Abstract
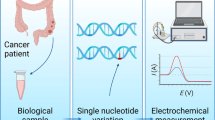
Two electrochemical bioplatforms were prepared based on thiolated hairpin DNA probes tethered to AuNP-modified screen-printed electrodes to detect T > G and T > C polymorphisms, namely rs1880269 and rs1800469, present the interleukin-6 (IL6) and transforming growth factor β1 (TGFβ1) genes. The electrochemical readout was ensured by the detection of the double-stranded DNA using methylene blue as a redox probe after treatment by EcoRI restrictase. The main parameters influencing the analytical response such as the thiolated DNA probe concentration, incubation time with electrode, DNA hybridization time, EcoRI enzyme load, and its cleavage time were optimized based on the current intensity and signal-to-blank (S/B) ratio as selection criteria. Using spiked buffer solutions, the IL6 and TGFβ1 E-bioplatforms display wide ranges of linearity (1 × 102–1 × 108 fM and 5 × 101–1 × 105 fM, respectively) and limits of detection (47.9 fM and 16.6 fM, respectively). The two bioelectrodes have also good discrimination toward 1-mismatched, two mismatched, and non-complementary sequences, when they were used 30-fold higher than the target sequences. More importantly, the two bioplatforms successfully detected the single nucleotide polymorphisms (SNPs) in scarcely diluted genomic DNA, collected from 52 donors, and showed they can reliably distinguish between heterozygous (TG and TC genotypes) and homozygous (GG and CC genotypes) patients with respect to the control subjects (TT genotype), where the differences are statistically highly significant (p-value < 0.0001). Thus, the designed devices could be used to conduct large cohort studies targeting these mutations or extended to other SNPs.
Graphical Abstract





Similar content being viewed by others
References
Erichsen HC, Chanock SJ (2004) SNPs in cancer research and treatment. Br J Cancer 90:747–751. https://doi.org/10.1038/sj.bjc.6601574
Mahdi KM, Nassiri MR, Nasiri K (2013) Hereditary genes and SNPs associated with breast cancer. Asian Pac J Cancer Prev 14:3403–3409. https://doi.org/10.7314/apjcp.2013.14.6.3403
Nassiri M, Kooshyar MM, Roudbar Z, Mahdavi M, Doosti M (2013) Genes and SNPs associated with non-hereditary and hereditary colorectal cancer. Asian Pac J Cancer Prev 14:5609–5614. https://doi.org/10.7314/apjcp.2013.14.10.5609
Ben Ahmed A, Zidi S, Sghaier I, Ghazouani E, Mezlini A, Almawi W et al (2017) Common variants in IL-1RN, IL-1beta and TNF-alpha and the risk of ovarian cancer: a case control study. Cent Eur J Immunol 42:150–155. https://doi.org/10.5114/ceji.2017.69356
Butler JM, Hall N, Narendran N, Yang YC, Paraoan L (2017) Identification of candidate protective variants for common diseases and evaluation of their protective potential. BMC Genomics 18:575. https://doi.org/10.1186/s12864-017-3964-3
Schwartz MLB, Williams MS, Murray MF (2017) Adding protective genetic variants to clinical reporting of genomic screening results: restoring balance. JAMA 317:1527–1528. https://doi.org/10.1001/jama.2017.1533
Gudmundsdottir K, Ashworth A (2006) The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene 25:5864–5874. https://doi.org/10.1038/sj.onc.1209874
Chen S, Parmigiani G (2007) Meta-analysis of BRCA1 and BRCA2 penetrance. J Clin Oncol 25:1329–1333. https://doi.org/10.1200/JCO.2006.09.1066
Antoniou A, Pharoah PD, Narod S, Risch HA, Eyfjord JE, Hopper JL et al (2003) Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 72:1117–1130. https://doi.org/10.1086/375033
Kuchenbaecker KB, Hopper JL, Barnes DR, Phillips KA, Mooij TM, Roos-Blom MJ et al (2017) Risks of Breast, Ovarian, and Contralateral Breast Cancer for BRCA1 and BRCA2 Mutation Carriers. JAMA 317:2402–2416. https://doi.org/10.1001/jama.2017.7112
Ben Ahmed A, Zidi S, Almawi W, Ghazouani E, Mezlini A, Loueslati BY et al (2020) Single nucleotide polymorphism of transforming growth factor-β1 and interleukin-6 as risk factors for ovarian cancer. Cent Eur J Immunol 45:267–275. https://doi.org/10.5114/ceji.2020.101242
