Abstract
Aims
Recent years have witnessed an increasing research interest in the roles of transcription factor (TF)-gene regulatory network in type 2 diabetes mellitus (T2DM). Thus, we sought to characterize the mechanistic insights based on the TF-gene regulatory network in skeletal muscle atrophy in T2DM.
Methods
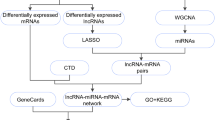
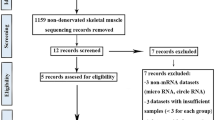
Differentially expressed TFs (DETFs) and mRNAs (DEmRNAs) were obtained in T2DM-related gene expression profiles (GSE12643, GSE55650, GSE166502, and GSE29221), followed by WGCNA, and GO and KEGG enrichment analyses. Next, the iRegulon plug-in unit of Cytoscape software was used to construct a TF-mRNA regulatory network. Besides, RT-qPCR and ChIP-seq were utilized to measure the expression of CEBPA and FGF21 in the skeletal muscle tissues or cells of T2DM rat models. At last, the effect of overexpression of FGF21 on the autophagy-lysosomal pathway was examined in skeletal muscle cells of T2DM rats.
Results
Totally, 12 DETFs and 102 DEmRNAs were found in the skeletal muscle tissues of T2DM samples. The DEmRNAs were mainly enriched in the autophagy-lysosomal pathway. CEBPA affected the skeletal muscle atrophy in T2DM by regulating 5 target genes via the autophagy-lysosomal pathway. CEBPA could target FGF21. In addition, the expression of CEBPA was elevated, while the expression of FGF21 was diminished in the skeletal muscle tissues or cells of T2DM rats. The CEBPA-FGF21 regulatory network promoted skeletal muscle atrophy in T2DM by activating the autophagy-lysosomal pathway.
Conclusion
The CEBPA-FGF21 regulatory network may participate in the T2DM-induced skeletal muscle atrophy by regulating the autophagy-lysosomal pathway. Thus, our study provides interesting targets for prevention of skeletal muscle atrophy in T2DM.







Similar content being viewed by others
References
Rachdaoui N (2020) Insulin: the friend and the foe in the development of type 2 diabetes mellitus. Int J Mol Sci 21(5):1770. https://doi.org/10.3390/ijms21051770
Farup J, Just J, de Paoli F, et al (2021) Human skeletal muscle CD90(+) fibro-adipogenic progenitors are associated with muscle degeneration in type 2 diabetic patients. Cell Metab 33(11):2201–2214 e2211. https://doi.org/10.1016/j.cmet.2021.10.001
Ato S, Kido K, Sato K, Fujita S (2019) Type 2 diabetes causes skeletal muscle atrophy but does not impair resistance training-mediated myonuclear accretion and muscle mass gain in rats. Exp Physiol 104(10):1518–1531. https://doi.org/10.1113/EP087585
Lee H, Song W (2018) Exercise and mitochondrial remodeling in skeletal muscle in type 2 diabetes. J Obes Metab Syndr 27(3):150–157. https://doi.org/10.7570/jomes.2018.27.3.150
Reddy SS, Shruthi K, Joy D, Reddy GB (2019) 4-PBA prevents diabetic muscle atrophy in rats by modulating ER stress response and ubiquitin-proteasome system. Chem Biol Interact 306:70–77. https://doi.org/10.1016/j.cbi.2019.04.009
Boada M, Catalan AI, Ottati C,et al (2021) Germline CEBPA mutation in familial acute myeloid leukemia. Hematol Rep 13(3):9114. https://doi.org/10.4081/hr.2021.9114
Wilhelmson AS, Porse BT (2020) CCAAT enhancer binding protein alpha (CEBPA) biallelic acute myeloid leukaemia: cooperating lesions, molecular mechanisms and clinical relevance. Br J Haematol 190(4):495–507. https://doi.org/10.1111/bjh.16534
