Abstract
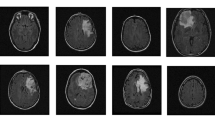
Malignant tumors can cause to brain tumors by the spreading to the brain. When the primary site of origin of brain metastases are differentiated by deep learning algorithms, doctors can begin treatment in time without performing biopsy. In this study, we aimed to classify the primary site of brain metastases from the MR images of patients with brain tumors. The MR images were read by radiologists from Kocaeli University Department of Radiology by using their Picture Archiving and Communication System (PACS). We used the three-dimensional convolutional neural networks technique, which is one of the artificial intelligence techniques that are frequently used to determine image classification and identify segmentation problems for the classification using a data set consisting of labeled MR images. Contrary to the models used in the studies performed on MR images in the literature, four different sequences were submitted to the classification model after four different convolutional processes were conducted, although not simultaneously. The classification scheme was designed as a binary classification and a three group classifications. The results showed that 81.48% accuracy rate was obtained in the binary classification and 74.07% accuracy was obtained in the three-group classification. To improve the accuracy rate of the three group classifications, data augmentation was applied and the accuracy rate improved to 75.92%.











Similar content being viewed by others
Data availability
Enquiries about data availability should be directed to the authors.
References
Achrol AS, Rennert RC, Anders C, Soffietti R, Ahluwalia MS, Nayak L, Chang SD (2019) Brain metastases. Nat Rev Dis Prim 5(1):1–26. https://doi.org/10.1038/s41572-018-0055-y
Ahmad F, Ahmad I, Dar WM (2017) Identification and classification of voxels of human brain for rewardless-related decision making using ANN technique. Neural Comput Appl 28(1):1035–1041. https://doi.org/10.1007/s00521-016-2413-6
Aizer AA, Lee EQ (2018) Brain metastases. Neurol Clin 36(3):557–577. https://doi.org/10.1016/j.ncl.2018.04.010
Al-Areqi F, Konyar MZ (2022) High accuracy classification of covid-19 from CT images using transfer learning architectures. DUJE Dicle Univ J Eng 13(3):457–466. https://doi.org/10.24012/dumf.1129870
Al-Areqi F, Konyar MZ (2022) Effectiveness evaluation of different feature extraction methods for classification of covid-19 from computed tomography images: a high accuracy classification study. Biomed Signal Process Control 76:103662. https://doi.org/10.1016/j.bspc.2022.103662
Amin J, Sharif M, Yasmin M, Fernandes SL (2018) Big data analysis for brain tumor detection: deep convolutional neural networks. Futur Gener Comput Syst 87:290–297. https://doi.org/10.1016/j.future.2018.04.065
Artzi M, Bressler I, Ben Bashat D (2019) Differentiation between glioblastoma, brain metastasis and subtypes using radiomics analysis. J Magn Reson Imaging 50(2):519–528. https://doi.org/10.1002/jmri.26643
Banik PP, Saha R, Kim KD (2020) An automatic nucleus segmentation and CNN model based classification method of white blood cell. Expert Syst Appl 149:113211. https://doi.org/10.1016/j.eswa.2020.113211
Charron O, Lallement A, Jarnet D, Noblet V, Clavier JB, Meyer P (2018) Automatic detection and segmentation of brain metastases on multimodal MR images with a deep convolutional neural network. Comput Biol Med 95:43–54. https://doi.org/10.1016/j.compbiomed.2018.02.004
Chollet F (2018a) Keras: the python deep learning library. Astrophysics Source Code Library. [Online]. https://keras.io/.
