Abstract
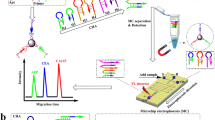
A versatile triple cascade amplification strategy was developed for ultrasensitive simultaneous detection of multiple cancer biomarkers using single particle inductively coupled plasma mass spectrometry (spICP-MS). The triple cascade amplification strategy consisted of an enhanced RecJf exonuclease-assisted target recycling amplification module, a hybridization chain reaction amplification module, and a signal amplification module based on DNA-templated multiple metal nanoclusters. In the enhanced RecJf exonuclease-assisted target recycling amplification module, the DNA bases at the 5′ ends of aptamers for specific recognition of biomarkers were deliberately replaced by the corresponding RNA bases to enhance amplification efficiency. The signal amplification module based on DNA-templated multiple metal nanoclusters was innovatively used to amplify the signals measured by spICP-MS and at the same time effectively suppress possible background interferences. The proposed spICP-MS platform achieved satisfactory quantitative results for both carcinoembryonic antigen (CEA) and a-fetoprotein (AFP) in human serum samples with accuracy comparable to that of the commercial ELISA kits. Moreover, it has wide dynamic ranges for both CEA (0.01–100 ng/mL) and AFP (0.01–200 ng/mL). The limit of detection for CEA and AFP was 0.6 and 0.5 pg/mL, respectively. Compared with conventional biomarkers detection methods, the proposed spICP-MS platform has the advantages of operational simplicity, ultra-high sensitivity, wide dynamic range, and low background. Therefore, it is reasonable to expect that the proposed spICP-MS platform can be further developed to be a promising alternative tool for biomarker detection in fields of clinical diagnosis and biomedical research.
Graphical Abstract








Similar content being viewed by others
References
Borrebaeck CAK, Wingren C (2009) Design of high-density antibody microarrays for disease proteomics: key technological issues. J Proteomics 72(6):928–935. https://doi.org/10.1016/j.jprot.2009.01.027
Poller W, Dimmeler S, Heymans S, Zeller T, Haas J, Karakas M, Leistner D-M, Jakob P, Nakagawa S, Blankenberg S, Engelhardt S, Thum T, Weber C, Meder B, Hajjar R, Landmesser U (2017) Non-coding RNAs in cardiovascular diseases: diagnostic and therapeutic perspectives. Eur Heart J 39(29):2704–2716. https://doi.org/10.1093/eurheartj/ehx165
Gao Y, Huo W, Zhang L, Lian J, Tao W, Song C, Tang J, Shi S, Gao Y (2019) Multiplex measurement of twelve tumor markers using a GMR multi-biomarker immunoassay biosensor. Biosens Bioelectron 123:204–210. https://doi.org/10.1016/j.bios.2018.08.060
Pereira LHM, Adebisi IN, Perez A, Wiebel M, Reis I, Duncan R, Goodwin WJ, Hu JJ, Lokeshwar VB, Franzmann EJ (2012) Salivary markers and risk factor data: a multivariate modeling approach for head and neck squamous cell carcinoma detection. Cancer Biomark 10:241–249. https://doi.org/10.3233/CBM-2012-0252
Ye S, Chen X, Yao Y, Li Y, Sun R, Zeng H, Shu Y, Yin H (2019) Thioredoxin reductase as a novel and efficient plasma biomarker for the detection of non-small cell lung cancer: a large-scale, multicenter study. Sci Rep 9(1):2652. https://doi.org/10.1038/s41598-018-38153-7
Lequin RM (2005) Enzyme immunoassay (EIA)/enzyme-linked immunosorbent assay (ELISA). Clin Chem 51(12):2415–2418. https://doi.org/10.1373/clinchem.2005.051532
He M, Herr AE (2010) Polyacrylamide gel photopatterning enables automated protein immunoblotting in a two-dimensional microdevice. J Am Chem Soc 132(8):2512–2513. https://doi.org/10.1021/ja910164d
Mishra M, Tiwari S, Gomes AV (2017) Protein purification and analysis: next generation Western blotting techniques. Expert Rev Proteomics 14(11):1037–1053. https://doi.org/10.1080/14789450.2017.1388167
