Abstract
An understanding of the factors that affect the spread of endemic bovine tuberculosis (bTB) is critical for the development of measures to stop and reverse this spread. Analyses of spatial data need to account for the inherent spatial heterogeneity within the data, or else spatial autocorrelation can lead to an overestimate of the significance of variables. This study used three methods of analysis—least-squares linear regression with a spatial autocorrelation term, geographically weighted regression (GWR) and boosted regression tree (BRT) analysis—to identify the factors that influence the spread of endemic bTB at a local level in England and Wales. The linear regression and GWR methods demonstrated the importance of accounting for spatial differences in risk factors for bTB, and showed some consistency in the identification of certain factors related to flooding, disease history and the presence of multiple genotypes of bTB. This is the first attempt to explore the factors associated with the spread of endemic bTB in England and Wales using GWR. This technique improves on least-squares linear regression approaches by identifying regional differences in the factors associated with bTB spread. However, interpretation of these complex regional differences is difficult and the approach does not lend itself to predictive models which are likely to be of more value to policy makers. Methods such as BRT may be more suited to such a task. Here we have demonstrated that GWR and BRT can produce comparable outputs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Bovine Tuberculosis (bTB) is a major challenge for the agricultural industry in Great Britain. When the disease is detected in a cattle herd the infected animals are culled and the herd is placed under movement restrictions which has considerable economic implications for the farmer. Surveillance and control of the disease is funded by the government and represents a considerable burden on public finances, being estimated to have cost the taxpayer £500 million in the past decade (Defra 2014). The distribution of the disease is not homogeneous across Great Britain, with incidence being highest in the south and west of England, along the Welsh/English border and in western counties of Wales (Lawes et al. 2016). Despite the spatial heterogeneity of the disease, control policies for bTB in England and Wales have traditionally been implemented at country level, with a move to more regional policies in the last few years such as the creation of risk areas in England (Defra 2011) and the Intensive Action Area in Wales (Welsh Government 2016). Both have applied different policies which have been designed to suit the level of risk in those areas such as increased testing and testing of contiguous herds following an incident where environmental or wildlife transmission is likely, or using a more sensitive test in low incidence areas to ensure the disease is eliminated before it becomes endemic. Recently, regional differences in a number of measures of bTB have been examined and have provided information on the effectiveness of bTB control polices in different areas (Moustakas and Evans 2016), but the reasons behind the spatial heterogeneity of bTB are not well understood.
BTB can be considered endemic in the high incidence areas of England and Wales, and these areas have been expanding over time. It is important to identify influential factors that affect the expansion of the endemic area, at regional and local levels, so that measures to stop and reverse the spread of bTB can be developed. The rate of this expansion has been shown to be non-uniform (Brunton et al. 2015) and it may be affected by many factors such as landscape characteristics, wildlife, climate, cattle movements, bTB testing and detection, and various other anthropogenic factors. Here we investigate what the drivers of spread are at a national level and at a local level.
Analyses of spatial data need to account for the inherent heterogeneity within the data, as discussed by Lennon (2000). Many factors related to disease spread, such as landscape factors and wildlife reservoirs of disease for example, vary geographically, showing spatial non-stationarity (Brunsdon et al. 1998). Statistical analyses which do not account for spatial autocorrelation can overestimate the significance of variables (Lennon 2000).
Various approaches for handling spatial heterogeneity in a regression model have been developed, including the use of a term that represents either spatial autocorrelation in the dependent variable or in the residuals from the independent variables (Crase et al. 2012) and the use of simultaneous autoregressive models (Pioz et al. 2012). If relationships are thought to alter spatially within the study area, geographically weighted regression (GWR) can be used to produce localised models for different spatial regions which take neighbouring observations into account (Brunsdon et al. 1998; Fotheringham et al. 1998). This is achieved through the use of a ‘moving window’ to identify a subset of the data to which a localised model is applied. GWR has been used across a wide variety of disciplines including agriculture, where it has been used, for example, to measure the spatial distribution of water requirement of crops in North China while adjusting for topographical and meteorological factors (Wang et al. 2013), and public health where it has been used to assess spatial patterns of leishmaniasis in the Middle East (Jaber et al. 2013).