Nurminen V, Neme A, Seuter S, Carlberg C (2019) Modulation of vitamin D signaling by the pioneer factor CEBPA. Biochim Biophys Acta Gene Regul Mech 1862(1):96–106. https://doi.org/10.1016/j.bbagrm.2018.12.004
Wicks K, Torbica T, Umehara T, Amin S, Bobola N, Mace KA (2015) Diabetes inhibits Gr-1+ myeloid cell maturation via Cebpa deregulation. Diabetes 64(12):4184–4197. https://doi.org/10.2337/db14-1895
Geng L, Lam KSL, Xu A (2020) The therapeutic potential of FGF21 in metabolic diseases: from bench to clinic. Nat Rev Endocrinol 16(11):654–667. https://doi.org/10.1038/s41574-020-0386-0
Yuan D, Wu BJ, Henry A, Rye KA, Ong KL (2019) Role of fibroblast growth factor 21 in gestational diabetes mellitus: a mini-review. Clin Endocrinol (Oxf) 90(1):47–55. https://doi.org/10.1111/cen.13881
Gasser E, Moutos CP, Downes M, Evans RM (2017) FGF1 - a new weapon to control type 2 diabetes mellitus. Nat Rev Endocrinol 13(10):599–609. https://doi.org/10.1038/nrendo.2017.78
Liu KP, Zhou D, Ouyang DY, et al (2013) LC3B-II deacetylation by histone deacetylase 6 is involved in serum-starvation-induced autophagic degradation. Biochem Biophys Res Commun 441(4):970–975. https://doi.org/10.1016/j.bbrc.2013.11.007
Saito S, Zhuang Y, Shan B, et al (2017) Tubastatin ameliorates pulmonary fibrosis by targeting the TGFbeta-PI3K-Akt pathway. PLoS ONE 12(10):e0186615. https://doi.org/10.1371/journal.pone.0186615
Stana F, Vujovic M, Mayaki D, et al (2017) Differential regulation of the autophagy and proteasome pathways in skeletal muscles in sepsis. Crit Care Med 45(9):e971–e979. https://doi.org/10.1097/CCM.0000000000002520
Kim KW, Baek MO, Choi JY, Son KH, Yoon MS (2019) Analysis of the molecular signaling signatures of muscle protein wasting between the intercostal muscles and the gastrocnemius muscles in db/db mice. Int J Mol Sci 20(23):6062. https://doi.org/10.3390/ijms20236062
Giron MD, Vilchez JD, Shreeram S, et al (2015) beta-Hydroxy-beta-methylbutyrate (HMB) normalizes dexamethasone-induced autophagy-lysosomal pathway in skeletal muscle. PLoS ONE 10(2):e0117520. https://doi.org/10.1371/journal.pone.0117520
Hirunsai M, Srikuea R (2021) Autophagy-lysosomal signaling responses to heat stress in tenotomy-induced rat skeletal muscle atrophy. Life Sci 275:119352. https://doi.org/10.1016/j.lfs.2021.119352
Lu Y, Li Y, Li G, Lu H (2020) Identification of potential markers for type 2 diabetes mellitus via bioinformatics analysis. Mol Med Rep 22(3):1868–1882. https://doi.org/10.3892/mmr.2020.11281
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinf 9:559. https://doi.org/10.1186/1471-2105-9-559
Ma X, Su J, Wang B, Jin X (2022) Identification of characteristic genes in whole blood of intervertebral disc degeneration patients by weighted gene coexpression network analysis (WGCNA). Comput Math Methods Med 2022:6609901. https://doi.org/10.1155/2022/6609901
Janky R, Verfaillie A, Imrichova H, et al (2014) iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLoS Comput Biol 10(7):e1003731. https://doi.org/10.1371/journal.pcbi.1003731
Mojani MS, Sarmadi VH, Vellasamy S, et al (2014) Evaluation of metabolic and immunological changes in streptozotocin-nicotinamide induced diabetic rats. Cell Immunol 289(1–2):145–149. https://doi.org/10.1016/j.cellimm.2014.04.004
Ortiz Mdel C, Lores-Arnaiz S, Albertoni Borghese MF, et al (2013) Mitochondrial dysfunction in brain cortex mitochondria of STZ-diabetic rats: effect of l-Arginine. Neurochem Res 38(12):2570–2580. https://doi.org/10.1007/s11064-013-1172-3
