Abstract
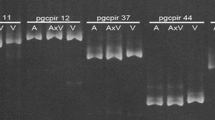
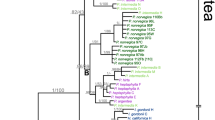
The hybrid origin of Miscanthus purpurascens has previously been proposed, primarily because of its intermediate morphology. In this study, phylogenies based on the DNA sequences from the internal transcribed spacer region of nuclear ribosomal DNA (nrDNA ITS), on the DNA sequences of the trnL intron and trnL-F intergenic spacer of chloroplast DNA, and on amplified fragment length polymorphism (AFLP) fingerprinting confirm that M. purpurascens originated through homoploid hybridization between M. sinensis and M. sacchariflorus. Two different types of ITS sequences were identified from almost all plants of M. purpurascens. One type was found to be closely related to M. sinensis and the other to M. sacchariflorus. Miscanthus purpurascens was found to possess many M. sinensis- and M. sacchariflorus-specific AFLP bands but no band specific to itself. Clustering with the Unweighted Pair Group Method with Arithmetic Mean and principal coordinate analysis based on the AFLP data also demonstrated that M. purpurascens is an approximate intermediate of the two species. In addition, M. purpurascens has the plastid genome of M. sinensis or M. sacchariflorus, suggesting that either species could be its maternal parent. All specimens of M. purpurascens and its coexisting parental species are identified as diploids (2n = 2x = 38). Possible mechanisms of natural hybridization, hybrid status, chloroplast DNA recombination, and evolutionary implications of this hybridization are also discussed.






Similar content being viewed by others
References
Abbott RJ, Hegarty MJ, Hiscock SJ, Brennan AC (2010) Homoploid hybrid speciation in action. Taxon 59:1375–1386
Akaike H (1974) A new look at the statistical model identification. IEEE T Automat Contr 19:716–723
Arnold ML (1997) Natural hybridization and evolution. Oxford University Press, New York
Arnold ML (2004) Transfer and origin of adaptations through natural hybridization: were Anderson and Stebbins right? Plant Cell 16:562–570
Bullard MJ, Nixon PMI, Heath MC (1997) Quantifying the yield of Miscanthus x giganteus in the UK. Asp Appl Biol 49:199–206
Burzyński A, Zbawicka M, Skibinski DO, Wenne R (2003) Evidence for recombination of mtDNA in the marine mussel Mytilus trossulus from the Baltic. Mol Biol Evol 20:388–392
Chen SL, Renvoize SA (2006) MISCANTHUS Andersson, Öfvers. Kongl. Vetensk.-Akad. Förh. 12:165. 1855. Flora of China, 2nd edn, vol 22, pp 581–583
Ciborowski KL, Consuegra S, de Leániz CG, Beaumont MA, Wang J, Jordan WC (2007) Rare and fleeting: an example of interspecific recombination in animal mitochondrial DNA. Biol Lett 3:554–557
Collura S, Azambre B, Finqueneisel G, Zimny T, Weber JV (2006) Miscanthus x giganteus straw and pellets as sustainable fuels: combination and emission tests. Environ Chem Lett 4:75–78
Corriveau JL, Coleman AW (1988) Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 angiosperm species. Am J Bot 75:1443–1458
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ferguson D, Sang T (2001) Speciation through homoploid hybridization between allotetraploids in peonies (Paeonia). Proc Natl Acad Sci USA 98:3915–3919
Franzke A, Mummenhoff K (1999) Recent hybrid speciation in Cardamine (Brassicaceae)—conversion of nuclear ribosomal ITS sequences in statu nascendi. Theor Appl Genet 98:831–834
Gammon MA, Grimsby JL, Tsirelson D, Kesseli R (2007) Molecular and morphological evidence reveals introgression in swarms of the invasive taxa Fallopia japonica, F. sachalinensis, and F. x bohemica (Polygonaceae) in the United States. Am J Bot 94:948–956
Gantenbein B, Fet V, Gantenbein-Ritter IA, Balloux F (2005) Evidence for recombination in scorpion mitochondrial DNA (Scorpiones: Buthidae). Proc R Soc B 272:697–704