Analyses of risk factors for bTB in Great Britain have historically been performed at the herd level using ordinary least squares (OLS) regression techniques (Reilly and Courtenay 2007; Carrique-Mas et al. 2008; Ramírez-Villaescusa et al. 2010; Johnston et al. 2011; Vial et al. 2011). While such analyses provide useful information about important risk factors for the spread of bTB, they may overlook, or be biased by, important spatial differences in risk factors. GWR may provide a useful alternative approach to gain insights into the spatial variation in the factors associated with the spread of bTB. Improved performance of GWR in comparison to OLS regression has been demonstrated for identifying the factors associated with urban flooding (Wang et al. 2016) and urban population growth (Liao and Wei 2014) in China.
In order to assess the usefulness of GWR in the context of understanding the spread of bTB we have used three methods of analysis—OLS linear regression with a spatial autocorrelation term, GWR and Boosted Regression Tree (BRT) analysis—to explore the factors that influence the spread of endemic bTB at a local level in England and Wales.
2 Methods
2.1 Estimates of rate of spread
The dependent variable for the analysis, rate of spread of endemic bTB per km, was calculated from the estimated location of the endemic front in successive years (the methodology used to generate these data has been described in Brunton et al. 2015). A grid of 6.25 km2 hexagonal cells was applied to England and Wales, and a rate of spread was obtained for all hexagons through which the endemic front was calculated to have spread between September 2001 and August 2012 (n = 2148).
2.2 Variable selection
An extensive dataset of 193 variables was compiled. Variables were selected if there was evidence they were associated with bTB in published literature, and if data were available to describe the variables at the geographical level required for the analysis. The large number of potential co-variates was rationalised by reviewing summary statistics and performing bi-variable least-squares linear regression against the dependent variable, fitting predicted values and visually assessing the residuals. Analyses were performed in Stata 12 (Stata Corporation, College Station, TX, USA), and a significance level of p < 0.05 was used throughout.
For many of the variables, the residuals were not normally distributed so transformation of the data was explored using Box Cox regression. Many of the variables used may have been acting as potential proxies for other factors, and thus be correlated with each other. In an attempt to avoid multicollinearity, pairwise Pearson product-moment correlations of all variables were produced and strong correlations identified ([r] > 0.8). Where two or more variables were highly correlated, the one with the highest correlation with the dependent variable and/or the greater biological plausibility was retained. This resulted in a reduced list of 75 independent variables. A list of these variables including the sources of the data can be found in Table S1 in the Supplementary Information. These variables were grouped under six themes: animal-level factors, farm-level factors, bTB history and testing, landscape characteristics, wildlife and climate. Two variables that were considered as a priori confounders and not grouped under the six themes were the time period in which spread occurred (TpS), and the number of different genotypes of M. bovis present within the hexagon or its neighbouring six hexagons during the time period of spread. TpS, a categorical variable, was coded as an indicator variable. Where appropriate, continuous variables were scaled by centring around the mean, subtracting the lowest observable value from each observation if an intercept of zero was not meaningful, or divided by a suitable constant (e.g. 100) to improve the unit change represented by coefficients. Missing data were examined to determine if they occurred at random or if the fact that they were missing was linked to the actual missing data.
2.3 Linear regression with spatial autocorrelation term
To account for spatial autocorrelation (SAC) between variables an autocorrelation term, calculated from neighbouring rates of spread using a kernel with a bandwidth of 10 km, was included as an independent variable. This SAC approach which was calculated from the dependent variable was preferred to the residual autocorrelation (RAC) method which is derived from the combination of predictor variables, as these changed with each of the multiple models that were developed. Both methods are well described and compared by Crase et al. (2012).
Because of the large number of variables available for inclusion in the model, a hierarchical stepwise approach was taken using six thematic models (co-variates grouped by theme are described in the supplementary information (S2)). The a priori confounders were not included in the thematic models but were forced into the final model. Principal components analysis was used to identify the components that contributed the most variance to the data within each thematic variable set. This was used to guide the selection of variables for inclusion in the modelling, rather than to create new variables from the components. This ensured that the model parameters could be easily interpreted. The variables with the strongest loading in each key component (preferentially those in component 1) were systematically added to a multivariable linear regression model with robust standard errors to allow for the presence of heteroscedasticity. The variance inflation factor for each variable in the thematic model was calculated using the “estat vif” command in Stata to assess whether collinearity was present in the model, and highly collinear variables (with a VIF > 10) were considered for exclusion. Beginning with this initial thematic model, a backward stepwise approach based on Akaike’s Information Criterion (AIC) was used to select the best fitting thematic model, with the least important variables (based on p values) being removed first (as recommended by Burnham and Anderson 2002). Following the approach taken by Pioz et al. (2012), models differing by less than two AIC points were considered to receive identical support from the data. In these instances the more parsimonious model was selected, unless there was good reason a priori for retaining a specific variable. Transformed variables were used where they improved the fit of the model.