Zheng Z, Geng W-C, Gao J, Wang Y-Y, Sun H, Guo D-S (2018) Ultrasensitive and specific fluorescence detection of a cancer biomarker via nanomolar binding to a guanidinium-modified calixarene. Chem Sci 9(8):2087–2091. https://doi.org/10.1039/C7SC04989G
Gallo V, Lai A, Pasquo A, Almaviva S, Iacobelli S, Persichetti L, Capellini G, Antonini G (2020) Surface-enhanced Raman scattering (SERS)–based immunosystem for ultrasensitive detection of the 90 K biomarker. Anal Bioanal Chem 412(27):7659–7667. https://doi.org/10.1007/s00216-020-02903-2
Krishnan S, Mani V, Wasalathanthri D, Kumar CV, Rusling JF (2011) Attomolar detection of a cancer biomarker protein in serum by surface plasmon resonance using superparamagnetic particle labels. Angew Chem Int Ed 50(5):1175–1178. https://doi.org/10.1002/anie.201005607
Cui W, He M, Mu L, Lin Z, Wang Y, Pang W, Reed M, Duan X (2018) Cellphone-enabled microwell-based microbead aggregation assay for portable biomarker detection. ACS sensors 3(2):432–440. https://doi.org/10.1021/acssensors.7b00866
Wang J, Wu L, Ren J, Qu X (2012) Visualizing human telomerase activity with primer-modified Au nanoparticles. Small 8(2):259–264. https://doi.org/10.1002/smll.201101938
Sawhney MA, Conlan RS (2019) POISED-5, a portable on-board electrochemical impedance spectroscopy biomarker analysis device. Biomed Microdevice 21(3):70. https://doi.org/10.1007/s10544-019-0406-9
Mitrano DM, Lesher EK, Bednar A, Monserud J, Higgins CP, Ranville JF (2012) Detecting nanoparticulate silver using single-particle inductively coupled plasma–mass spectrometry. Environ Toxicol Chem 31(1):115–121. https://doi.org/10.1002/etc.719
Montoro Bustos AR, Petersen EJ, Possolo A, Winchester MR (2015) Post hoc interlaboratory comparison of single particle ICP-MS size measurements of NIST gold nanoparticle reference materials. Anal Chem 87(17):8809–8817. https://doi.org/10.1021/acs.analchem.5b01741
Peters R, Herrera-Rivera Z, Undas A, van der Lee M, Marvin H, Bouwmeester H, Weigel S (2015) Single particle ICP-MS combined with a data evaluation tool as a routine technique for the analysis of nanoparticles in complex matrices. J Anal At Spectrom 30(6):1274–1285. https://doi.org/10.1039/C4JA00357H
Han G, Xing Z, Dong Y, Zhang S, Zhang X (2011) One-step homogeneous DNA assay with single-nanoparticle detection. Angew Chem Int Ed 50(15):3462–3465. https://doi.org/10.1002/anie.201006838
Hu J, Deng D, Liu R, Lv Y (2018) Single nanoparticle analysis by ICPMS: a potential tool for bioassay. J Anal At Spectrom 33(1):57–67. https://doi.org/10.1039/C7JA00235A
Li B-R, Tang H, Yu R-Q, Jiang J-H (2020) Single-nanoparticle ICPMS DNA assay based on hybridization-chain-reaction-mediated spherical nucleic acid assembly. Anal Chem 92(3):2379–2382. https://doi.org/10.1021/acs.analchem.9b05741
Liu X, Zhang S-Q, Cheng Z-H, Wei X, Yang T, Yu Y-L, Chen M-L, Wang J-H (2018) Highly sensitive detection of microRNA-21 with ICPMS via hybridization accumulation of upconversion nanoparticles. Anal Chem 90(20):12116–12122. https://doi.org/10.1021/acs.analchem.8b03038
Wu N, Wang K, Wang Y-T, Chen M-L, Chen X-W, Yang T, Wang J-H (2020) Three-dimensional DNA nanomachine biosensor by integrating DNA walker and rolling machine cascade amplification for ultrasensitive detection of cancer-related gene. Anal Chem 92(16):11111–11118. https://doi.org/10.1021/acs.analchem.0c01074
Xing Y, Han J, Wu X, Pierce DT, Zhao JX (2020) Graphene/gold nanoparticle composites for ultrasensitive and versatile biomarker assay using single-particle inductively-coupled plasma/mass spectrometry. Analyst 145(24):7932–7940. https://doi.org/10.1039/D0AN01019G
Xu X, Chen J, Li B, Tang L, Jiang J (2019) Single particle ICP-MS-based absolute and relative quantification of E. coli O157 16S rRNA using sandwich hybridization capture. The Analyst 144(5):1725–1730. https://doi.org/10.1039/C8AN02063A
