Summary
Background
Osteoporosis is the most frequent bone metabolic disease. In order to improve early detection, prediction, prevention, diagnosis, and treatment of the disease, a new model of P4 medicine (personalized, predictive, preventive, and participatory medicine) could be applied. The aim of this work was to systematically review the publications of four different types of “omics” studies related to osteoporosis, in order to discover novel predictive, preventive, diagnostic, and therapeutic targets for better management of the geriatric population.
Methods
To systematically search the PubMed database, we created specific groups of criteria for four different types of “omics” information on osteoporosis: genomic, transcriptomic, proteomic, and metabolomic. We then analyzed the intersections between them in order to find correlations and common pathways or molecules with important roles in osteoporosis, and with a potential application in disease prediction, prevention, diagnosis, or treatment.
Results
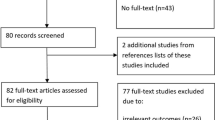
Altogether, 180 publications of “omics” studies in the field of osteoporosis were found and reviewed at first selection. After introducing the inclusion and exclusion criteria (the secondary selection), 46 papers were included in the systematic review.
Conclusions
The intersection of reviewed papers identified five genes (ESR1, IBSP, CTNNB1, SOX4, and IDUA) and processes like the Wnt pathway, JAK/STAT signaling, and ERK/MAPK, which should be further validated for their predictive, diagnostic, or other clinical value in osteoporosis. Such molecular insights will enable us to fit osteoporosis into the P4 strategy and could increase the effectiveness of disease prediction and prevention, with a decrease in morbidity in the geriatric population.




Similar content being viewed by others
References
Košnik MMF, Štajer D, Koželj M, Černelč P. Interna medicina, 4th ed. Ljubljana: Littera picta; 2011.
Mafi Golchin M, Heidari L, Ghaderian SM, Akhavan-Niaki H. Osteoporosis: a silent disease with complex genetic contribution. J Genet Genomics. 2016;43(2):49–61.
Hui SL, Slemenda CW, Johnston CC Jr.. Age and bone mass as predictors of fracture in a prospective study. J Clin Invest. 1988;81(6):1804–9.
De Laet C, Kanis JA, Oden A, Johanson H, Johnell O, Delmas P, et al. Body mass index as a predictor of fracture risk: a meta-analysis. Osteoporos Int. 2005;16(11):1330–8.
Klotzbuecher CM, Ross PD, Landsman PB, Abbott TA, Berger M. Patients with prior fractures have an increased risk of future fractures: a summary of the literature and statistical synthesis. J Bone Miner Res. 2000;15(4):721–39.
Kanis JA, Johnell O, Oden A, Johansson H, De Laet C, Eisman JA, et al. Smoking and fracture risk: a meta-analysis. Osteoporos Int. 2005;16(2):155–62.
Kanis JA, Johansson H, Johnell O, Oden A, De Laet C, Eisman JA, et al. Alcohol intake as a risk factor for fracture. Osteoporos Int. 2005;16(7):737–42.
Kanis JA, Johansson H, Oden A, Johnell O, de Laet C, Melton IL, et al. A meta-analysis of prior corticosteroid use and fracture risk. J Bone Miner Res. 2004;19(6):893–9.
Ugurlu U, Nayki U, Nayki C, Ulug P, Kulhan M, Yildirim Y. Assessment of smoking for low bone mineral density in postmenopausal Turkish women. Wien Klin Wochenschr. 2016;128(3):114–9.
Hood L, Rowen L, Galas DJ, Aitchison JD. Systems biology at the Institute for Systems Biology. Brief Funct Genomic Proteomic. 2008;7(4):239–48.
Sobradillo P, Pozo F, Agusti A. P4 medicine: the future around the corner. Arch Bronconeumol. 2011;47(1):35–40.
Younesi E, Hofmann-Apitius M. From integrative disease modeling to predictive, preventive, personalized and participatory (P4) medicine. EPMA J. 2013;4(1):23.
Flores M, Glusman G, Brogaard K, Price ND, Hood L. P4 medicine: how systems medicine will transform the healthcare sector and society. Per Med. 2013;10(6):565–76.
National Research Council (US) Committee on A Framework for Developing a New Taxonomy of Disease. Toward precision medicine: building a knowledge network for biomedical research and a new taxonomy of disease. Washington DC: National Academies Press; 2011.