Lucarelli F, Tombelli S, Minunni M, Marrazza G, Mascini M (2008) Electrochemical and piezoelectric DNA biosensors for hybridisation detection. Anal Chim Acta 609:139–159. https://doi.org/10.1016/j.aca.2007.12.035
Babamiri B, Bahari D, Salimi A (2019) Highly sensitive bioaffinity electrochemiluminescence sensors: recent advances and future directions. Biosens Bioelectron 142:111530. https://doi.org/10.1016/j.bios.2019.111530
Campuzano S, Yanez-Sedeno P, Pingarron JM (2017) Electrochemical genosensing of circulating biomarkers. Sensors-Basel 17:866. https://doi.org/10.3390/s17040866
Campuzano S, Torrente-Rodriguez RM, Lopez-Hernandez E, Conzuelo F, Granados R, Sanchez-Puelles JM et al (2014) Magnetobiosensors based on viral protein p19 for microRNA determination in cancer cells and tissues. Angew Chem Int Ed Engl 53:6168–6171. https://doi.org/10.1002/anie.201403270
Vargas E, Povedano E, Montiel VR, Torrente-Rodriguez RM, Zouari M, Montoya JJ et al. (2018) Single-step incubation determination of miRNAs in cancer cells using an amperometric biosensor based on competitive hybridization onto magnetic beads. Sensors-Basel 18:863. https://doi.org/10.3390/s18030863
Zouari M, Campuzano S, Pingarron JM, Raouafi N (2018) Amperometric biosensing of miRNA-21 in serum and cancer cells at nanostructured platforms using anti-DNA-RNA hybrid antibodies. ACS Omega 3:8923–8931. https://doi.org/10.1021/acsomega.8b00986
Zouari M, Campuzano S, Pingarron JM, Raouafi N (2017) Competitive RNA-RNA hybridization-based integrated nanostructured-disposable electrode for highly sensitive determination of miRNAs in cancer cells. Biosens Bioelectron 91:40–45. https://doi.org/10.1016/j.bios.2016.12.033
Zouari M, Campuzano S, Pingarrón JM, Raouafi N (2020) Femtomolar direct voltammetric determination of circulating miRNAs in sera of cancer patients using an enzymeless biosensor. Anal Chim Acta 1104:188–198. https://doi.org/10.1016/j.aca.2020.01.016
Yammouri G, Mohammadi H, Amine A (2019) A highly sensitive electrochemical biosensor based on carbon black and gold nanoparticles modified pencil graphite electrode for microRNA-21 detection. Chem Afr 2:291–300. https://doi.org/10.1007/s42250-019-00058-x
Zouari M, Campuzano S, Pingarron JM, Raouafi N (2020) Determination of miRNAs in serum of cancer patients with a label- and enzyme-free voltammetric biosensor in a single 30-min step Microchim Acta 187:444. https://doi.org/10.1007/s00604-020-04400-w
Zayani R, Rabti A, Ben Aoun S, Raouafi N (2021) Fluorescent and electrochemical bimodal bioplatform for femtomolar detection of microRNAs in blood sera. Sensor Actuat B-Chem 327:128950. https://doi.org/10.1016/j.snb.2020.128950
Rabti A, Zayani R, Meftah M, Salhi I, Raouafi N (2020) Impedimetric DNA E-biosensor for multiplexed sensing of Escherichia coli and its virulent f17 strains. Microchim Acta 187:635. https://doi.org/10.1007/s00604-020-04614-y
Haddaoui M, Sola C, Raouafi N, Korri-Youssoufi H (2019) E-DNA detection of rpoB gene resistance in Mycobacterium tuberculosis in real samples using Fe3O4/polypyrrole nanocomposite. Biosens Bioelectron 128:76–82. https://doi.org/10.1016/j.bios.2018.11.045
Qicai L, Qiang Y, Wennan W, Yu W, Liqing L, Chengfei Z et al (2015) DNA electrochemical sensor for detection of PRSS1 point mutation based on restriction endonuclease technique. Prep Biochem 45:430–437. https://doi.org/10.1080/10826068.2014.940971
Lin L, Liu A, Zhao C, Weng S, Lei Y, Liu Q et al (2013) A chronocoulometric LNA sensor for amplified detection of K-ras mutation based on site-specific DNA cleavage of restriction endonuclease. Biosens Bioelectron 42:409–414. https://doi.org/10.1016/j.bios.2012.09.063
Sun X, Wang S, Zhang Y, Tian Y, Zhou N (2017) Ultrasensitive detection of DNA based on target-triggered hairpin assembly and exonuclease-assisted recycling amplification. Sens Actuat B: Chem 252:306–312. https://doi.org/10.1016/j.snb.2017.06.014
Szczelkun MD, Halford SE (2013) Restriction endonuclease. In: Maloy S, Hughes K (eds) Brenner’s encyclopedia of genetics, 2nd edn. Academic Press, San Diego, pp 184–189