The six thematic models were then sequentially added into one overall model starting with the one with the smallest root mean squared error (RMSE). The F test was used to determine whether each thematic set of variables contributed significantly to the overall model. If a p value greater than 0.05 was obtained from the F test, all variables in that group were removed. Finally, using the same backward stepwise approach based on AIC as applied to the thematic models, the overall model was developed using the remaining variables from the thematic models and the a priori confounders. Significant variables at the level of p ≤ 0.05 were retained in the model. Variables which had been removed were added back into the model one at a time and reconsidered for inclusion if they generated a p value less than 0.05. The likelihood ratio test was then used to determine whether the model including the previously dropped variables gave a better fit to the data than one excluding the variables.
The residuals of the model were assessed using the Breusch-Pagan/Cook-Weisberg test for heteroscedasticity (Breusch and Pagan 1979). This generated a p value of less than 0.001 indicating that there was sufficient evidence to reject the null hypothesis that the residuals were homogeneous, and thus it was appropriate to use robust standard errors.
2.4 GWR
GWR analyses were performed in R version 3.0.1 (2013-05-16) utilising the GWModel package for all geographically weighted analyses. R packages ‘RColorBrewer’ and ‘foreign’ were also used to display and export the analysis outputs. A statistical significance level of p ≤ 0.05 was used in all analyses. The methodology loosely followed a workflow for the GWModel package outlined by Gollini et al. (2013) and can be split into three steps: Geographically Weighted (GW) summary statistics, GW-Principal Component Analysis and GW Regression analysis.
For this work we utilised the geographic weighting in its simplest form applying a simple moving subset of records to the analysis. For each of the 2148 hexagons we selected its closest 215 hexagons (i.e. 10 % of the total data set) to run a localised model. The size of this subset is termed the bandwidth of the GWR analysis. A bandwidth of 215 was shown to fit natural regions within our irregular shaped study area, as well as showing no significant change in outputs during the GW PCA analysis when compared with the automatically calculated bandwidth of 348 generated by the bw.gwpca function of GWModel.
GW summary statistics such as plotting regionalised standard deviations (and GW inter-quartile ranges) were used to highlight areas of high variability for variables, and identify where application of GW analysis may warrant close scrutiny. The GW Principal Components Analysis (PCA) identified those variables which accounted for the greatest variation within the 10 % subset at each location, applying PCA in a similar way as it was utilised in the linear regression analysis.
The variable selection performed prior to the linear regression determined the variables offered to the GW analysis (as described in Table 2). Further variable selection was conducted to eliminate variables where significant regional co-linearity occurred before selecting the final model using a stepwise selection approach based on the AIC.
The rationale for using the same variables offered to the linear regression analysis as a starting point for variable selection for the GW analysis was that the original complete covariate dataset collated for the project contained too many variables to model. It included a number of alternative measures of similar environmental or farm characteristics meaning that strong relationships were found between similar groups of predictors. Additionally, while the GW analysis was intended to be used to assess regional variation in the drivers of the rate of spread of endemic bTB, the ultimate goal of the project was to provide information that could be practically used to inform national bTB control policies. It made sense to start with the variables that were also used to model the rate of spread at a national scale, since these were likely to be of most importance to policy makers, and to see how their significance and relationship varied in different areas using the GW approach.
A number of criteria were used to deal with multicollinearity; primarily estimates of correlation, complimented by GW PCA analysis to identify which of the variables accounted for the majority of the variance within the predictor database. Where further variable selection was required, decisions were based upon biological plausibility and the suitability of variables as targets for policy development for practical interventions.
The variables that were most influential on the rate of spread according to this model were mapped to illustrate the geographical variation in key variables. The number of hexagons where a variable had the most influence on the rate of spread (as determined by the size of the p value) was calculated for each variable.
