Abstract
Background
To establish a treatment option for liver fibrosis, the possibility of the drug repurposing theory was investigated, with a focus on the off-target effects of active pharmaceutical ingredients.
Methods
First, several active pharmaceutical ingredients were screened for their effects on the gene expression in the hepatocytes using chimeric mice with humanized hepatocytes. As per the gene expression-based screening assay for 36 medications, we assessed the mechanism of the antifibrotic effect of letrozole, a third-generation aromatase inhibitor, in mouse models of liver fibrosis induced by carbon tetrachloride (CCl4) and a methionine choline-deficient (MCD) diet. We assessed liver histology, serum biochemical markers, and fibrosis-related gene and protein expressions in the hepatocytes.
Results
A gene expression-based screening assay revealed that letrozole had a modifying effect on fibrosis-related gene expression in the hepatocytes, including YAP, CTGF, TGF-β, and CYP26A1. Letrozole was administered to mouse models of CCl4- and MCD-induced liver fibrosis and it ameliorated the liver fibrosis. The mechanisms involved the inhibition of the Yap-Ctgf profibrotic pathway following a decrease in retinoic acid levels in the hepatocytes caused by suppression of the hepatic retinol dehydrogenase, Hsd17b13 and activation of the retinoic acid hydrogenase, Cyp26a1.
Conclusions
Letrozole slowed the progression of liver fibrosis by inhibiting the Yap-Ctgf pathway. The mechanisms involved the modification of the Hsd17b13 and Cyp26a1 expressions led to the suppression of retinoic acid in the hepatocytes, which contributed to the activation of Yap-Ctgf pathway. Because of its off-target effect, letrozole could be repurposed for the treatment of liver fibrosis.
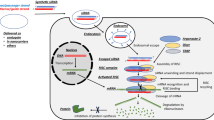
Graphical abstract

The third-generation aromatase inhibitor letrozole ameliorated liver fibrosis by suppressing the Yap-Ctgf pathway by partially modifying the Hsd17b13 and Cyp26a1 expressions, which reduced the retinoic acid level in the hepatocytes. The gene expression analysis using chimeric mice with humanized liver revealed that the mechanisms are letrozole specific and, therefore, may be repurposed for the treatment of liver fibrosis.







Similar content being viewed by others
Abbreviations
- ALT:
-
Alanine aminotransferase
- Acta2:
-
Actin alpha 2, α-smooth muscle actin
- AI:
-
Aromatase inhibitor
- AST:
-
Aspartate aminotransferase
- BW:
-
Body weight
- CCl4:
-
Carbon tetrachloride
- COL1A1:
-
Collagen type I alpha 1
- CTGF:
-
Connective tissue growth factor
- CYP26A1:
-
Cytochrome P450 26A1
- H&E:
-
Hematoxylin and eosin
- HSC:
-
Hepatic stellate cell
- HSD17B13:
-
17β-Hydroxysteroid dehydrogenase 13
- NAFLD:
-
Nonalcoholic fatty liver disease
- NASH:
-
Nonalcoholic steatohepatitis
- LW:
-
Liver weight
- M2BP:
-
Mac-2 binding protein
- MMP:
-
Matrix metalloproteinase
- MCD:
-
Methionine choline-deficient
- NS:
-
No statistical significance
- RA:
-
Retinoic acid
- RDH:
-
Retinol dehydrogenase
- SERM:
-
Selective estrogen receptor modulator
- SCFA:
-
Short-chain fatty acid
- T-Bil:
-
Total bilirubin
- TC:
-
Total cholesterol
- TGF-β:
-
Transforming growth factor-β
- TG:
-
Triglycerides
- YAP:
-
Yes-associated protein
References
Ibrahim SH, Hirsova P, Gores GJ. Non-alcoholic steatohepatitis pathogenesis: sublethal hepatocyte injury as a driver of liver inflammation. Gut. 2018;67:963–72.