Hughes DC, Turner DC, Baehr LM et al (2021) Knockdown of the E3 ubiquitin ligase UBR5 and its role in skeletal muscle anabolism. Am J Physiol Cell Physiol 320(1):C45–C56. https://doi.org/10.1152/ajpcell.00432.2020
Hughes DC, Baehr LM, Driscoll JR, Lynch SA, Waddell DS, Bodine SC (2020) Identification and characterization of Fbxl22, a novel skeletal muscle atrophy-promoting E3 ubiquitin ligase. Am J Physiol Cell Physiol 319(4):C700–C719. https://doi.org/10.1152/ajpcell.00253.2020
Queiroz AL, Lessard SJ, Ouchida AT, et al (2021) The MicroRNA miR-696 is regulated by SNARK and reduces mitochondrial activity in mouse skeletal muscle through Pgc1alpha inhibition. Mol Metab 51:101226. https://doi.org/10.1016/j.molmet.2021.101226
Yu G, Wang LG, He QY (2015) ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31(14):2382–2383. https://doi.org/10.1093/bioinformatics/btv145
Ajawatanawong P, Atkinson GC, Watson-Haigh NS, Mackenzie B, Baldauf SL (2012) SeqFIRE: a web application for automated extraction of indel regions and conserved blocks from protein multiple sequence alignments. Nucleic Acids Res 40(Web Server issue):W340–347. https://doi.org/10.1093/nar/gks561
Liu Z, Sun J, Li C, Xu L, Liu J (2021) MKL1 regulates hepatocellular carcinoma cell proliferation, migration and apoptosis via the COMPASS complex and NF-kappaB signaling. BMC Cancer 21(1):1184. https://doi.org/10.1186/s12885-021-08185-w
Ying L, Wang L, Guo K, et al (2021) Paracrine FGFs target skeletal muscle to exert potent anti-hyperglycemic effects. Nat Commun 12(1):7256. https://doi.org/10.1038/s41467-021-27584-y
Liu WJ, Ye L, Huang WF, et al (2016) p62 links the autophagy pathway and the ubiqutin-proteasome system upon ubiquitinated protein degradation. Cell Mol Biol Lett 21:29. https://doi.org/10.1186/s11658-016-0031-z
Mu J, Zhang D, Tian Y, Xie Z, Zou MH (2020) BRD4 inhibition by JQ1 prevents high-fat diet-induced diabetic cardiomyopathy by activating PINK1/Parkin-mediated mitophagy in vivo. J Mol Cell Cardiol 149:1–14. https://doi.org/10.1016/j.yjmcc.2020.09.003
Cividini F, Scott BT, Suarez J, et al (2021) Ncor2/PPARalpha-dependent upregulation of MCUb in the type 2 diabetic heart impacts cardiac metabolic flexibility and function. Diabetes 70(3):665–679. https://doi.org/10.2337/db20-0779
Ming Y, Yin Y, Sun Z (2020) Interaction of Nuclear Receptor Subfamily 4 Group A Member 1 (Nr4a1) and Liver Linase B1 (LKB1) Mitigates Type 2 Diabetes Mellitus by Activating Monophosphate-Activated Protein Kinase (AMPK)/Sirtuin 1 (SIRT1) Axis and Inhibiting Nuclear Factor-kappa B (NF-kappaB) Activation. Med Sci Monit 26:e920278. https://doi.org/10.12659/MSM.920278
Duesing K, Charpentier G, Marre M, et al (2008) Evaluating the association of common PBX1 variants with type 2 diabetes. BMC Med Genet 9:14. https://doi.org/10.1186/1471-2350-9-14
Liu W, Meridew JA, Aravamudhan A, et al (2019) Targeted regulation of fibroblast state by CRISPR-mediated CEBPA expression. Respir Res 20(1):281. https://doi.org/10.1186/s12931-019-1253-1
Zhu Y, Ding X, She Z, et al (2020) Exploring shared pathogenesis of alzheimer’s disease and type 2 diabetes mellitus via co-expression networks analysis. Curr Alzheimer Res 17(6):566–575. https://doi.org/10.2174/1567205017666200810164932