Buckhout-White S, Person C, Medintz IL, Goldman ER (2018) Restriction enzymes as a target for DNA-based sensing and structural rearrangement. ACS Omega 3:495–502. https://doi.org/10.1021/acsomega.7b01333
Yang H, Peng Y, Xu M, Xu S, Zhou Y (2021) Development of DNA biosensors based on DNAzymes and nucleases. Crit Rev Anal Chem 1–16 (under press). https://doi.org/10.1080/10408347.2021.1944046
Smith M, Smith K, Olstein A, Oleinikov A, Ghindilis A (2020) Restriction endonuclease-based assays for DNA detection and isothermal exponential signal amplification. Sensors-Basel 20:3873
Lou J, Liu S, Tu W, Dai Z (2015) Graphene quantums dots combined with endonuclease cleavage and bidentate chelation for Highly Sensitive Electrochemiluminescent DNA Biosensing. Anal Chem 87:1145–1151. https://doi.org/10.1021/ac5037318
Smith MW, Ghindilis AL, Seoudi IA, Smith K, Billharz R, Simon HM (2014) A new restriction endonuclease-based method for highly-specific detection of DNA targets from methicillin-resistant Staphylococcus aureus. PLoS ONE 9:e97826. https://doi.org/10.1371/journal.pone.0097826
Rabti A, Zayani R, Meftah M, Salhi I, Raouafi N (2020) Impedimetric DNA E-biosensor for multiplexed sensing of Escherichia coli and its virulent f17 strains. Microchim Acta 187:635. https://doi.org/10.1007/s00604-020-04614-y
Argoubi W, Saadaoui M, Ben Aoun S, Raouafi N (2015) Optimized design of a nanostructured SPCE-based multipurpose biosensing platform formed by ferrocene-tethered electrochemically-deposited cauliflower-shaped gold nanoparticles. Beilstein J Nanotechnol 6:1840–1852. https://doi.org/10.3762/bjnano.6.187
Oliveira DA, Silva JV, Flauzino JMR, Castro ACH, Moço ACR, Soares MMCN et al (2018) Application of nanomaterials for the electrical and optical detection of the hepatitis B virus. Anal Biochem 549:157–163. https://doi.org/10.1016/j.ab.2018.03.023
Mannelli I, Minunni M, Tombelli S, Wang R, Michela Spiriti M, Mascini M (2005) Direct immobilisation of DNA probes for the development of affinity biosensors. Bioelectrochemistry 66:129–138. https://doi.org/10.1016/j.bioelechem.2004.04.008
Sahli R, Fave C, Raouafi N, Boujlel K, Schollhorn B, Limoges B (2013) Switching on/off the chemisorption of thioctic-based self-assembled monolayers on gold by applying a moderate cathodic/anodic potential. Langmuir 29:5360–5368. https://doi.org/10.1021/la401117u
Zouari M, Campuzano S, Pingarron JM, Raouafi N (2018) Ultrasensitive determination of microribonucleic acids in cancer cells with nanostructured-disposable electrodes using the viral protein p19 for recognition of ribonucleic acid/microribonucleic acid homoduplexes. Electrochim Acta 262:39–47. https://doi.org/10.1016/j.electacta.2017.12.190
Chérif N, Zouari M, Amdouni F, Mefteh M, Ksouri A, Bouhaouala-Zahar B, Raouafi N (2021) Direct Amperometric Sensing of Fish Nodavirus RNA Using Gold Nanoparticle/DNA-Based Bioconjugates
Zayani R, Rezig D, Fares W, Marrakchi M, Essafi M, Raouafi N (2021) Multiplexed magnetofluorescent bioplatform for the sensitive detection of SARS-CoV-2 viral RNA without nucleic acid amplification. Anal Chem 93:11225–11232. https://doi.org/10.1021/acs.analchem.1c01950
Xu H, Wang L, Ye H, Yu L, Zhu X, Lin Z et al (2012) An ultrasensitive electrochemical impedance sensor for a special BRCA1 breast cancer gene sequence based on lambda exonuclease assisted target recycling amplification. Chem Commun 48:6390–6392. https://doi.org/10.1039/c2cc31588b
Funding
The authors acknowledge the financial support of the Tunisian PRF program (NanofastResponse, grant ref.: PRF2017D4P1 and SmartBioSens, grant ref.: PRFCOV19–D2P2).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meftah, M., Habel, A., Baachaoui, S. et al. Sensitive electrochemical detection of polymorphisms in IL6 and TGFβ1 genes from ovarian cancer DNA patients using EcoRI and DNA hairpin-modified gold electrodes. Microchim Acta 190, 15 (2023). https://doi.org/10.1007/s00604-022-05595-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05595-w