Eslam M, Newsome PN, Sarin SK, et al. A new definition for metabolic dysfunction-associated fatty liver disease: an international expert consensus statement. J Hepatol. 2020;73:202–9.
Sumida Y, Yoneda M. Current and future pharmacological therapies for NAFLD/NASH. J Gastroenterol. 2018;53:362–76.
Angulo P, Kleiner DE, Dam-Larsen S, et al. Liver Fibrosis, but no other histologic features, is associated with long-term outcomes of patients with nonalcoholic fatty liver disease. Gastroenterology. 2015;149:389-97.e10.
Ramachandran P, Dobie R, Wilson-Kanamori JR, et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature. 2019;575:512–8.
Seki E, Brenner DA. Recent advancement of molecular mechanisms of liver fibrosis. J Hepatobil Pancreat Sci. 2015;22:512–8.
Gressner OA, Lahme B, Demirci I, et al. Differential effects of TGF-beta on connective tissue growth factor (CTGF/CCN2) expression in hepatic stellate cells and hepatocytes. J Hepatol. 2007;47:699–710.
Abe H, Kamimura K, Kobayashi Y, et al. Effective prevention of liver fibrosis by liver-targeted hydrodynamic gene delivery of matrix metalloproteinase-13 in a rat liver fibrosis model. Mol Ther Nucleic Acids. 2016;5: e276.
Napolitano F, Zhao Y, Moreira VM, et al. Drug repositioning: a machine-learning approach through data integration. J Cheminform. 2013;5:30.
Nakashima M, Tachiki H, Enomoto H, et al. Changes in hepatic gene expression induced by various statin formulations in chimeric PXB-mouse with highly humanized liver. Toxicol Lett. 2013;221:S194.
Nakashima M, Enomoto H, Tachiki H, et al. Effects of irinotecan on hepatic gene expression in chimeric PXB-mouse® with highly humanized liver. Toxicol Lett. 2014;229:S235.
Sanoh S, Horiguchi A, Sugihara K, et al. Prediction of in vivo hepatic clearance and half-life of drug candidates in human using chimeric mice with humanized liver. Drug Metab Dispos. 2012;40:322–8.
Sakuma T, Kawasaki Y, Jarukamjorn K, et al. Sex differences of drug-metabolizing enzyme: Female predominant expression of human and mouse cytochrome P450 3A isoforms. J Health Sci. 2009;55:325–37.
Yamada T, Obata A, Kashiwagi Y, et al. Gd-EOB-DTPA-enhanced-MR imaging in the inflammation stage of nonalcoholic steatohepatitis (NASH) in mice. Magn Res Imaging. 2016;34:724–9.
Brigstock DR. Strategies for blocking the fibrogenic actions of connective tissue growth factor (CCN2): From pharmacological inhibition in vitro to targeted siRNA therapy in vivo. J Cell Commun Signal. 2009;3:5–18.
Vrekoussis T, Chaniotis V, Navrozoglou I, et al. Image analysis of breast cancer immunohistochemistry-stained sections using ImageJ: an RGB-based model. Anticancer Res. 2009;29:4995–8.
Iwata A, Kamada Y, Ebisutani Y, et al. Establishment of mouse Mac-2 binding protein enzyme-linked immunosorbent assay and its application for mouse chronic liver disease models. Hepatol Res. 2016;47:902–9.
Sierra-Filardi E, Nieto C, Domínguez-Soto A, et al. CCL2 shapes macrophage polarization by GM-CSF and M-CSF: identification of CCL2/CCR2-dependent gene expression profile. J Immunol. 2014;192:3858–67.
Lan T, Li C, Yang G, et al. Sphingosine kinase 1 promotes liver fibrosis by preventing miR-19b-3p-mediated inhibition of CCR2. Hepatology. 2018;68:1070–86.
Marra F, Tacke F. Roles for chemokines in liver disease. Gastroenterology. 2014;147:577–94.
