Abstract
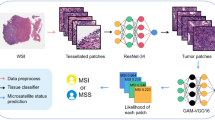
Cancers have emerged as a significant concern due to their impact on public health and society. The examination and interpretation of tissue sections stained with Hematoxylin and Eosin (H&E) play a crucial role in disease assessment, particularly in cases like gastric cancer. Microsatellite instability (MSI) is suggested to contribute to the carcinogenesis of specific gastrointestinal tumors. However, due to the nonspecific morphology observed in H&E-stained tissue sections, MSI determination often requires costly evaluations through various molecular studies and immunohistochemistry methods in specialized molecular pathology laboratories. Despite the high cost, international guidelines recommend MSI testing for gastrointestinal cancers. Thus, there is a pressing need for a new diagnostic modality with lower costs and widespread applicability for MSI detection. This study aims to detect MSI directly from H&E histology slides in gastric cancer, providing a cost-effective alternative. The performance of well-known deep convolutional neural networks (DCNNs) and a proposed architecture are compared. Medical image datasets are typically smaller than benchmark datasets like ImageNet, necessitating the use of off-the-shelf DCNN architectures developed for large datasets through techniques such as transfer learning. Designing an architecture proportional to a custom dataset can be tedious and may not yield desirable results. In this work, we propose an automatic method to extract a lightweight and efficient architecture from a given heavy architecture (e.g., well-known off-the-shelf DCNNs) proportional to a specific dataset. To predict MSI instability, we extracted the MicroNet architecture from the Xception network using the proposed method and compared its performance with other well-known architectures. The models were trained using tiles extracted from whole-slide images, and two evaluation strategies, tile-based and whole-slide image (WSI)-based, were employed and compared. Additionally, a visual explanation of the best convolutional neural network model is presented to validate numerical results. The MicroNet architecture achieved the best accuracy (0.85) and area under the curve-receiver operating characteristic curve (0.93), outperforming previous works for the study dataset. The proposed method can be utilized by developers to design lightweight and efficient problem-based neural network architectures, such as MicroNet, for MSI prediction.































Similar content being viewed by others
Data availability
The datasets analyzed during the current study are available in the zenodo repository, https://zenodo.org/record/2530835#.Y9zqn3YzZPY.
Code availability
Source codes of Micronet and utility methods are available at: https://github.com/habibrostami/stad.
References
Cancer (2022). https://www.who.int/news-room/fact-sheets/detail/cancer. Accessed 02 Feb 2022
Demir C, Yener B (2005) Automated cancer diagnosis based on histopathological images: a systematic survey. Rensselaer Polytech Inst Tech Rep
Ahmedt-Aristizabal D, Armin MA, Denman S, Fookes C, Petersson L (2021) A survey on graph-based deep learning for computational histopathology. Comput Med Imaging Graph 95:102027
van der Laak J, Litjens G, Ciompi F (2021) Deep learning in histopathology: the path to the clinic. Nat Med 27(5):775–784
Cho KO, Lee SH, Jang HJ (2020) Feasibility of fully automated classification of whole slide images based on deep learning. Korean J Physiol Pharmacol 24(1):89–99
Arvaniti E et al (2018) Automated Gleason grading of prostate cancer tissue microarrays via deep learning. Sci Rep 8(1):1–11
Steiner DF et al (2018) Impact of deep learning assistance on the histopathologic review of lymph nodes for metastatic breast cancer. Am J Surg Pathol 42(12):1636
Jang H-J, Song IH, Lee SH (2021) Generalizability of deep learning system for the pathologic diagnosis of various cancers. Appl Sci 11(2):808
Echle A et al (2020) Clinical-grade detection of microsatellite instability in colorectal tumors by deep learning. Gastroenterology 159(4):1406–1416
Li K, Luo H, Huang L, Luo H, Zhu X (2020) Microsatellite instability: a review of what the oncologist should know. Cancer Cell Int 20(1):1–13
Kim YB et al (2016) Microsatellite instability of gastric and colorectal cancers as a predictor of synchronous gastric or colorectal neoplasms. Gut Liver 10(2):220
Russakovsky O et al (2015) Imagenet large scale visual recognition challenge. Int J Comput Vis 115(3):211–252
Liu W, Wang Z, Liu X, Zeng N, Liu Y, Alsaadi FE (2017) A survey of deep neural network architectures and their applications. Neurocomputing 234:11–26
Rajpurkar P et al. (2017) Chexnet: radiologist-level pneumonia detection on chest x-rays with deep learning. ArXiv Prepr. ArXiv171105225
Shen D, Wu G, Suk H-I (2017) Deep learning in medical image analysis. Annu Rev Biomed Eng 19:221–248