Garnhart NJ (2001) Binthere v1.0, A program to bin AFLP data. University of New Hampshire
Greef JM, Deuter M (1993) Syntaxonomy of Miscanthus x giganteus Greef et Deu. Angew Bot 67:87–90
Gross BL, Schwarzbach AE, Rieseberg LH (2003) Origin (s) of the diploid hybrid species Helianthus deserticola (Asteraceae). Am J Bot 90:1708–1719
Heaton EA, Long SP, Voigt TB, Jones M, Clifton-Brown J (2004) Miscanthus for renewable energy generation: European Union experience and projections for Illinois. Mitig Adapt Strat Glob Change 9:433–451
Heaton EA, Dohleman FG, Long SP (2008) Meeting US biofuel goals with less land: the potential of Miscanthus. Global Change Biol 14:2000–2014
Hillis DM, Moritz C, Porter CA, Baker RJ (1991) Evidence for biased gene conversion in concerted evolution of ribosomal DNA. Science 251:308–310
Hoarau G, Holla S, Lescasse R, Stam WT, Olsen JL (2002) Heteroplasmy and evidence for recombination in the mitochondrial control region of the flatfish Platichthys flesus. Mol Biol Evol 19:2261–2264
Hodkinson TR, Renvoize SA (2001) Nomenclature of Miscanthus x giganteus (Poaceae). Kew Bull 56:759–760
Hodkinson TR, Chase MW, Renvoize SA (2002a) Characterization of a genetic resource collection for Miscanthus (Saccharinae, Andropogoneae, Poaceae) using AFLP and ISSR PCR. Ann Bot 89:627–636
Hodkinson TR, Chase MW, Takahashi C, Leitch IJ, Bennett MD, Renvoize SA (2002b) The use of DNA sequencing (ITS and trnL-F), AFLP, and fluorescent in situ hybridization to study allopolyploid Miscanthus (Poaceae). Am J Bot 89:279–286
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hughes KW, Petersen RH (2001) Apparent recombination or gene conversion in the ribosomal ITS region of a Flammulina (Fungi, Agaricales) hybrid. Mol Biol Evol 18:94–96
Jaramillo-Correa JP, Bousquet J (2005) Mitochondrial genome recombination in the zone of contact between two hybridizing conifers. Genetics 171:1951–1962
Jordan WC, Courtney MW, Neigel JE (1996) Low levels of intraspecific genetic variation at a rapidly evolving chloroplast DNA locus in North American duckweeds (Lemnaceae). Am J Bot 83:430–439
Jug T, Dovc P, Pohar J, Snoj A (2004) RAPD analysis as a tool for discriminating marble trout from hybrids (marble trout × brown trout) in the zones of hybridization. J Anim Breed Genet 121:156–162
Kameyama Y, Toyama M, Ohara M (2005) Hybrid origins and F1 dominance in the free-floating, sterile bladderwort, Utricularia australis f. australis (Lentibulariaceae). Am J Bot 92:469–476
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kraytsberg Y, Schwartz M, Brown TA, Ebralidse K, Kunz WS, Clayton DA, Vissing J, Khrapko K (2004) Recombination of human mitochondrial DNA. Science 304:981–983
Lemieux C, Turmel M, Lee RW (1981) Physical evidence for recombination of chloroplast DNA in hybrid progeny of Chlamydomonas eugametos and C. moewusii. Curr Genet 3:97–103
Lemieux C, Turmel M, Seligy VL, Lee RW (1984) Chloroplast DNA recombination in interspecific hybrids of Chlamydomonas: linkage between a nonmendelian locus for streptomycin resistance and restriction fragments coding for 16S rRNA. Proc Natl Acad Sci USA 81:1164–1168
Les DH, Philbrick CT (1993) Studies of hybridization and chromosome number variation in aquatic angiosperms: evolutionary implications. Aquat Bot 44:181–228
Lewandowski I, Clifton-Brown JC, Scurlock JMO, Huisman W (2000) Miscanthus: European experience with a novel energy crop. Biomass Bioenerg 19:209–227
Linde-Laursen I (1993) Cytogenetic analysis of Miscanthus ‘Giganteus’, an interspecific hybrid. Hereditas 119:297–300
Liu L (1997) Miscanthus Anderss. Flora of China, 1st edn, vol 10, pp 4–9. (in Chinese)
Medgyesy P, Fejes E, Maliga P (1985) Interspecific chloroplast recombination in a Nicotiana somatic hybrid. Proc Natl Acad Sci USA 82:6960–6964