Hood L, Flores M. A personal view on systems medicine and the emergence of proactive P4 medicine: predictive, preventive, personalized and participatory. N Biotechnol. 2012;29(6):613–24.
David JG, Leroy H. Systems biology and emerging technologies will catalyze the transition from reactive medicine to predictive, personalized, preventive and participatory (P4) medicine. Interdiscipl Bio Cent. 2009;1:6.
Hood L. Systems biology and p4 medicine: past, present, and future. Rambam Maimonides Med J. 2013;4(2):e0012.
Hood L, Balling R, Auffray C. Revolutionizing medicine in the 21st century through systems approaches. Biotechnol J. 2012;7(8):992–1001.
Estrada K, Styrkarsdottir U, Evangelou E, Hsu YH, Duncan EL, Ntzani EE, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet. 2012;44(5):491–501.
Guo Y, Tan LJ, Lei SF, Yang TL, Chen XD, Zhang F, et al. Genome-wide association study identifies ALDH7A1 as a novel susceptibility gene for osteoporosis. PLOS Genet. 2010;6(1):e1000806.
Rivadeneira F, Styrkarsdottir U, Estrada K, Halldorsson BV, Hsu YH, Richards JB, et al. Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat Genet. 2009;41(11):1199–206.
Duncan EL, Danoy P, Kemp JP, Leo PJ, McCloskey E, Nicholson GC, et al. Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLOS Genet. 2011;7(4):e1001372.
Zhang L, Choi HJ, Estrada K, Leo PJ, Li J, Pei YF, et al. Multistage genome-wide association meta-analyses identified two new loci for bone mineral density. Hum Mol Genet. 2014;23(7):1923–33.
Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. Multiple genetic loci for bone mineral density and fractures. N Engl J Med. 2008;358(22):2355–65.
Guo Y, Zhang LS, Yang TL, Tian Q, Xiong DH, Pei YF, et al. IL21R and PTH may underlie variation of femoral neck bone mineral density as revealed by a genome-wide association study. J Bone Miner Res. 2010;25(5):1042–8.
Kung AW, Xiao SM, Cherny S, Li GH, Gao Y, Tso G, et al. Association of JAG1 with bone mineral density and osteoporotic fractures: a genome-wide association study and follow-up replication studies. Am J Hum Genet. 2010;86(2):229–39.
Richards JB, Rivadeneira F, Inouye M, Pastinen TM, Soranzo N, Wilson SG, et al. Bone mineral density, osteoporosis, and osteoporotic fractures: a genome-wide association study. Lancet. 2008;371(9623):1505–12.
Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. New sequence variants associated with bone mineral density. Nat Genet. 2009;41(1):15–7.
Zheng HF, Duncan EL, Yerges-Armstrong LM, Eriksson J, Bergstrom U, Leo PJ, et al. Meta-analysis of genome-wide studies identifies MEF2C SNPs associated with bone mineral density at forearm. J Med Genet. 2013;50(7):473–8.
Guo Y, Wang JT, Liu H, Li M, Yang TL, Zhang XW, et al. Are bone mineral density loci associated with hip osteoporotic fractures? A validation study on previously reported genome-wide association loci in a Chinese population. Genet Mol Res. 2012;11(1):202–10.
Xiao SM, Kung AW, Gao Y, Lau KS, Ma A, Zhang ZL, et al. Post-genome wide association studies and functional analyses identify association of MPP7 gene variants with site-specific bone mineral density. Hum Mol Genet. 2012;21(7):1648–57.
Styrkarsdottir U, Thorleifsson G, Gudjonsson SA, Sigurdsson A, Center JR, Lee SH, et al. Sequence variants in the PTCH1 gene associate with spine bone mineral density and osteoporotic fractures. Nat Commun. 2016;7:10129.
Paternoster L, Lorentzon M, Vandenput L, Karlsson MK, Ljunggren O, Kindmark A, et al. Genome-wide association meta-analysis of cortical bone mineral density unravels allelic heterogeneity at the RANKL locus and potential pleiotropic effects on bone. PLOS Genet. 2010;6(11):e1001217.