Hintermann E, Bayer M, Pfeilschifter JM, et al. CXCL10 promotes liver fibrosis by prevention of NK cell mediated hepatic stellate cell inactivation. J Autoimmun. 2010;35:424–35.
Schmidt-Arras D, Rose-John S. IL-6 pathway in the liver: From physiopathology to therapy. J Hepatol. 2016;64:1403–15.
Rong X, Liu J, Yao X, et al. Human bone marrow mesenchymal stem cells-derived exosomes alleviate liver fibrosis through the Wnt/β-catenin pathway. Stem Cell Res Ther. 2019;10:98.
Mezquita B, Mezquita P, Pau M, et al. All-trans-retinoic acid activates the pro-invasive Src-YAP-interleukin 6 axis in triple-negative MDA-MB-231 breast cancer cells while cerivastatin reverses this action. Sci Rep. 2018;8:7047.
Ma Y, Belyaeva OV, Brown PM, et al. 17-Beta hydroxysteroid dehydrogenase 13 is a hepatic retinol dehydrogenase associated with histologic features of nonalcoholic fatty liver disease. Hepatology. 2019;69:1504–19.
Su W, Mao Z, Liu Y, et al. Role of HSD17B13 in the liver physiology and pathophysiology. Mol Cell Endocrinol. 2019;489:119–25.
Sookoian S, Pirola CJ, Valenti L, et al. Genetic pathways in nonalcoholic fatty liver disease: insights from systems biology. Hepatology. 2020;72:330–46.
Stender S, Romeo S. HSD17B13 as a promising therapeutic target against chronic liver disease. Liver Int. 2020;40:756–7.
Yang W, Han W, Qin A, et al. The emerging role of Hippo signaling pathway in regulating osteoclast formation. J Cell Physiol. 2018;233:4606–17.
Lipson KE, Wong C, Teng Y, et al. CTGF is a central mediator of tissue remodeling and fibrosis and its inhibition can reverse the process of fibrosis. Fibrogenesis Tissue Rep. 2012;5:S24.
Makino Y, Hikita H, Kodama T, et al. CTGF mediates tumor-stroma interactions between hepatoma cells and hepatic stellate cells to accelerate HCC progression. Cancer Res. 2018;78:4902–14.
Napoli JL. Physiological insights into all-trans-retinoic acid biosynthesis. Biochim Biophys Acta. 2012;1821:152–67.
Qin XY, Suzuki H, Honda M, et al. Prevention of hepatocellular carcinoma by targeting MYCN-positive liver cancer stem cells with acyclic retinoid. Proc Natl Acad Sci U S A. 2018;115:4969–74.
Shimizu H, Tsubota T, Kanki K, et al. All-trans retinoic acid ameliorates hepatic stellate cell activation via suppression of thioredoxin interacting protein expression. J Cell Physiol. 2018;233:607–16.
Lee KA, Song YC, Kim GY, et al. Retinoic acid alleviates Con A-induced hepatitis and differentially regulates effector production in NKT cells. Eur J Immunol. 2012;42:1685–94.
Su W, Wang Y, Jia X, et al. Comparative proteomic study reveals 17β-HSD13 as a pathogenic protein in nonalcoholic fatty liver disease. Proc Natl Acad Sci U S A. 2014;111:11437–42.
Saeed A, Bartuzi P, Heegsma J, et al. Impaired hepatic vitamin a metabolism in NAFLD mice leading to vitamin a accumulation in hepatocytes. Cell Mol Gastroenterol Hepatol. 2021;11:309-325.e3.
Verma R, Krishna A. Effect of letrozole, a selective aromatase inhibitor, on testicular activities in adult mice: both in vivo and in vitro study. Gen Comp Endocrinol. 2017;241:57–68.
Poutanen M, Penning TM. Biology and clinical relevance of hydroxysteroid (17β) dehydrogenase enzymes. Mol Cell Endocrinol. 2019;489:1–2.
