Abstract
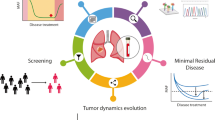
A tissue biopsy is the “golden standard” for molecular profiling that is essential in decision-making regarding treatment for malignant tumors, including primary lung cancer. However, tumor biopsies are associated with several limitations, including invasiveness and difficulty in achieving access. Liquid biopsies have several potential advantages over tissue biopsies, and recent advances in molecular technologies have enabled liquid biopsies to be introduced into daily clinical practice. Cell-free blood-based liquid biopsies to detect mutations in the epidermal growth factor receptor (EGFR) gene in the plasma have been approved and may be useful in selecting patients for treatment with tyrosine kinase inhibitors of EGFR. We herein describe blood-based liquid biopsies and review the current status and future perspectives of plasma genotyping in primary lung cancer.





Similar content being viewed by others
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Herbst RS, Heymach JV, Lippman SM. Lung cancer. N Engl J Med. 2008;359:1367–80.
Rudin CM, Ismaila N, Hann CL, Malhotra N, Movsas B, Norris K, et al. Treatment of small-cell lung cancer: American Society of Clinical Oncology endorsement of the American College of Chest Physicians guideline. J Clin Oncol. 2015;33:4106–11.
Bunn PA Jr, Minna JD, Augustyn A, Bunn PA Jr, Minna JD, Augustyn A, et al. Small cell lung cancer: can recent advances in biology and molecular biology be translated into improved outcomes?. J Thorac Oncol. 2016;11:453–74.
Non-Small Cell Lung Cancer Collaborative Group. Chemotherapy and supportive care versus supportive care alone for advanced non-small cell lung cancer. Cochrane Database Syst Rev 2010;12(5):CD007309.
Hanahan D, Weingberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Swanton C, Govindan R. Clinical implications of genomic discoveries in lung cancer. N Engl J Med. 2016;374:1864–73.
Hirsch FR, Scagliotti GV, Mulshine JL, Kwon R, Curran WJ Jr, Wu YL, et al. Lung cancer: current therapies and new targeted treatments. Lancet. 2017;389:299–311.
Hanna N, Johnson D, Temin S, Baker S Jr, Brahmer J, Ellis PM, et al. Systemic therapy for stage IV non-small-cell lung cancer: American Society of Clinical Oncology Clinical practice guideline update. J Clin Oncol. 2017;35:3484–515.
Yoneda K, Tanaka F. Molecular diagnosis and targeting for lung cancer. In: Shinomiya N, Kataoka H, Shimada Y, editors. Molecular diagnosis and targeting for thoracic and gastrointestinal malignancy. Singapore: Springer; 2018. pp. 1–32.
Saito M, Shiraishi K, Kunitoh H, Takenoshita S, Yokota J, Kohno T. Gene aberrations for precision medicine against lung adenocarcinoma. Cancer Sci. 2016;107:713–20.
Barlesi F, Mazieres J, Merlio JP, Debieuvre D, Mosser J, Lena H, et al. Routine molecular profiling of patients with advanced non-small-cell lung cancer: results of a 1-year nationwide programme of the French Cooperative Thoracic Intergroup (IFCT). Lancet. 2016;387:1415–26.
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol 2017;14:531–48.
Sacher AG, Komatsubara KM, Oxnard GR. Application of plasma genotyping technologies in non-small cell lung cancer: a practical review. J Thorac Oncol. 2017;12:1344–56.
Diaz LA Jr, Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014;32:579–86.
Heitzer E, Ulz P, Geigl JB. Circulating tumor DNA as a liquid biopsy for cancer. Clin Chem. 2015;61:112–23.
Tanaka F, Yoneda K. Adjuvant therapy following surgery in non-small cell lung cancer (NSCLC). Surg Today. 2016;46:25–37.
Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017;17:223–38.
Zhang W, Xia W, Lv Z, Ni C, Xin Y, Yang L. Liquid biopsy for cancer: circulating tumor cells, circulating free DNA or exosomes? Cell Physiol Biochem. 2017;41:755–68.
Wang J, Chang S, Li G, Sun Y. Application of liquid biopsy in precision medicine: opportunities and challenges. Front Med. 2017;11:522–7.
Riethdorf S, O’Flaherty L, Hille C, Pantel K. Clinical applications of the CellSearch platform in cancer patients. Adv Drug Deliv Rev. 2018. https://doi.org/10.1016/j.addr.2018.01.011 (pii: S0169-409X(18)30011-5).
Allard WJ, Matera J, Miller MC, Repollet M, Connelly MC, Rao C, et al. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy or patients with nonmalignant diseases. Clin Cancer Res. 2004;10:6897–904.
