737. https://doi.org/10.1093/sysbio/syt034
De Boer H, Newman MF, Poulsen AD, Droop AJ, Fer T, Hiền LTT, Hlavatá K, Lamxay V, Richardson JE, Steffen K, Leong-Škorničková J (2018) Convergent morphology in Alpinieae (Zingiberaceae): Recircumscribing Amomum as a monophyletic genus. Taxon 67:6–36. https://doi.org/10.12705/671.2
Borowiec ML (2016) AMAS: a fast tool for alignment manipulation and computing of summary statistics. PeerJ 4:e1660. https://doi.org/10.7717/peerj.1660
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
Fér T, Schmickl R (2018) HybPhyloMaker: target enrichment data analysis from raw reads to species trees. Evol Bioinf 14:1–9. https://doi.org/10.1177/1176934317742613
Foster PG (2004) Modelling compositional heterogeneity. Syst Biol 53:485–495. https://doi.org/10.1080/10635150490445779
Guindon S, Gascuel O (2003) A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hijmans RJ, Guarino L, Cruz M, Rojas E (2001) Computer tools for spatial analysis of plant genetic resources data: 1. DIVA-GIS. Pl Genet Res Newslett 127:15–19
Horaninow P (1862) Prodromus Monographiae Scitaminearum. Academiae Caesareae Scientiarum, St. Petersburg
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molec Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
IUCN Standards and Petitions Committee (2019) Guidelines for Using the IUCN Red List Categories and Criteria. Version 14. IUCN, Cambridge. Available at: www.iucnredlist.org/documents/RedList Guidelines.pdf. Accessed June 2021
Junier T, Zdobnov EM (2010) The Newick utilities: high-throughput phylogenetic tree processing in the UNIX shell. Bioinformatics 26:1669–1670. https://doi.org/10.1093/bioinformatics/btq243
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molec Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: A fast, scalable, and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455. https://doi.org/10.1093/bioinformatics/btz305
Kress WJ, Prince LM, Williams KJ (2002) The phylogeny and a new classification of the gingers (Zingiberaceae): evidence from molecular data. Amer J Bot 89:1682–1696. https://doi.org/10.3732/ajb.89.10.1682
Larsen K (1972) Studies in the genus Globba in Thailand. Notes Roy Bot Gard Edinburgh 31:229–241
Mabberley DJ (2017) Mabberley’s Plant Book, 4th edn. Cambridge University Press, Cambridge. https://doi.org/10.1017/9781316335581
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A (2010) How many bootstrap replicates are necessary? J Computat Biol 17:337–354. https://doi.org/10.1089/cmb.2009.0179
Price MN, Dehal PS, Arkin AP (2010) FastTree 2 – Approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
QGIS.org (2021) QGIS Geographic Information System. QGIS Association. Available at: http://www.qgis.org
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. Available at: https://www.R-project.org/.
Rambaut A (2014) FigTree v1.4.2. A graphical viewer of phylogenetic trees. Available at: http://tree.bio.ed.ac.uk/software/figtree/
Revell LJ (2012) phytools: An R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 3:217–223. https://doi.org/10.1111/j.2041-210X.2011.00169.x
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Sakai S, Kato M, Inoue T (1999) Three pollination guilds and variation in floral characteristics of Bornean gingers (Zingiberaceae and Costaceae). Amer J Bot 86:646–658. https://doi.org/10.2307/2656573
Sangvirotjanapat S, Denduangboriphant J, Newman MF (2019a) A taxonomic revision of Globba subsect. Nudae (Zingiberaceae). Eur J Taxon 503:1–37. https://doi.org/10.5852/ejt.2019.503
Sangvirotjanapat S, Denduangboriphant J, Newman MF (2019b) A taxonomic revision of Globba sect. Nudae subsect. Mediocalcaratae (Zingiberaceae). Edinburgh J Bot 77:1–63. https://doi.org/10.1017/S0960428619000258
Sangvirotjanapat S, Newman MF (2021) Eight new species of Globba (Zingiberaceae) from Thailand and Lao PDR. Phytotaxa 505:139–156. https://doi.org/10.11646/phytotaxa.505.2.2
Sangvirotjanapat S, Trần HĐ, Newman MF (2020) Ten new species of Globba section Globba from continental South-East Asia. Thai Forest Bull 48:212–233. https://doi.org/10.20531/tfb.2020.48.2.15
Schumann KM (1904) Zingiberaceae. In: Engler HGA (ed), Das Pflanzenreich IV. 46 (Heft 20). Engelmann, Berlin, pp 132–161
Secretariat of the Convention on Biological Diversity (2020) Global Biodiversity Outlook 5. ICAO, Montreal
Smitinand T (1958) The genus Dipterocarps Gaertn. f. in Thailand. Thai Forest Bull, Bot 4:1–50
