Abstract
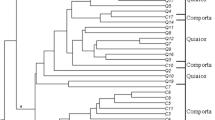
As a popular flowering species with many cultivars, Cymbidium ensifolium (L.) is commercially important in horticulture. However, so far little has been known about genetic diversity and conservation genetics of this species. Understanding of the genetic variation and relationships in cultivars of C. ensifolium is a prerequisite for development of future germplasm conservation and cultivar improvement. Here we report assessment of genetic variations in C. ensifolium cultivars using the DNA fingerprinting technique of inter-simple sequence repeats (ISSR). A total of 239 ISSR loci were identified and used for evaluation of genetic variation with a selection of 19 ISSR primers. Among these ISSR loci, 99.16% were polymorphic with wide genetic variation as shown by Nei’s gene diversity (H = 0.2431) among 85 tested cultivars. ISSR fingerprinting profiles showed that each cultivar had its characteristic DNA pattern, indicating unequivocal cultivar identification at molecular level. Eighteen cultivar-specific ISSR markers were identified in seven cultivars. The cultivar Sijiwenhan was confirmed as hybrid by four ISSR primers. Several cultivars with same name but different geographical origins were distinguished based on their ISSR profiles. A dendrogram generated with ISSR markers could group 73 of 85 cultivars into four major clusters. Further analysis of ISSR variation revealed that about 69% of total genetic variation in this species is due to genetic divergence inside geographical groups. Our results suggest that both germplasm collection and in situ conservation are important for future planning of C. ensifolium species conservation.



Similar content being viewed by others
Abbreviations
- ISSR:
-
Inter-simple sequence repeats
- PCA:
-
Principal component analysis
- PCR:
-
Polymerase chain reaction
- UPGMA:
-
Unweighted pair-group method with arithmetic means
References
Campbell VV, Rowe G, Beebee TJC, Hutchings MJ (2002) Isolation and characterization of microsatellite primers for the fragrant orchid Gymnadenia conopsea (L.) R. Brown (Orchidaceae). Conserv Genet 3:209–210
Capesius I (1976) Isolation and characterization of native at-rich satellite DNA from nuclei of the orchid Cymbidium. FEBS Lett 68:255–258
Choi SH, Kim MJ, Lee JS, Ryu KH (2006) Genetic diversity and phylogenetic relationships among and within species of oriental cymbidiums based on RAPD analysis. Sci Hortic 108:79–85
Dressler RL (1993) Phylogeny and classification of the orchid family. Cambridge University Press, Cambridge
Du Puy D, Cribb P (2007) The genus Cymbidium. Royal Botanic Gardens, Kew
Fang DQ, Roose ML (1997) Identification of closely related citrus cultivars with inter-simple sequence repeat markers. Theor Appl Genet 95:408–417
Hew CS (2001) Ancient Chinese orchid cultivation: a fresh look at an age-old practice. Sci Hortic 87:1–10
Hieber AD, Mudalige-Jayawickrama RG, Kuehnle AR (2006) Color genes in the orchid oncidium gower ramsey: identification, expression, and potential genetic instability in an interspecific cross. Planta 223:521–531
Jarne P, Lagoda PJL (1996) Microsatellites, from molecules to populations and back. Trends Ecol Evol 11:424–429
Kar P, Kar P, Srivastava P, Awasthi A, Urs S (2008) Genetic variability and association of ISSR markers with some biochemical traits in mulberry (Morus spp.) genetic resources available in India. Tree Genet Genom 4:75–83
Kimura M, Crow JC (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Lewontin RC (1972) Apportionment of human diversity. Evol Biol 6:381–398
Liu JJ, Podila GK (1997) Characterization of a MADS box gene (Y09611) from immature female cone of red pine (PGR97-032). Plant Physiol 113:665