Tanaka F, Yoneda K, Kondo N, Hashimoto M, Takuwa T, Matsumoto S, et al. Circulating tumor cell as a diagnostic marker in primary lung cancer. Clin Cancer Res. 2009;15:6980–6.
Naito T, Tanaka F, Ono A, Yoneda K, Takahashi T, Murakami H, et al. Prognostic impact of circulating tumor cells in patients with small cell lung cancer. J Thorac Oncol. 2012;7:512–9.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
Holdhoff M, Schmidt K, Donehower R, Diaz LA Jr. Analysis of circulating tumor DNA to confirm somatic KRAS mutations. J Natl Cancer Inst. 2009;101:1284–5.
Thierry AR, El Messaoudi S, Gahan PB, Anker P, Stroun M. Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev. 2016;35:347–76.
El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: preanalytical considerations. Clin Chim Acta. 2013;424:222–30.
Normanno N, Denis MG, Thress KS, Ratcliffe M, Reck M. Guide to detecting epidermal growth factor receptor (EGFR) mutations in ctDNA of patients with advanced non-small-cell lung cancer. Oncotarget. 2017;8:12501–16.
Garcia J, Dusserre E, Cheynet V, Bringuier PP, Brengle-Pesce K, Wozny AS, et al. Evaluation of pre-analytical conditions and comparison of the performance of several digital PCR assays for the detection of major EGFR mutations in circulating DNA from non-small cell lung cancers: the CIRCAN_0 study. Oncotarget. 2017;8:87980–96.
Vallée A, Marcq M, Bizieux A, Kouri CE, Lacroix H, Bennouna J, et al. Plasma is a better source of tumor-derived circulating cell-free DNA than serum for the detection of EGFR alterations in lung tumor patients. Lung Cancer. 2013;82:373–4.
Li X, Ren R, Ren S, Chen X, Cai W, Zhou F, et al. Peripheral blood for epidermal growth factor receptor mutation detection in non-small cell lung cancer patients. Transl Oncol. 2014;7:341–8.
Luo J, Shen L, Zheng D. Diagnostic value of circulating free DNA for the detection of EGFR mutation status in NSCLC: a systematic review and meta-analysis. Sci Rep. 2014;4:6269.
Reck M, Hagiwara K, Han B, Tjulandin S, Grohé C, Yokoi T, et al. ctDNA Determination of EGFR mutation status in European and Japanese patients with advanced NSCLC: the ASSESS study. J Thorac Oncol. 2016;11:1682–9.
Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, et al. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem. 2012;84:1003–11.
Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O’Connell A, Messineo MM, et al. Noninvasive detection of response and resistance in EGFR‑mutant lung cancer using quantitative next‑generation genotyping of cell‑free plasma DNA. Clin Cancer Res. 2014;20:1698–705.
Dressman D, Yan H, Traverso G, Kinzler KW, Vogelstein B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc Natl Acad Sci USA. 2003;100:8817–22.
Li M, Diehl F, Dressman D, Vogelstein B, Kinzler KW. BEAMing up for detection and quantification of rare sequence variants. Nat Methods. 2006;3:95–7.
Yohe S, Thyagarajan B. Review of clinical next-generation sequencing. Arch Pathol Lab Med. 2017;141:1544–57.
Vendrell JA, Grand D, Rouquette I, Costes V, Icher S, Selves J, et al. High-throughput detection of clinically targetable alterations using next-generation sequencing. Oncotarget. 2017;8:40345–58.
Vendrell JA, Mau-Them FT, Béganton B, Godreuil S, Coopman P, Solassol J. Circulating cell free tumor DNA detection as a routine tool for lung cancer patient management. Int J Mol Sci. 2017;18:E264.
Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20:548–54.
Lanman RB, Mortimer SA, Zill OA, Sebisanovic D, Lopez R, Blau S, et al. Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS One. 2015;10:e0140712.
Qiu M, Wang J, Xu Y, Ding X, Li M, Jiang F, et al. Circulating tumor DNA is effective for the detection of EGFR mutation in non-small cell lung cancer: a meta-analysis. Cancer Epidemiol Biomark Prev. 2015;24:206–12.
Kim E, Feldman R, Wistuba II. Update on EGFR mutational testing and the potential of noninvasive liquid biopsy in non-small-cell lung cancer. Clin Lung Cancer. 2017. https://doi.org/10.1016/j.cllc.2017.08.001 (pii: S1525-7304(17)30229-2).
Singh AP, Li S, Cheng H. Circulating DNA in EGFR-mutated lung cancer. Ann Transl Med. 2017;5:379.
Duréndez-Sáez E, Azkárate A, Meri M, Calabuig-Fariñas S, Aguilar-Gallardo C, Blasco A, et al. New insights in non-small-cell lung cancer: circulating tumor cells and cell-free DNA. J Thorac Dis. 2017;9(Suppl 13):S1332-45.