Sakuma T, Takai M, Endo Y, et al. A novel female-specific member of the CYP3A gene subfamily in the mouse liver. Arch Biochem Biophys. 2000;377:153–62.
Sakuma T, Endo Y, Mashino M, et al. Regulation of the expression of two female-predominant CYP3A mRNAs (CYP3A41 and CYP3A44) in mouse liver by sex and growth hormones. Arch Biochem Biophys. 2002;404:234–42.
Delvoux B, D’Hooghe T, Kyama C, et al. Inhibition of type 1 17β-hydroxysteroid dehydrogenase impairs the synthesis of 17β-estradiol in endometriosis lesions. J Clin Endocrinol Metab. 2014;99:276–84.
Wang XQ, Aka JA, Li T, et al. Inhibition of 17β-hydroxysteroid dehydrogenase type 7 modulates breast cancer protein profile and enhances apoptosis by down-regulating GRP78. J Steroid Biochem Mol Biol. 2017;172:188–97.
Abul-Husn NS, Cheng X, Li AH, et al. A protein-truncating HSD17B13 variant and protection from chronic liver disease. N Engl J Med. 2018;378:1096–106.
Carlsson B, Lindén D, Brolén G, et al. Review article: the emerging role of genetics in precision medicine for patients with non-alcoholic steatohepatitis. Aliment Pharmacol Ther. 2020;51:1305–20.
Su W, Peng J, Li S, et al. Liver X receptor alpha induces 17β-hydroxysteroid dehydrogenase-13 expression through SREBP-1c. Am J Physiol Endocrinol Metab. 2017;312:E357–67.
Gellert-Kristensen H, Nordestgaard BG, Tybjaerg-Hansen A, et al. High risk of fatty liver disease amplifies the alanine transaminase-lowering effect of a HSD17B13 variant. Hepatology. 2020;71:56–66.
Luukkonen PK, Tukiainen T, Juuti A, et al. Hydroxysteroid 17-beta dehydrogenase 13 variant increases phospholipids and protects against fibrosis in nonalcoholic fatty liver disease. JCI Insight. 2020;5: e132158.
Isoherranen N, Zhong G. Biochemical and physiological importance of the CYP26 retinoic acid hydroxylases. Pharmacol Ther. 2019;204: 107400.
Wang M, Li J, Li H, et al. Down-regulating the high level of 17-beta-hydroxysteroid dehydrogenase 13 plays a therapeutic role for non-alcoholic fatty liver disease. Int J Mol Sci. 2022;23:5544.
Ma Y, Brown PM, Lin DD, et al. Hsd17b13 deficiency does not protect mice from obesogenic diet injury. Hepatology. 2021;73:1701–16.
Wolbold R, Klein K, Burk O, et al. Sex is a major determinant of CYP3A4 expression in human liver. Hepatology. 2003;38:978–88.
Anstee QM, Darlay R, Cockell S, et al. Genome-wide association study of non-alcoholic fatty liver and steatohepatitis in a histologically characterised cohort. J Hepatol. 2020;73:505–15.
Eslam M, George J. Genetic contributions to NAFLD: leveraging shared genetics to uncover systems biology. Nat Rev Gastroenterol Hepatol. 2020;17:40–52.
Acknowledgements
The authors would like to thank Takao Tsuchida in the Division of Gastroenterology and Hepatology at the Niigata University for his excellent assistance in histological analyses. The authors would also like to thank Nobuyoshi Fujisawa, Yoshitaka Maeda, Kanako Oda, Shuko Adachi, Toshikuni Sasaoka, and all staff members at the Division of Laboratory Animal Resources in Niigata University. They also thank Enago for the critical reading of the manuscript and English language review. The authors declare that they have no conflict of interest.
Funding
Kamimura K, Sakai N, and Terai S of Niigata University received research funds from Towa Pharmaceutical Co., Ltd., and Nakashima M, Ohyama K, Miyamoto H, Inamine T, and Tokunaga A of Nagasaki University received research funds from Towa Pharmaceutical Co., Ltd.