Travers ME, Mackay DJ, Dekker Nitert M, et al (2013) Insights into the molecular mechanism for type 2 diabetes susceptibility at the KCNQ1 locus from temporal changes in imprinting status in human islets. Diabetes 62(3):987–992. https://doi.org/10.2337/db12-0819
Stoynev N, Dimova I, Rukova B, Hadjidekova S, Nikolova D, Toncheva D, Tankova T (2014) Gene expression in peripheral blood of patients with hypertension and patients with type 2 diabetes. J Cardiovasc Med (Hagerstown) 15(9):702–709. https://doi.org/10.2459/JCM.0b013e32835dbcc8
Jimenez V, Jambrina C, Casana E, et al (2018) FGF21 gene therapy as treatment for obesity and insulin resistance. EMBO Mol Med 10(8):e8791. https://doi.org/10.15252/emmm.201708791
Lu W, Li X, Luo Y (2021) FGF21 in obesity and cancer: new insights. Cancer Lett 499:5–13. https://doi.org/10.1016/j.canlet.2020.11.026
Guo C, Zhao L, Li Y, Deng X, Yuan G (2021) Relationship between FGF21 and drug or nondrug therapy of type 2 diabetes mellitus. J Cell Physiol 236(1):55–67. https://doi.org/10.1002/jcp.29879
Fangmann D, Geisler C, Schlicht K, et al (2021) Differential effects of protein intake versus intake of a defined oligopeptide on FGF-21 in obese human subjects in vivo. Clin Nutr 40(2):600–607. https://doi.org/10.1016/j.clnu.2020.06.006
Kharitonenkov A, Shiyanova TL, Koester A, et al(2005) FGF-21 as a novel metabolic regulator. J Clin Invest 115(6):1627–1635. https://doi.org/10.1172/JCI23606
Yun HR, Jo YH, Kim J, Shin Y, Kim SS, Choi TG (2020) Roles of Autophagy in Oxidative Stress. Int J Mol Sci 21(9):3289. https://doi.org/10.3390/ijms21093289
Henriksen TI, Wigge LV, Nielsen J, Pedersen BK, Sandri M, Scheele C (2019) Dysregulated autophagy in muscle precursor cells from humans with type 2 diabetes. Sci Rep 9(1):8169. https://doi.org/10.1038/s41598-019-44535-2
Perry BD, Caldow MK, Brennan-Speranza TC, et al (2016) Muscle atrophy in patients with Type 2 Diabetes Mellitus: roles of inflammatory pathways, physical activity and exercise. Exerc Immunol Rev 22:94–109
Huang X, Kuang S, Shen Z, Liang M, Lin Z (2020) High glucose disrupts autophagy lysosomal pathway in gingival epithelial cells via ATP6V0C. J Periodontol 91(5):705–714. https://doi.org/10.1002/JPER.19-0262
Funding
This study was funded by the National Natural Science Foundation of China (81702241), the Natural Science Foundation of Hunan Province (2018JJ3847), China Postdoctoral Science Foundation (2020M682597) and the Natural Science Foundation of Hunan Province (2021JJ31129).
Author information
Authors and Affiliations
Contributions
KW and SH designed the study. FZ and YL collated the data, designed and developed the database, carried out data analyses and produced the initial draft of the manuscript. KW and SH contributed to drafting the manuscript. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Ethics statement
This study protocol was approved by the Experimental Animal Ethics Committee of Xiangya Hospital of Central South University.
Informed consent
Clinical data was primarily obtained from public databases.
Additional information
Managed By Fabrizio Barbetti.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, K., Huang, S., Zheng, F. et al. The CEBPA-FGF21 regulatory network may participate in the T2DM-induced skeletal muscle atrophy by regulating the autophagy-lysosomal pathway. Acta Diabetol 60, 1491–1503 (2023). https://doi.org/10.1007/s00592-023-02131-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00592-023-02131-x