Abstract
A sensor array is described that consists of a carbon quantum dot (CQD) and metal ions, including Hg2+, Cu2+, Fe3+, Ag+, Cd2+, and Pb2+. The CQDs display blue fluorescence with excitation/emission maxima at 370/440 nm. It is shown that the array can be applied to the determination of all natural amino acids (NAAs). Metal ions can quench the fluorescence of the CQDs, while NAAs can take metal ions away or co-bind to the CQD@metal-ion complex, which enhances or depresses the fluorescence of the CQDs. Based on the differential fluorescence variation, the CQD@metal-ion@NAA array exhibits a unique pattern for NAAs. Principal component analysis (PCA) and hierarchical cluster analysis (HCA) were carried out to generate visualized datagrams for NAA discrimination. The design and construction of the sensor array is convenient and economical. The sensor array can distinguish NAAs at a concentration of as low as 30 μM, and can distinguish NAAs into acidic, neutral and basic NAAs. Semi-quantitative assay of alanine and threonine was also realized. Based on the low limit of detection and multi-NAA detection capability, the array can differentiate healthy cases from acute leukemia, chronic leukemia and lymphoma by analyzing the NAA status of serum samples.

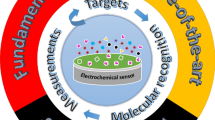
Schematic representation of a fluorometric (FL) sensor array based on single CQD (carbon quantum dot) interacting with metal ions for differentiating all NAAs (natural amino acids) into acidic, neutral and basic NAAs (ANs, NNs and BNs) through PCA (principle component analysis).







Similar content being viewed by others
Abbreviations
- CQDs:
-
Carbon quantum dots
- NAA:
-
Natural amino acids
References
Wu G (2009) Amino acids: metabolism, functions, and nutrition. Amino Acids 37:1–17
Alvarez B, Radi R (2003) Peroxynitrite reactivity with amino acids and proteins. Amino Acids 25:295–311
Zaia DAM (2004) A review of adsorption of amino acids on minerals: was it important for origin of life? Amino Acids 27:113–118
Fernstrom JD (2013) Large neutral amino acids: dietary effects on brain neurochemistry and function. Amino Acids 45:419–430
Wang WW, Qiao SY, Li DF (2009) Amino acids and gut function. Amino Acids 37:105–110
Trevino SR, Scholtz JM, Pace CN (2007) Amino acid contribution to protein solubility: asp, Glu, and Ser contribute more favorably than the other hydrophilic amino acids in RNase Sa. J Mol Biol 366:449–460
Wu G (2013) Functional amino acids in nutrition and health. Amino Acids 3:407–411
Kuhl A, Hahn MG, Dumic M, Mittendorf J (2005) Alicyclic β-amino acids in medicinal chemistry. Amino Acids 29:89–100
Blalock JE, Smith EM (1984) Hydropathic anti-complementarity of amino acids based on the genetic code. Biochem Biophys Res Commun 121:203–207
Zhou Y, Yoon J (2012) Recent progress in fluorescent and colorimetric chemosensors for detection of amino acids. Chem Soc Rev 41:52–67
Brückner H, Westhauser T (2003) Chromatographic determination of L- and D-amino acids in plants. Amino Acids 24:43–55
Roth M (1971) Fluorescence reaction for amino acids. Anal Chem 43:880–882
Clarkson TW, Kench JE (1956) Urinary excretion of amino acids by men absorbing heavy metals. Biochem J 62:361–372
Minami T, Esipenko NA, Zhang B, Kozelkova ME, Isaacs L, Nishiyabu R, Kubo Y, Jr PA (2012) Supramolecular sensor for Cancer-associated nitrosamines. J Am Chem Soc 134:20021–20024
Xu S, Su Z, Zhuo Z, Nie Y, Wang J, Ge G, Luo X (2017) Rapid synthesis of nitrogen doped carbon dots and their application as a label free sensor array for simultaneous discrimination of multiple proteins. J Mater Chem B 5:8748–8753
Palacios MA, Wang Z, Montes VA, Zyryanov GV, Jr PA (2008) Rational Design of a Minimal Size Sensor Array for metal ion detection. J Am Chem Soc 130:10307–10314
Wu Y, Liu X, Wu Q, Yi J, Zhang G (2017) Carbon Nanodots-based fluorescent turn-on sensor Array for biothiols. Anal Chem 89:7084–7089
Wang B, Han J, Bojanowski NM, Bender M, Ma C, Seehafer K, Herrmann AH, Bunz UHF (2018) An optimized sensor Array identifies all natural amino acids. ACS Sensors 3:1562–1568