Chen JH, Asch SM (2017) Machine learning and prediction in medicine—beyond the peak of inflated expectations. N Engl J Med 376(26):2507
Liu F et al (2018) Deep learning approach for evaluating knee MR images: achieving high diagnostic performance for cartilage lesion detection. Radiology 289(1):160–169
A Survey on Deep Reinforcement Learning--«Chinese Journal of Computers» 2018年01期 (2022). http://en.cnki.com.cn/Article_en/CJFDTOTAL-JSJX201801001.htm. Accessed 18 Apr 2022
Hornik K, Stinchcombe M, White H (1989) Multilayer feedforward networks are universal approximators. Neural Netw 2(5):359–366
Zhou J, Xiao D, Zhang M (2019) Feature correlation loss in convolutional neural networks for image classification. In: 2019 IEEE 3rd information technology, networking, electronic and automation control conference (ITNEC), pp 219–223. https://doi.org/10.1109/ITNEC.2019.8729534
Chollet F (2017) Xception: deep learning with depthwise separable convolutions. In: 2017 proceedings of the IEEE conference on computer vision and pattern recognition. CVPR, pp 1800–1807. https://doi.org/10.1109/CVPR.2017.195
Sabour S, Frosst N, Hinton GE (2017) Dynamic routing between capsules. Neural Inf Process Syst. https://doi.org/10.1371/journal.pone.0035195
Alber M et al (2019) iNNvestigate neural networks! J Mach Learn Res 20(93):1–8
Montavon G, Samek W, Müller K-R (2018) Methods for interpreting and understanding deep neural networks. Digit Signal Process 73:1–15
Kindermans PJ et al. (2017) Learning how to explain neural networks: patternnet and patternattribution. ArXiv Prepr. ArXiv170505598
Camburu OM (2020) Explaining deep neural networks. ArXiv Prepr. ArXiv201001496
Hildebrand LA, Pierce CJ, Dennis M, Paracha M, Maoz A (2021) Artificial intelligence for histology-based detection of microsatellite instability and prediction of response to immunotherapy in colorectal cancer. Cancers 13(3):391
Pressman IS, Xu H, Kang J, Cha YJ, Lee SH, Hwang TH (2020) Deep learning can predict microsatellite instability from histology in colorectal cancer across different ethnic groups. Cancer Res 80:2100
Yamashita R et al (2021) Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study. Lancet Oncol 22(1):132–141
Kather JN (2019) Histological images for MSI vs. MSS classification in gastrointestinal cancer, FFPE samples. ZENODO.
Zhang W et al (2020) MRI-based deep learning analysis can predict microsatellite instability in rectal cancer. SSRN Electron J. https://doi.org/10.2139/ssrn.3569821
Wang T et al. (2020) Microsatellite instability prediction of uterine corpus endometrial carcinoma based on H&E histology whole-slide imaging. In: 2020 IEEE 17th international symposium on biomedical imaging (ISBI), IEEE, pp 1289–1292
Hong R, Liu W, DeLair D, Razavian N, Fenyö D (2021) Predicting endometrial cancer subtypes and molecular features from histopathology images using multi-resolution deep learning models. Cell Rep Med 2(9):100400. https://doi.org/10.1016/j.xcrm.2021.100400
Schmauch B et al (2020) A deep learning model to predict RNA-Seq expression of tumours from whole slide images. Nat Commun 11(1):1–15
Zhu J et al. (2020) Computational analysis of pathological image enables interpretable prediction for microsatellite instability. ArXiv Prepr. ArXiv201003130
Muti HS et al (2021) Development and validation of deep learning classifiers to detect Epstein-Barr virus and microsatellite instability status in gastric cancer: a retrospective multicentre cohort study. Lancet Digit Health 3(10):e654–e664
Kather JN et al (2019) Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med 25(7):1054–1056. https://doi.org/10.1038/s41591-019-0462-y
Xu TB, Liu CL (2022) Deep neural network self-distillation exploiting data representation invariance. IEEE Trans Neural Netw Learn Syst 33(1):257–269. https://doi.org/10.1109/TNNLS.2020.3027634
Ayinde BO, Inanc T, Zurada JM (2019) Redundant feature pruning for accelerated inference in deep neural networks. Neural Netw 118:148–158. https://doi.org/10.1016/j.neunet.2019.04.021
Xu C, Gao W, Li T, Bai N, Li G, Zhang Y (2023) Teacher-student collaborative knowledge distillation for image classification. Appl Intell 53(2):1997–2009
van Erven T, Harremoes P (2014) Rényi divergence and Kullback-Leibler divergence. IEEE Trans Inf Theory 60(7):3797–3820. https://doi.org/10.1109/TIT.2014.2320500
Anghel A et al (2019) A high-performance system for robust stain normalization of whole-slide images in histopathology. Front Med 6:193
Reinhard E, Adhikhmin M, Gooch B, Shirley P (2001) Color transfer between images. IEEE Comput Graph Appl 21(5):34–41
Khan AM, Rajpoot N, Treanor D, Magee D (2014) A nonlinear mapping approach to stain normalization in digital histopathology images using image-specific color deconvolution. IEEE Trans Biomed Eng 61(6):1729–1738
Vahadane A et al (2016) Structure-preserving color normalization and sparse stain separation for histological images. IEEE Trans Med Imaging 35(8):1962–1971