Bai H, Mao L, Wang HS, Zhao J, Yang L, An TT, et al. Epidermal growth factor receptor mutations in plasma DNA samples predict tumor response in Chinese patients with stages IIIB to IV non-small-cell lung cancer. J Clin Oncol. 2009;27:2653–9.
Brevet M, Johnson ML, Azzoli CG, Ladanyi M. Detection of EGFR mutations in plasma DNA from lung cancer patients by mass spectrometry genotyping is predictive of tumor EGFR status and response to EGFR inhibitors. Lung Cancer. 2011;73:96–102.
Hu C, Liu X, Chen Y, Sun X, Gong Y, Geng M, et al. Direct serum and tissue assay for EGFR mutation in non-small cell lung cancer by high-resolution melting analysis. Oncol Rep. 2012;28:1815–21.
Kim HR, Lee SY, Hyun DS, Lee MK, Lee HK, Choi CM, et al. Detection of EGFR mutations in circulating free DNA by PNA‑mediated PCR clamping. J Exp Clin Cancer Res. 2013;32:50.
Karachaliou N, Mayo-de las Casas C, Queralt C, de Aguirre I, Melloni B, Cardenal F, et al. Association of EGFR L858R mutation in circulating free DNA with survival in the EURTAC trial. JAMA Oncol. 2015;1:149–57.
Kimura H, Suminoe M, Kasahara K, Sone T, Araya T, Tamori S, et al. Evaluation of epidermal growth factor receptor mutation status in serum DNA as a predictor of response to gefitinib (IRESSA). Br J Cancer 2007;97:778–84.
Goto K, Ichinose Y, Ohe Y, Yamamoto N, Negoro S, Nishio K, et al. Epidermal growth factor receptor mutation status in circulating free DNA in serum: From IPASS, a phase III study of gefitinib or carboplatin/paclitaxel in non‑small cell lung cancer. J Thorac Oncol 2012;7:115–21.
Douillard JY, Ostoros G, Cobo M, Ciuleanu T, Cole R, McWalter G, et al. Gefitinib treatment in EGFR mutated Caucasian NSCLC: Circulating‑free tumor DNA as a surrogate for determination of EGFR status. J Thorac Oncol 2014;9:1345–53.
Wu YL, Sequist LV, Hu CP, Feng J, Lu S, Huang Y, et al. EGFR mutation detection in circulating cell-free DNA of lung adenocarcinoma patients: analysis of LUX-Lung 3 and 6. Br J Cancer. 2017;116:175–85.
Weber B, Meldgaard P, Hager H, Wu L, Wei W, Tsai J, et al. Detection of EGFR mutations in plasma and biopsies from non-small cell lung cancer patients by allele-specific PCR assays. BMC Cancer. 2014;14:294.
Mok T, Wu YL, Lee JS, Yu CJ, Sriuranpong V, Sandoval-Tan J, et al. Detection and dynamic changes of EGFR mutations from circulating tumor DNA as a predictor of survival outcomes in NSCLC patients treated with first-line intercalated erlotinib and chemotherapy. Clin Cancer Res. 2015;21:3196–203.
Reckamp KL, Melnikova VO, Karlovich C, Sequist LV, Camidge DR, Wakelee H, et al. A highly sensitive and quantitative test platform for detection of NSCLC EGFR mutations in urine and plasma. J Thorac Oncol. 2016;11:1690–700.
US Food and Drug Administration. cobas EGFR Mutation Test v2. 2015. http://www.fda.gov/drugs/informationondrugs/approveddrugs/ucm504540.htm. Accessed 31 Jan 2018.
Yung TK, Chan KC, Mok TS, Tong J, To KF, Lo YM. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non-small cell lung cancer patients. Clin Cancer Res. 2009;15:2076–84.
Lee JY, Qing X, Xiumin W, Yali B, Chi S, Bak SH, et al. Longitudinal monitoring of EGFR mutations in plasma predicts outcomes of NSCLC patients treated with EGFR TKIs: Korean Lung Cancer Consortium (KLCC‑12‑02). Oncotarget 2016;7:6984–93.
Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O’Connell A, Feeney N, et al. Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2016;2:1014–22.
Couraud S, Vaca-Paniagua F, Villar S, Oliver J, Schuster T, Blanché H, et al. Noninvasive diagnosis of actionable mutations by deep sequencing of circulating free DNA in lung cancer from never‑smokers: a proof‑of‑concept study from BioCAST/IFCT‑1002. Clin Cancer Res 2014;20:4613–24.
Uchida J, Kato K, Kukita Y, Kumagai T, Nishino K, Daga H, et al. Diagnostic accuracy of noninvasive genotyping of EGFR in lung cancer patients by deep sequencing of plasma cell-Free DNA. Clin Chem. 2015;61:1191–6.