Zhang F, Lu C, Wang M, Yu X, Wei W, Xia Z (2018) A chiral sensor Array for peptidoglycan biosynthesis monitoring based on MoS2 Nanosheets-supported host-guest recognitions. ACS Sensors 3:304–312
Wang B, Han J, Ma C, Bender M, Seehafer K, Herrmann A, Bunz UHF (2017) A simple optoelectronic tongue discriminates amino acids. Chem Eur J 23:12471–12474
Zhang W, Gao N, Cui J, Wang C, Wang S, Zhang G, Dong X, Zhang D, Li G (2017) AIE-doped poly[ionic liquid] photonic sphere: a single-sphere based customizable sensing platform for discrimination of Multianalytes. Chem Sci 8:6281–6289
Xiao H, Ting B, Zhou N, Zhuang M, Yang L, Shou Z (2016) Nitrogen-doped carbon nanoparticle modulated turn-on fluorescent probes for histidine detection and its imaging in living cells. Nanoscale. 8:2205–2211
Li X, Rui M, Song J, Shen Z, Zeng H (2015) Carbon and graphene quantum dots for optoelectronic and energy devices: a review. Adv Funct Mater 25:4929–4947
Zhu J, Bai X, Zhai Y, Chen X, Zhu Y, Pan G, Zhang H, Dong B, Song H (2017) Carbon dots with efficient solid-state photoluminescence towards white light-emitting diodes. J Mater Chem C 5:11416–11420
Liu S, Tian J, Wang L, Zhang Y, Qin X, Luo Y, Asiri AM, Al-Youbi AO, Sun X (2012) Hydrothermal treatment of grass: a low-cost, green route to nitrogen-doped, carbon-rich, Photoluminescent polymer Nanodots as an effective fluorescent sensing platform for label-free detection of cu(II) ions. Adv Mater 24:2037–2041
Zhu X, Zhao T, Nie Z, Mao Z, Liu Y, Yao S (2016) Nitrogen-doped carbon nanoparticle modulated turn-on fluorescent probes for histidine detection and its imaging in living cells. Nanoscale 8:2205–2211
Akdeniz A, Mosca L, Minami T, Jr. PA (2015) Sensing of enantiomeric excess in chiral carboxylic acids, Chem. Commun. 51:5770–5773
Hou J, Zhang F, Yan X, Wang L, Yan J, Ding H, Ding L (2015) Sensitive detection of biothiols and histidine based on the recovered fluorescence of the carbon quantum dots–hg(II) system. Anal Chim Acta 859:72–78
Fong JFY, Chin SF, Ng SM (2016) A unique “turn-on” fluorescence signalling strategy for highly specific detection of ascorbic acid using carbon dots as sensing probe. Biosens Bioelectron 85:844–852
Kermit M, Tomic O (2003) Independent component analysis applied on gas sensor Array measurement data. IEEE Sensors J 3:218–228
Williams B, Onsman A, Brown T (2010) Exploratory factor analysis: a five-step guide for novices. J Emerg Primary Health Care (JEPHC) 8(3):990399
Askim JR, Mahmoudi M, Suslick KS (2013) Optical sensor arrays for chemical sensing: the optoelectronic nose. Chem Soc Rev 42:8649–8682
Lim SY, Shen W, Gao Z (2015) Carbon quantum dots and their applications. Chem Soc Rev 44:362–381
Rudman D, Vogler WR, Howard CH, Gerron GG (1971) Observations on the plasma amino acids of patients with acute leukemia. Cancer Res 31:1159–1165
Lu T, Chen L, Lai S, Li J, Zhou Y (1991) Changes of plasma free amino acid concentration in patients with leukemia and lymphoma. Clin Med China 7:136–137
Acknowledgements
This work was supported by National Natural Science Foundation of China (51873085), Natural Science Foundation of Liaoning Province of China (20180510023 and 20180550947), Natural Science Foundation for Education Department of Liaoning Province of China (LYB201603) and Shenyang Municipal Program for the Top Young and Middle-aged Innovative Talents of Liaoning Province of China (RC170258).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no financial and personal relationships with other people or organizations that can inappropriately influence our work, there is no professional or other personal interest of any nature or kind in any product, service and/or company that may be construed as influencing the position presented in, or the review of, the manuscript entitled, “Sensor array based on single carbon quantum dot for differentiating all natural amino acids”.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 418 kb)
Rights and permissions
About this article
Cite this article
Gao, Y., Gao, F., Zhang, G. et al. Sensor array based on single carbon quantum dot for fluorometric differentiation of all natural amino acids. Microchim Acta 186, 858 (2019). https://doi.org/10.1007/s00604-019-3864-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-3864-0