Nagamitsu T, Kawahara T, Kanazashi A (2006) Endemic dwarf birch Betula apoiensis (Betulaceae) is a hybrid that originated from Betula ermanii and Betula ovalifolia. Plant Spec Biol 21:19–29
Nagata N (2010) Mechanisms for independent cytoplasmic inheritance of mitochondria and plastids in angiosperms. J Plant Res 123:193–199
Nylander JAA, Wilgenbusch JC, Warren DL, Swofford DL (2008) AWTY (are we there yet?): a system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics 24:581–583
Pan J, Zhang D, Sang T (2007) Molecular phylogenetic evidence for the origin of a diploid hybrid of Paeonia (Paeoniaceae). Am J Bot 94:400–408
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Rayburn AL, Crawford J, Rayburn CM, Juvik JA (2009) Genome size of three Miscanthus species. Plant Mol Biol Rep 27:184–188
Rieseberg LH (1997) Hybrid origins of plant species. Annu Rev Ecol Syst 28:359–389
Rokas A, Ladoukakis E, Zouros E (2003) Animal mitochondrial DNA recombination revisited. Trends Ecol Evol 18:411–417
Seehausen O (2004) Hybridization and adaptive radiation. Trends Ecol Evol 19:198–207
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381
Städler T, Delph LF (2002) Ancient mitochondrial haplotypes and evidence for intragenic recombination in a gynodioecious plant. Proc Natl Acad Sci USA 99:11730–11735
Sun Q, Lin Q, Yi ZL, Yang ZR, Zhou FS (2010) A taxonomic revision of Miscanthus s.l. (Poaceae) from China. Bot J Linn Soc 164:178–220
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Takahashi C, Shibata F (2002) Analysis of Miscanthus sacchariflorus and M. sinensis chromosomes by fluorescence in situ hybridization using rDNA and total genomic DNA probes. Chromosome Sci 6:7–11
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Teo LL, Kiew R, Set O, Lee SK, Gan YY (2002) Hybrid status of kuwini, Mangifera odorata Griff. (Anacardiaceae) verified by amplified fragment length polymorphism. Mol Ecol 11:1465–1469
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tsaousis AD, Martin DP, Ladoukakis ED, Posada D, Zouros E (2005) Widespread recombination in published animal mtDNA sequences. Mol Biol Evol 22:925–933
Ujvari B, Dowton M, Madsen T (2007) Mitochondrial DNA recombination in a free-ranging Australian lizard. Biol Lett 3:189–192
Wang S, Zhang LL, Hu JJ, Bao ZM, Liu ZJ (2010) Molecular and cellular evidence for biased mitotic gene conversion in hybrid scallop. BMC Evol Biol 10:6
Wendel JF, Schnabel A, Seelanan T (1995) Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium). Proc Natl Acad Sci USA 92:280–284
White TJ, Bruns TL, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yong ND, Healy J (2003) Gapcoder automates the use of indel characters in phylogenetic analysis. BMC Bioinformatics 4:6
Zhang Q, Sodmergen LY (2003) Examination of the cytoplasmic DNA in male reproductive cells to determine the potential for cytoplasmic inheritance in 295 angiosperm species. Plant Cell Physiol 44:941–951
Zhou R, Shi S, Wu C (2005) Molecular criteria for determining new hybrid species—An application to the Sonneratia hybrids. Mol Phylogenet Evol 35:595–601
Zub HW, Brancourt-Hulmel M (2010) Agronomic and physiological performances of different species of Miscanthus, a major energy crop.A review. Agron Sustain Dev 30:201–214
Acknowledgments
This research was funded by the National High-tech R&D Program (863 Program) of China (2011AA10020903, 2012AA10180104) and the Mendel Biotechnology Inc., USA
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, J., Zhu, M., Ai, X. et al. Molecular evidence for a natural diploid hybrid between Miscanthus sinensis (Poaceae) and M. sacchariflorus . Plant Syst Evol 299, 1367–1377 (2013). https://doi.org/10.1007/s00606-013-0801-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-013-0801-2