Paweletz CP, Sacher AG, Raymond CK, Alden RS, O’Connell A, Mach SL, et al. Bias-corrected targeted next-generation sequencing for rapid, multiplexed detection of actionable alterations in cell-free DNA from advanced lung cancer patients. Clin Cancer Res. 2016;22:915–22.
Thompson JC, Yee SS, Troxel AB, Savitch SL, Fan R, Balli D, et al. Detection of therapeutically targetable driver and resistance mutations in lung cancer patients by next-generation sequencing of cell-free circulating tumor DNA. Clin Cancer Res. 2016;22:5772–82.
Jenkins S, Yang JC, Ramalingam SS, Yu K, Patel S, Weston S, et al. Plasma ctDNA analysis for detection of the EGFR T790M mutation in patients with advanced non-small cell lung cancer. J Thorac Oncol. 2017;12:1061–70.
Takahama T, Sakai K, Takeda M, Azuma K, Hida T, Hirabayashi M, et al. Detection of the T790M mutation of EGFR in plasma of advanced non–small cell lung cancer patients with acquired resistance to tyrosine kinase inhibitors (West Japan oncology group 8014LTR study). Oncotarget. 2016;7:58492–9.
Mok TSK, Kim SW, Wu YL, Nakagawa K, Yang JJ, Ahn MJ, et al. Gefitinib plus chemotherapy versus chemotherapy in epidermal growth factor receptor mutation-positive non-small-cell lung cancer resistant to first-Line gefitinib (IMPRESS): overall survival and biomarker analyses. J Clin Oncol. 2017;35:4027–34.
Karlovich C, Goldman JW, Sun JM, Mann E, Sequist LV, Konopa K, et al. Assessment of EGFR mutation status in matched plasma and tumor tissue of NSCLC patients from a phase I study of rociletinib (CO-1686). Clin Cancer Res. 2016;22:2386–95.
Oxnard GR, Thress KS, Alden RS, Lawrance R, Paweletz CP, Cantarini M, et al. Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non-small-cell lung cancer. J Clin Oncol. 2016;34:3375–82.
Wu YL, Zhou C, Liam CK, Wu G, Liu X, Zhong Z, et al. First-line erlotinib versus gemcitabine/cisplatin in patients with advanced EGFR mutation-positive non-small-cell lung cancer: analyses from the phase III, randomized, open-label, ENSURE study. Ann Oncol. 2015;26:1883–9.
Oztan A, Fischer S, Schrock AB, Erlich RL, Lovly CM, Stephens PJ, et al. Emergence of EGFR G724S mutation in EGFR-mutant lung adenocarcinoma post progression on osimertinib. Lung Cancer. 2017;111:84–7.
Kobayashi S, Boggon TJ, Dayaram T, Jänne PA, Kocher O, Meyerson M, et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med. 2005;352:786–92.
Sequist LV, Waltman BA, Dias-Santagata D, Digumarthy S, Turke AB, Fidias P, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med. 2011;3:75ra26.
Ohashi K, Maruvka YE, Michor F, Pao W. Epidermal growth factor receptor tyrosine kinase inhibitor-resistant disease. J Clin Oncol. 2013;31:1070–80.
Ramalingam SS, Yang JC, Lee CK, Kurata T, Kim DW, John T, et al. Osimertinib as first-line treatment of EGFR mutation-positive advanced non-small-cell lung cancer. J Clin Oncol 2017;25:JCO2017747576. https://doi.org/10.1200/JCO.2017.74.7576. (Epub ahead of print).
Goss G, Tsai CM, Shepherd FA, Bazhenova L, Lee JS, Chang GC, et al. Osimertinib for pretreated EGFR Thr790Met-positive advanced non-small-cell lung cancer (AURA2): a multicentre, open-label, single-arm, phase 2 study. Lancet Oncol. 2016;17:1643–52.
Mok TS, Wu Y-L, Ahn M-J, Garassino MC, Kim HR, Ramalingam SS, et al. Osimertinib or platinum-pemetrexed in EGFR T790M-positive lung cancer. N Engl J Med. 2017;376:629–40.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Kazue Yoneda has no conflicts of interest; Naoko Imanishi has no conflicts of interest; Yoshinobu Ichiki has no conflicts of interest; Fumihiro Tanaka received a research grant from Astra Zeneca, Chugai Pharmaceutical, Taiho Pharmaceutical and Ono Pharmaceutical and received honoraria from Astra Zeneca, Chugai Pharmaceutical and Taiho Pharmaceutical.
Rights and permissions
About this article
Cite this article
Yoneda, K., Imanishi, N., Ichiki, Y. et al. A liquid biopsy in primary lung cancer. Surg Today 49, 1–14 (2019). https://doi.org/10.1007/s00595-018-1659-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00595-018-1659-2