Trost Z, Trebse R, Prezelj J, Komadina R, Logar DB, Marc J. A microarray based identification of osteoporosis-related genes in primary culture of human osteoblasts. Bone. 2010;46(1):72–80.
Mak YT, Hampson G, Beresford JN, Spector TD. Variations in genome-wide gene expression in identical twins – a study of primary osteoblast-like culture from female twins discordant for osteoporosis. BMC Genet. 2004;5:14.
Hopwood B, Tsykin A, Findlay DM, Fazzalari NL. Gene expression profile of the bone microenvironment in human fragility fracture bone. Bone. 2009;44(1):87–101.
Liu YZ, Dvornyk V, Lu Y, Shen H, Lappe JM, Recker RR, et al. A novel pathophysiological mechanism for osteoporosis suggested by an in vivo gene expression study of circulating monocytes. J Biol Chem. 2005;280(32):29011–6.
Chen XD, Xiao P, Lei SF, Liu YZ, Guo YF, Deng FY, et al. Gene expression profiling in monocytes and SNP association suggest the importance of the STAT1 gene for osteoporosis in both Chinese and Caucasians. J Bone Miner Res. 2010;25(2):339–55.
Liu YZ, Zhou Y, Zhang L, Li J, Tian Q, Zhang JG, et al. Attenuated monocyte apoptosis, a new mechanism for osteoporosis suggested by a transcriptome-wide expression study of monocytes. PLOS ONE. 2015;10(2):e0116792.
Xiao P, Chen Y, Jiang H, Liu YZ, Pan F, Yang TL, et al. In vivo genome-wide expression study on human circulating B cells suggests a novel ESR1 and MAPK3 network for postmenopausal osteoporosis. J Bone Miner Res. 2008;23(5):644–54.
Chen Y, Xia RG. Screening and functional microarray analysis of differentially expressed genes related to osteoporosis. Genet Mol Res. 2014;13(2):3228–36.
Yan B, Li J, Zhang L. Identification of B cells participated in the mechanism of postmenopausal women osteoporosis using microarray analysis. Int J Clin Exp Med. 2015;8(1):1027–34.
Ma M, Chen X, Lu L, Yuan F, Zeng W, Luo S, et al. Identification of crucial genes related to postmenopausal osteoporosis using gene expression profiling. Aging Clin Exp Res. 2015; doi:10.1007/s40520-015-0509-y.
Liu L, Zhu Q, Wang J, Xi Q, Zhu H, Gu M. Gene expression changes in human mesenchymal stem cells from patients with osteoporosis. Mol Med Rep. 2015;12(1):981–7.
Zhou Z, Gao M, Liu Q, Tao MD. Comprehensive transcriptome analysis of mesenchymal stem cells in elderly patients with osteoporosis. Aging Clin Exp Res. 2015;27(5):595–601.
Xie W, Ji L, Zhao T, Gao P. Identification of transcriptional factors and key genes in primary osteoporosis by DNA microarray. Med Sci Monit. 2015;21:1333–44.
Wu X, Guo S, Shen G, Ma X, Tang C, Xie K, et al. Screening of osteoprotegerin-related feature genes in osteoporosis and functional analysis with DNA microarray. Eur J Med Res. 2013;18:15.
Wen Y, Guo X, Hao J, Xiao X, Wang W, Wu C, et al. Integrative analysis of genome-wide association studies and gene expression profiles identified candidate genes for osteoporosis in Kashin-Beck disease patients. Osteoporos Int. 2016;27(3):1041–6.
Zhang Y, Wang N, Ma J, Chen XC, Li Z, Zhao W. Expression profile analysis of new candidate genes for the therapy of primary osteoporosis. Eur Rev Med Pharmacol Sci. 2016;20(3):433–40.
Pineda B, Serna E, Laguna-Fernandez A, Noguera I, Panach L, Hermenegildo C, et al. Gene expression profile induced by ovariectomy in bone marrow of mice: a functional approach to identify new candidate genes associated to osteoporosis risk in women. Bone. 2014;65:33–41.
He H, Cao S, Niu T, Zhou Y, Zhang L, Zeng Y, et al. Network-based meta-analyses of associations of multiple gene expression profiles with bone mineral density variations in women. PLOS ONE. 2016;11(1):e0147475.