Chollet F (2018b) Deep learning with python. Manning Publications, Shelter Island, p 384
Çinar A, Yıldırım M (2020) Detection of tumors on brain MRI images using the hybrid convolutional neural network architecture. Med Hypotheses 139:109684. https://doi.org/10.1016/j.mehy.2020.109684
Havaei M, Davy A, Warde-Farley D, Biard A, Courville A, Bengio Y, Pal C, Jodoin P-M, Larochelle H (2017) Brain tumor segmentation with deep neural networks. Med Image Anal 35:18–31. https://doi.org/10.1016/j.media.2016.05.004
Kamnitsas K, Ledig C, Newcombe VF, Simpson JP, Kane AD, Menon DK, Rueckert D, Glocker B (2016) Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation, Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med Image Anal 36:61–78. https://doi.org/10.1016/j.media.2016.10.004
Kaplan K, Kaya Y, Kuncan M, Minaz MR, Ertunç HM (2020) An improved feature extraction method using texture analysis with LBP for bearing fault diagnosis. Appl Soft Comput 87:106019. https://doi.org/10.1016/j.asoc.2019.106019
Kniep HC, Madesta F, Schneider T, Hanning U, Schönfeld MH, Schön G, Gellissen S (2018) Radiomics of Brain MRI: utility in prediction of metastatic tumor type. Radiology 290(2):479–487. https://doi.org/10.1148/radiol.2018180946
Liao P, Wu H, Yu T (2017) ROC curve analysis in the presence of imperfect reference standards. Stat Biosci 9(1):91–104. https://doi.org/10.1007/s12561-016-9159-7
Liu BJ, Huang HK (2020) Picture archiving and communication systems and electronic medical records for the healthcare enterprise. Biomedical information technology. Academic Press, USA, pp 105–164
Mohsen H, El-Dahshan EA, El-Horbaty EM, and Salem AM (2017) Brain tumor type classification based on support vector machine in magnetic resonance images. Annals of “Dunarea De Jos” University of Galati, Mathematics, Physics, Theoretical mechanics, Fascicle II, Year IX (XL), (1)
Momeny M, Jahanbakhshi A, Jafarnezhad K, Zhang YD (2020) Accurate classification of cherry fruit using deep CNN based on hybrid pooling approach. Postharvest Biol Technol 166:111204. https://doi.org/10.1016/j.postharvbio.2020.111204
Ortiz-Ramon R, Larroza A, Ruiz-Espana S, Arana E, Moratal D, Member S (2017) Identifying the primary site of origin of mri brain metastases from lung and breast cancer following a 2D radiomics approach. In: 2017 IEEE 14th international symposium on biomedical imaging (ISBI 2017), Melbourne, 18-21, pp. 12123-1216. https://doi.org/10.1109/ISBI.2017.7950735
Ortiz-Ramón R, Larroza A, Ruiz-España S, Arana E, Moratal D (2018) Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur Radiol 28(11):4514–4523. https://doi.org/10.1007/s00330-018-5463-6
Ortiz-Ramon R., Larroza A., Ruiz-Espana S., Arana E., Moratal D. ve Member S., (2017) A radiomics evaluation of 2D and 3D MRI texture features to classify brain metastases from lung cancer and melanoma. In: Proceedings intenational annual conference of IEEE engineering in medicine and biology society (EMBC), Jeju Island, 11–15, pp 493–496. https://doi.org/10.1109/EMBC.2017.8036869
Pham CH, Ducournau A, Fablet R, and Rousseau F (2017) Brain MRI super-resolution using deep 3D convolutional networks. In: 2017 IEEE 14th international symposium on biomedical imaging (ISBI 2017), Melbourne, 8–21, pp 197–200. https://doi.org/10.1109/ISBI.2017.7950500
Qian Z, Li Y, Wang Y, Li L, Li R, Wang K, Li S, Tang K, Zhang C, Fan X, Chen B, Li W (2019) Differentiation of glioblastoma from solitary brain metastases using radiomic machine-learning classifiers. Cancer Lett 451:128–135. https://doi.org/10.1016/j.canlet.2019.02.054
Rajagopalan R, Wallace RA, Periasamy MP (1992) U.S. Patent No. 5,141,740. Washington, DC: U.S. Patent and Trademark Office.