Liu JJ, Ekramoddoullah AKM, Podila GK (2003) A MADS-box gene specifically expressed in the reproductive tissues of red pine (Pinus resinosa) is a homologue to floral homeotic genes with C-function in angiosperms. Physiol Mol Bio Plants 9:197–206
Liu ZJ, Chen SC, Ru ZZ, Chen LJ (2006a) Chinese Cymbidium plants. Science, Beijing
Liu JJ, Ekramoddoullah AKM, Hunt RS, Zamani A (2006b) Identification and characterization of random amplified polymorphic DNA markers linked to a major gene (Cr2) for resistance to Cronartium ribicola in Pinus monticola. Phytopathology 96:395–399
Mondragón-Palomino M, Theißen G (2009) Why are orchid flowers so diverse? Reduction of evolutionary constraints by paralogues of class B floral homeotic genes. Ann Bot 104:583–594
Nash N (1996) Flavar of the month, Cymbidium ensifolium. Orchids 9:972–974
Nei M (1973) Analysis of gene diversity in subdivided population. Proc Natl Acad Sci USA 70:3321–3323
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 74:5269–5273
Obara-Okeyo P, Kako S (1998) Genetic diversity and identification of Cymbidium cultivars as measured by random amplified polymorphic DNA (RAPD) markers. Euphytica 99:95–101
Obara-Okeyo P, Fujii K, Kako S (1998) Isozyme variation in Cymbidium species (Orchidaceae). Hort Sci 33:133–135
Powell W, Morgante M, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Raina SN, Rani V, Kojima T, Ogihara Y, Singh KP, Devarumath RM (2001) RAPD and ISSR fingerprints as useful genetic markers for analysis of genetic diversity, varietal identification, and phylogenetic relationships in peanut (Arachis hypogaea) cultivars and wild species. Genome 44:763–772
Van den Berg C, Ryan A, Cribb P, Chase MW (2002) Molecular phylogenetics of Cymbidium (Orchidaceae: Maxillarieae): sequence data from internal transcribed spacers (ITS) of nuclear ribosomal DNA and plastid matK. Lindleyana 17:102–111
Wang HZ, Wang YD, Zhou XY, Ying QC, Zheng KL (2004) Analysis of genetic diversity of 14 species of Cymbidium based on RAPDs and AFLPs. Acta Biol Exp Sin 37:482–486
Wang HZ, Feng SG, Lu JJ, Shi NN, Liu JJ (2009a) Phylogenetic study and molecular identification of 31 Dendrobium species using inter-simple sequence repeat (ISSR) markers. Sci Hortic 122:440–447
Wang HZ, Wu ZX, Lu JJ, Shi NN, Zhao Y, Zhang ZT, Liu JJ (2009b) Molecular diversity and relationships among Cymbidium goeringii cultivars based on inter-simple sequence repeat (ISSR) markers. Genetica 136:391–399
Weber JL (1990) Informativeness of human (dC-dA)n · (dG-dT)n polymorphisms. Genomics 7:524–530
Wolff K, Morgan-Richards M (1998) PCR markers distinguish Plantago major subspecies. Theor Appl Genet 96:282–286
Wu YS (1993) Cymbidium plants in China, 2nd edn. Chinese Forestry, Beijing
Yeh FC, Yang RC, Boyle TBJ, Ye ZH, Mao JX (2000) POPGENE ver. 1.32: the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre University of Alberta Canada
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome Fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Acknowledgments
This research was funded in part by the National Natural Science Foundation (Nos. 30670199, 30870180 and 31070298), the Natural Science Foundation of Zhejiang Province (Nos. X305692 and X301406), the Zhejiang Scientific and Technological Program (2008C12081), the Hangzhou Scientific and Technological Program (2005132H06), and the Qianjiang Scholar Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, HZ., Lu, JJ., Hu, X. et al. Genetic variation and cultivar identification in Cymbidium ensifolium . Plant Syst Evol 293, 101–110 (2011). https://doi.org/10.1007/s00606-011-0429-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-011-0429-z