Author information
Authors and Affiliations
Contributions
NS, KK, HM, and KO contributed to the study conception and design. Material preparation, data collection, and analysis were performed by NS, KK, HM, MK, TN, YN, TS, AS, TY, HK, HS, AT, TI, MN, HE, KK, and HT. The first draft of the manuscript was written by NS, KK, TI, KK, KO, and ST. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
535_2022_1929_MOESM2_ESM.tif
Supplementary file2 (TIF 2084 KB) Figure S1. The heat map and gene expression list changed significantly with LET administration. Six chimeric mice with humanized liver were used to test all 36 medicines
535_2022_1929_MOESM3_ESM.tif
Supplementary file3 (TIF 811 KB) Figure S2. (A), (B). Effect of LET on the hepatic steatosis of CCl4- and MCD-treated mice. The scale bar represents 100 µm. (C), (D). The relative gene expressions of Hsd17b13 and Cyp26a1 in the female mice treated with CCl4 and MCD. The values represent mean ± SD (n=5 for each group), * p <0.05, One-way ANOVA followed by Bonferroni’s multiple comparison test
535_2022_1929_MOESM4_ESM.tif
Supplementary file4 (TIF 1872 KB) Figure S3. The heat map and list of chemokine and cytokine gene expressions after 36 medicines were tested on six chimeric mice
535_2022_1929_MOESM5_ESM.tif
Supplementary file5 (TIF 4428 KB) Figure S4. The heat map and list of oxidative stress-related gene expressions after 36 medicines were tested on six chimeric mice
535_2022_1929_MOESM6_ESM.tif
Supplementary file6 (TIF 207 KB) Figure S5. Correlation between liver fibrosis and serum RA, and ATRA in the liver tissue. (A). The relationship between liver fibrosis (%) and serum RA. (B). The relationship between liver fibrosis (%) and ATRA in the liver tissue of the CCl4 model mice. The black circles indicate data from animals. The bold black line shows the trend line, and correlation analyses were performed. ** p <0.01, *** p <0.001, and NS, no statistical significance. r, correlation coefficient
535_2022_1929_MOESM7_ESM.tif
Supplementary file7 (TIF 469 KB) Figure S6. Effect of LET on estradiol in female mice. (A). Urine estradiol. (B). Urine creatinine. (C) Creatinine adjusted urinary estradiol. The values represent mean ± SD (n=5 for each group), N.S., not significant, the Student's t-test was used to evaluate the differences in gene expression between the control and drug-treated groups
535_2022_1929_MOESM8_ESM.tif
Supplementary file8 (TIF 659 KB) Figure S7. The dose-dependent effect of LET on gene expression in PXB cells. Relative gene expressions of CTGF, YAP, ACTA2, TGF-β, and HSD17B13 in PXB cells treated with different doses of LET. The values represent mean ± SD (n=4 for each group), * p <0.05, ** p <0.01. The Student's t-test was used to evaluate the differences in gene expression between the control and drug-treated groups
535_2022_1929_MOESM9_ESM.tif
Supplementary file9 (TIF 385 KB) Figure S8. Effect of LET on gene expression in human hepatic stellate cells. Relative gene expressions of CTGF, YAP, ACTA2, TGF-β, and HSD17B13 in PXB cells treated with LET (1 μg/ml). The values represent mean ± SD (n=8 for each group), NS Not statistically significant. The Student's t-test was used to evaluate the differences in gene expression between the control and drug-treated groups
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sakai, N., Kamimura, K., Miyamoto, H. et al. Letrozole ameliorates liver fibrosis through the inhibition of the CTGF pathway and 17β-hydroxysteroid dehydrogenase 13 expression. J Gastroenterol 58, 53–68 (2023). https://doi.org/10.1007/s00535-022-01929-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-022-01929-w