Ryu A, Cho NJ, Kim YS, Lee EY (2019) Predictive value of serum uric acid levels for adverse perinatal outcomes in preeclampsia. Medicine. https://doi.org/10.1097/MD.0000000000015462
Sajjad M, Khan S, Muhammad K, Wu W, Ullah A, Baik SW (2019) Multi-grade brain tumor classification using deep CNN with extensive data augmentation. J Comput Sci 30:174–182. https://doi.org/10.1016/j.jocs.2018.12.003
Saleh NAA, Konyar MZ, Kaplan K, Ongir S and Ertunç HM (2022b) Detection of Air Bubbles from Tire Shearography Images. In: 2022b International congress on human-computer interaction, optimization and robotic applications (HORA), 2022b, pp. 1-4, https://doi.org/10.1109/HORA55278.2022.9799926
Saleh RAA, Konyar MZ, Kaplan K and Ertunç HM (2022a) Tire defect detection model using machine learning. In: 2022a 2nd International conference on emerging smart technologies and applications (eSmarTA), pp 1–5. https://doi.org/10.1109/eSmarTA56775.2022.9935140.
Toğaçar M, Ergen B, Cömert Z (2020) BrainMRNet: Brain tumor detection using magnetic resonance images with a novel convolutional neural network model. Med Hypotheses 134:109531. https://doi.org/10.1016/j.mehy.2019.109531
Toğaçar M, Ergen B, Cömert Z (2020) Application of breast cancer diagnosis based on a combination of convolutional neural networks, ridge regression and linear discriminant analysis using invasive breast cancer images processed with autoencoders. Med Hypotheses 135:109503. https://doi.org/10.1016/j.mehy.2019.109503
Tsao MN, Xu W, Wong RK, Lloyd N, Laperriere N, Sahgal A, Chow E (2018) Whole brain radiotherapy for the treatment of newly diagnosed multiple brain metastases. Cochrane Database Syst Rev 52:1. https://doi.org/10.1002/14651858.CD003869.pub3
Tuan TA, Kim JY, Bao PT (2018) 3D brain magnetic resonance imaging segmentation by using bitplane and adaptive fast marching. Int J Imaging Syst Technol 28(3):223–230. https://doi.org/10.1002/ima.22273
Viergever MA, Maintz JA, Klein S, Murphy K, Staring M, Pluim JP (2016) A survey of medical image registration–under review. Med Image Anal 33(10):140–144. https://doi.org/10.1016/j.media.2016.06.030
Wang C, Cheng M, Sohel F, Bennamoun M, Li J (2019) NormalNet: a voxel-based CNN for 3D object classification and retrieval. Neurocomputing 323:139–147. https://doi.org/10.1016/j.neucom.2018.09.075
Yan L (2018) DICOM standard and its application in PACS system. Med Imaging Process Technol 1(1):34–41. https://doi.org/10.24294/mipt.v1i1.221
Zhang YD, Dong Z, Chen X, Jia W, Du S, Muhammad K, Wang SH (2019) Image based fruit category classification by 13-layer deep convolutional neural network and data augmentation. Multimed Tools Appl 78(3):3613–3632. https://doi.org/10.1007/s11042-017-5243-3
Zou L, Zheng J, Miao C, Mckeown MJ, Wang ZJ (2017) 3D CNN based automatic diagnosis of attention deficit hyperactivity disorder using functional and structural MRI. IEEE Access 5:23626–23636. https://doi.org/10.1109/ACCESS.2017.2762703
Acknowledgements
This study was conducted in Sensor Laboratory of Mechatronics Engineering Department in Kocaeli University. We would like to thank to Kocaeli University Department of Radiology for their support and efforts.
Funding
The authors have not disclosed any funding
Author information
Authors and Affiliations
Contributions
YC and KK developed the theory and performed the computations. BA conceived the presented idea and prepared the dataset. HME verified the analytical methods. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cuşkun, Y., Kaplan, K., Alparslan, B. et al. Classification of the brain metastases based on a new 3D deep learning architecture. Soft Comput 27, 17243–17256 (2023). https://doi.org/10.1007/s00500-023-08051-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00500-023-08051-w