Specht CD, Yockteng R, Almeida AM, Kirchoff BK, Kress WJ (2012) Homoplasy, pollination, and emerging complexity during the evolution of floral development in the tropical gingers (Zingiberales). Bot Rev 78:440–462. https://doi.org/10.1007/s12229-012-9111-6
Steele KP, Vigalys R (1994) Phylogenetic analyses of Polemoniaceae using nucleotide sequences of the plastid gene matK. Syst Bot 19:126–142. https://doi.org/10.2307/2419717
Takano A, Okada H (2002) Multiple occurrences of triploid formation in Globba (Zingiberaceae) from molecular evidence. Pl Syst Evol 230: 143–159. http://www.jstor.org/stable/23644198
Van Welzen P, Madern A, Raes N, Parnell JAN, Simpson DA, Byrne C, Curtis T, Macklin J, Trias-Blasi A, Prajaksood A, Bygrave P, Dransfield S, Kirkup DW, Moat J, Wilkin P, Couch C, Boyce PC, Chayamarit K, Chantaranothai P, Esser H-J, Jebb MHP, Larsen K, Larsen SS, Nielsen I, Meade C, Middleton DJ, Pendry CA, Muasya AM, Pattharahirantricin N, Pooma R, Suddee S, Staples GW, Sungkaew S, Teerawatananon A (2011) The current and future status of floristic provinces in Thailand. In: Trisurat Y, Shreshta RP, Alkemade R (eds) Land use, climate change and biodiversity modelling: perspectives and applications. Information Science Reference, Hershey, pp 219–247
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA Genes for Phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White T (eds) PCR Protocols. Academic Press, San Diego, pp 315–322
Williams KJ (2004) The phylogeny, evolution and classification of the genus Globba and tribe Globbeae (Zingiberaceae): appendages do matter. Amer J Bot 91:100–114. https://doi.org/10.3732/ajb.91.1.100
Záveská E, Fér T, Šída O, Krak K, Marhold K, Leong-Škorničková J (2012) Phylogeny of Curcuma (Zingiberaceae) based on plastid and nuclear sequences: proposal of the new subgenus Ecomata. Taxon 61:747–763. https://doi.org/10.1002/tax.614004
Záveská E, Fér T, Šída O, Marhold K, Leong-Škorničková J (2016) Hybridization among distantly related species: Examples from the polyploid genus Curcuma (Zingiberaceae). Molec Phylogen Evol 100:303–321. https://doi.org/10.1016/j.ympev.2016.04.017
Acknowledgements
This work was carried out as part of the first author’s PhD thesis at Chulalongkorn University. We are grateful to the staff of the Office of the Forest Herbarium (BKF), Department of National Parks, Wildlife and Plant Conservation for permission to collect in national parks and for accompanying us in the field, and to Dr Oudomphone Insisiengmay (The Cabinet of Lao Academy of Science and Technology (CAST), Ministry of Education and Sport, Lao PDR) and Mrs Peou Youleang (RUPP) for the same cooperation in Lao PDR and Cambodia.
Funding
This work is supported financially by the 100th Anniversary Chulalongkorn University Fund for Doctoral Scholarship and the 90th Anniversary Chulalongkorn Fund (Ratchadaphiseksomphot Endowment Fund). The Royal Botanic Garden Edinburgh (RBGE) is supported by the Scottish Government’s Rural and Environmental Science and Analytical Services Division. Computational resources were supplied by the project "e-Infrastruktura CZ" (e-INFRA LM2018140) provided within the programme Projects of Large Research, Development and Innovations Infrastructures. Field work in Cambodia and Laos was supported financially by The Sud Expert Plantes Developpement Durable programme (SEP2D), project AAP3-96 and Institut de Systématique Évolution Biodiversité (ISYEB), Muséum national d’histoire naturelle, Paris, France.
Author information
Authors and Affiliations
Contributions
SS collected the data and carried out the morphological analysis. MN and SS wrote the manuscript. TF and SS carried out analysis of molecular data and wrote relevant parts of the text. JD supervised the PhD thesis of SS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Availability of data and material (data transparency)
All sequence data are available in GenBank, morphological data are given in Table 2.
Additional information
Handling editor: Livia Wanntorp.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sangvirotjanapat, S., Fér, T., Denduangboripant, J. et al. Phylogeny of Globba section Nudae and taxonomic revision of the new Globba subsection Pelecantherae. Plant Syst Evol 308, 5 (2022). https://doi.org/10.1007/s00606-021-01789-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-021-01789-6