Macenko M et al. (2009) A method for normalizing histology slides for quantitative analysis. In: 2009 IEEE international symposium on biomedical imaging: from nano to macro, IEEE, pp 1107–1110
The Cancer Genome Atlas - Colorectal Carcinoma Study - National Cancer Institute (2022). https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga/studied-cancers/colorectal. Accessed 03 Feb 2022
UNKNOWN (2022) sklearn: a set of python modules for machine learning and data mining. https://pypi.python.org/pypi/scikit-learn/. Accessed 03 Feb 2022
Keras: the Python deep learning API (2022). https://keras.io/. Accessed 03 Feb 2022
TensorFlow (2022). https://www.tensorflow.org/. Accessed 03 Feb 2022
Wang N, Zeng NN, Zhu W (2010) Sensitivity, specificity, accuracy, associated confidence interval and ROC analysis with practical SAS implementations. p 9
Dahiya N, Gupta S, Garg M (2021) Microsatellite instability in gastrointestinal cancer using deep learning: a review. In: IOP conference series: materials science and engineering, IOP Publishing, p 012025
Kuntz S et al (2021) Gastrointestinal cancer classification and prognostication from histology using deep learning: systematic review. Eur J Cancer 155:200–215
Rawla P, Barsouk A (2019) Epidemiology of gastric cancer: global trends, risk factors and prevention. Przeglad Gastroenterol 14(1):26–38. https://doi.org/10.5114/pg.2018.80001
Behzadi-Khormouji H et al (2020) Deep learning, reusable and problem-based architectures for detection of consolidation on chest X-ray images. Comput Methods Progr Biomed 185:105162
Hu B, El Hajj N, Sittler S, Lammert N, Barnes R (2012) Gastric cancer: classification, histology and application of molecular pathology. AJ Gastrointest Oncol 3(3):251–261
Musallam AS, Sherif AS, Hussein MK (2022) Efficient framework for detecting COVID-19 and pneumonia from chest X-ray using deep convolutional network. Egypt Inf J 23:247–257
Mehrotra R, Agrawal R, Ansari MA (2022) Diagnosis of hypercritical chronic pulmonary disorders using dense convolutional network through chest radiography. Multimed Tools Appl 81(6):7625–7649
Hou J, Gao T (2021) Explainable DCNN based chest X-ray image analysis and classification for COVID-19 pneumonia detection. Sci Rep 11(1):1–15
Jin W, Li X, Hamarneh G (2022) Evaluating explainable AI on a multi-modal medical imaging task: can existing algorithms fulfill clinical requirements?
Grün F, Rupprecht C, Navab N, Tombari F (2016) A taxonomy and library for visualizing learned features in convolutional neural networks. ArXiv Prepr. ArXiv160607757
Zhang Y, Tiňo P, Leonardis A, Tang K (2021) A survey on neural network interpretability. IEEE Trans Emerg Top Comput Intell 5:726–742
Bodria F, Giannotti F, Guidotti R, Naretto F, Pedreschi D, Rinzivillo S (2021) Benchmarking and survey of explanation methods for black box models. ArXiv Prepr. ArXiv210213076
Wickramanayake S, Hsu W, Lee ML (2021) Towards fully interpretable deep neural networks: are we there yet?. ArXiv Prepr. ArXiv210613164
Schwalbe G, Finzel B (2021) XAI method properties: a (Meta-) study. ArXiv Prepr. ArXiv210507190
Behzadi-Khormouji H, Rostami H (2021) Fast multi-resolution occlusion: a method for explaining and understanding deep neural networks. Appl Intell 51(4):2431–2455
Zeiler MD, Fergus R (2014) Visualizing and understanding convolutional networks. European conference on computer vision. Springer, Cham, pp 818–833
Ribeiro MT, Singh S, Guestrin C (2016) Why should i trust you? Explaining the predictions of any classifier. In: Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining, pp 1135–1144
Petsiuk V, Das A, Saenko K (2018) Rise: randomized input sampling for explanation of black-box models. ArXiv Prepr. ArXiv180607421
Simonyan K, Vedaldi A, Zisserman A (2013) Deep inside convolutional networks: visualising image classification models and saliency maps. ArXiv Prepr. ArXiv13126034
Springenberg JT, Dosovitskiy A, Brox T, Riedmiller M (2014) Striving for simplicity: the all convolutional net. ArXiv Prepr. ArXiv14126806
Zhou B, Khosla A, Lapedriza A, Oliva A, Torralba A (2016) Learning deep features for discriminative localization. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 2921–2929
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D (2017) Grad-cam: visual explanations from deep networks via gradient-based localization. In: Proceedings of the IEEE international conference on computer vision, pp 618–626
Qi Z, Khorram S, Li F (2019) Visualizing deep networks by optimizing with integrated gradients. In: CVPR workshops
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rostami, H., Ashkpour, M., Behzadi-Khormouji, H. et al. Automatic extraction of lightweight and efficient neural network architecture of heavy convolutional architectures to predict microsatellite instability from hematoxylin and eosin histology in gastric cancer. Neural Comput & Applic (2024). https://doi.org/10.1007/s00521-024-09882-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00521-024-09882-w