Zhang S, Han G, Xing Z, Zhang S, Zhang X (2014) Multiplex DNA assay based on nanoparticle probes by single particle inductively coupled plasma mass spectrometry. Anal Chem 86(7):3541–3547. https://doi.org/10.1021/ac404245z
Du X, Jin R (2019) Atomically precise metal nanoclusters for catalysis. ACS Nano 13(7):7383–7387. https://doi.org/10.1021/acsnano.9b04533
Jin R, Zeng C, Zhou M, Chen Y (2016) Atomically precise colloidal metal nanoclusters and nanoparticles: fundamentals and opportunities. Chem Rev 116(18):10346–10413. https://doi.org/10.1021/acs.chemrev.5b00703
Chen C, Zhao J, Lu Y, Sun J, Yang X (2018) Fluorescence immunoassay based on the phosphate-triggered fluorescence turn-on detection of alkaline phosphatase. Anal Chem 90(5):3505–3511. https://doi.org/10.1021/acs.analchem.7b05325
Chen Y, Liu H, Jiang J, Gu C, Zhao Z, Jiang T (2020) Immunoassay of tumor markers based on graphene surface-enhanced Raman spectroscopy. ACS Appl Bio Mater 3(11):8012–8022. https://doi.org/10.1021/acsabm.0c01098
Huang Z, Li Z, Jiang M, Liu R, Lv Y (2020) Homogeneous multiplex immunoassay for one-step pancreatic cancer biomarker evaluation. Anal Chem 92(24):16105–16112. https://doi.org/10.1021/acs.analchem.0c03780
Jie G, Li C, Zhao Y, Kuang Q, Niu S (2019) Fluorescent Mn:ZnCdS@ZnS and CdTe quantum dots probes on SiO2 microspheres for versatile detection of carcinoembryonic antigen and monitoring T4 polynucleotide kinase activity. ACS Applied Nano Materials 2(7):4637–4645. https://doi.org/10.1021/acsanm.9b01003
Li C, Ma X, Guan Y, Tang J, Zhang B (2019) Microcantilever array biosensor for simultaneous detection of carcinoembryonic antigens and α-fetoprotein based on real-time monitoring of the profile of cantilever. ACS sensors 4(11):3034–3041. https://doi.org/10.1021/acssensors.9b01604
Liu Y, Du M, Zhu J, Hu X, Ji X, He Z (2018) Three-dimensional immunosensing platform based on a hybrid nanoflower for sensitive detection of α-fetoprotein and enterovirus 71. ACS Applied Nano Materials 1(9):4964–4971. https://doi.org/10.1021/acsanm.8b01109
Qiu Z, Shu J, Tang D (2018) Near-infrared-to-ultraviolet light-mediated photoelectrochemical aptasensing platform for cancer biomarker based on core–shell NaYF4:Yb, Tm@TiO2 upconversion microrods. Anal Chem 90(1):1021–1028. https://doi.org/10.1021/acs.analchem.7b04479
Song Y, Zhang W, He S, Shang L, Ma R, Jia L, Wang H (2019) Perylene diimide and luminol as potential-resolved electrochemiluminescence nanoprobes for dual targets immunoassay at low potential. ACS Appl Mater Interfaces 11(37):33676–33683. https://doi.org/10.1021/acsami.9b11416
Xiao L, Zhu A, Xu Q, Chen Y, Xu J, Weng J (2017) Colorimetric biosensor for detection of cancer biomarker by au nanoparticle-decorated Bi2Se3 nanosheets. ACS Appl Mater Interfaces 9(8):6931–6940. https://doi.org/10.1021/acsami.6b15750
Zhang C, Gao Y, Yang N, You T, Chen H, Yin P (2018) Direct determination of the tumor marker AFP via silver nanoparticle enhanced SERS and AFP-modified gold nanoparticles as capturing substrate. Microchim Acta 185(2):90. https://doi.org/10.1007/s00604-017-2652-y
Acknowledgements
The authors acknowledge the financial support of the National Natural Science Foundation of China (no. 22273020), the Leading Plan of Science and Technology Innovation in Hunan High-Tech Industry (2020SK2029), and the Research Foundation of Education Bureau of Hunan Province (no. 21C0474).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, YL., Wang, JK., Chen, ZP. et al. Ultrasensitive detection of multiple cancer biomarkers by a triple cascade amplification strategy in combination with single particle inductively coupled plasma mass spectrometry. Microchim Acta 190, 20 (2023). https://doi.org/10.1007/s00604-022-05604-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05604-y