Deng FY, Liu YZ, Li LM, Jiang C, Wu S, Chen Y, et al. Proteomic analysis of circulating monocytes in Chinese premenopausal females with extremely discordant bone mineral density. Proteomics. 2008;8(20):4259–72.
Daswani B, Gupta MK, Gavali S, Desai M, Sathe GJ, Patil A, et al. Monocyte proteomics reveals involvement of phosphorylated HSP27 in the pathogenesis of osteoporosis. Dis Markers. 2015;2015:196589. doi:10.1155/2015/196589.
Zhang L, Liu YZ, Zeng Y, Zhu W, Zhao YC, Zhang JG, et al. Network-based proteomic analysis for postmenopausal osteoporosis in Caucasian females. Proteomics. 2016;16(1):12–28.
Zeng Y, Zhang L, Zhu W, Xu C, He H, Zhou Y, et al. Quantitative proteomics and integrative network analysis identified novel genes and pathways related to osteoporosis. J Proteomics. 2016;142:45–52.
Qundos U, Drobin K, Mattsson C, Hong MG, Sjoberg R, Forsstrom B, et al. Affinity proteomics discovers decreased levels of AMFR in plasma from Osteoporosis patients. Proteomics Clin Appl. 2016;10(6):681–90.
Fan Y, Liu J, Wang S, Wang H, Shi F, Xiong L, et al. Functional proteome of bones in rats with osteoporosis following ovariectomy. Life Sci. 2005;76(25):2893–901.
You YS, Lin CY, Liang HJ, Lee SH, Tsai KS, Chiou JM, et al. Association between the metabolome and low bone mineral density in Taiwanese women determined by (1)H NMR spectroscopy. J Bone Miner Res. 2014;29(1):212–22.
Qi H, Bao J, An G, Ouyang G, Zhang P, Wang C, et al. Association between the metabolome and bone mineral density in pre- and post-menopausal Chinese women using GC-MS. Mol Biosyst. 2016;12(7):2265–75. doi:10.1039/c6mb00181e.
Ma B, Liu J, Zhang Q, Ying H, Jiye A, Sun J, et al. Metabolomic profiles delineate signature metabolic shifts during estrogen deficiency-induced bone loss in rat by GC-TOF/MS. PLOS ONE. 2013;8(2):e54965.
Liu X, Liu Y, Cheng M, Zhang X, Xiao H. A metabolomics study of the inhibitory effect of 17-beta-estradiol on osteoclast proliferation and differentiation. Mol Biosyst. 2015;11(2):635–46.
NCBI. SMG6, nonsense mediated mRNA decay factor Gene ID: 23293 2016. https://www.ncbi.nlm.nih.gov/gene/23293. Accessed 26 June 2016.
Moore SC, Matthews CE, Sampson JN, Stolzenberg-Solomon RZ, Zheng W, Cai Q, et al. Human metabolic correlates of body mass index. Metabolomics. 2013;10(2):259–69.
NCBI. ESR1 estrogen receptor 1 (Homo sapiens [human]), Gene ID: 2099 2016. http://www.ncbi.nlm.nih.gov/gene/2099. Accessed 21 June 2016.
NCBI IBSP integrin binding sialoprotein (Homo sapiens [human]) Gene ID: 3381 2008. https://www.ncbi.nlm.nih.gov/gene/?term=3381. Accessed 26 June 2016.
NCBI. CTNNB1 catenin beta 1 (Homo sapiens [human]) Gene ID: 1499 2009. https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=1499. Accessed 28 June 2016.
NCBI SOX4 SRY-box 4 (Homo sapiens [human]) Gene ID: 6659 2008. https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=6659. Accessed 28 June 2016.
NCBI IDUA iduronidase, alpha-L- (Homo sapiens [human]) Gene ID: 3425 2008. https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=3425. Accessed 28 June 2016.
Acknowledgement
The study was financially supported by the Slovenian Research Agency (J3-7245, P3-298, and MR38167).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
K. Kodrič, K. Čamernik, D. Černe, R. Komadina, and J. Marc declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Kodrič, K., Čamernik, K., Černe, D. et al. P4 medicine and osteoporosis: a systematic review. Wien Klin Wochenschr 128 (Suppl 7), 480–491 (2016). https://doi.org/10.1007/s00508-016-1125-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00508-016-1125-3