2.5 BRT
Boosted regression trees (BRT) modelling was used to perform a preliminary validation of the GWR outputs and predictor variable selection. This method is now widely used in spatial modelling. It is an iterative machine learning technique based on regression trees, that attempts to minimise a loss function (deviance) and does not assume a defined starting distribution (Elith and Graham 2008). As such it is suited to the use of a large number of covariates and a large number of observations, and is particularly effective at accounting for non-linear relationships with the response variable. The models were offered the same covariates as the final GWR model and implemented using the VECMAP® software suite. Three area wide models were run, each for a specific region where GWR showed a different and consistent relationship with the most important predictor covariates, defined as those with the largest number of hexagons in which they were the most important variable according to the size of the p value.
3 Results
3.1 Linear regression with SAC term
The key components of each thematic set of variables that were identified by the principal components analysis are presented in Table 1. The variables that were included in the thematic models are presented in Table 2. Animal management factors such as movements, and testing appeared to be important, as did the presence of badgers. Variables such as clay or sandy soil and elevation were important in component 1 of the landscape characteristics set, and could be related to the suitability of the environment for badgers or cattle.
Four of the six thematic sets of variables were affected by missing values. The worst affected was the farm-level model where missing values in three variables (the percentage of permanent pasture within the convex hull that belonged to the farm (651 missing), movements to slaughter (158 missing) and the size of the primary market (581 missing)) reduced the number of observations by more than half from 2148 to 1042. Inspection of the missing values showed them to be randomly distributed within each variable so we proceeded with a smaller sample size.
The results of the F tests to assess the significance of the contribution of each set of variables to the final model are presented in Table 3. The tests indicated that the climate variables and the wildlife variables were not contributing significantly to the model so they were removed. The animal-level and bTB history and testing variables were bordering significance so were retained.
The final model is presented in Table 4. Nine variables were included in the final model, including the time period of spread which was split into eight categories and the spatial autocorrelation term. In addition to the autocorrelation term which had the strongest relationship, five variables were positively associated with the rate of spread. The large amount of variation explained by the inclusion of the autocorrelation term highlighted the need for a more geographically robust analysis to take account of the spatial dependence. The observed values of the rate of spread of endemic bTB versus the predicted values of the rate of spread from the final model are presented in Fig. 1.
3.2 GWR
GW summary statistics were generated for all variables selected as part of the linear regression to help understand the dataset. Important spatial characteristics of the data were identified through these statistics as demonstrated in Fig. 2 which shows substantial regional variation of the dependent variable.
Figure 3 illustrates the dominant variable contributing to the GW-PCA component 1, and shows clear regional variations in the most prominent variable. Further investigations proved this was indicative of the fact that the predictor dataset altered in structure in different regions.
The final GWR model utilising 16 out of the 26 variables taken from those identified by the global linear regression is described in Table 5. The automated selection procedure tested 276 possible models. The adjusted R-squared value of the selected model was 0.284. Strong evidence of spatial non-stationarity was obtained for all explanatory variables in explaining the rate of spread, except the number of cattle aged between 30 and 60 months, and the occurrence of OTF-W incidents in the year prior to spread which had weak evidence of non-stationarity. Figure 4 maps the most influential variable per hexagon (defined as that with the smallest p value), which demonstrates that a relatively small number of variables (~4) dominated the map, with distance to market being the most frequently identified predictor, followed by testing interval in the time period of spread, and the number of genotypes in a hexagon and its surrounding six hexagons. Figure 5 shows the total number of hexagons per variable where each variable is the most influential factor for rate of spread.
Distance to nearest market was the most frequently identified “winning variable” (i.e. the variable with the smallest p value) across the northern regions and southern regions but not in the central and eastern regions (Fig. 4). We also examined individual variables to see how their relationship with the rate of spread varied with location. This is presented in Fig. 6 for the distance to market variable, which showed marked variation in the slope of the relationship, being positive in the north and negative in the south. Examination of the data found no clear evidence of clustering of markets which might drive this pattern.
3.3 BRT
BRT models were produced for three distinct areas of spread in order to assess whether similar results to the GWR would be obtained. The areas assessed were the northern and southern areas where distance to nearest market was the most influential variable (though positively and negatively correlated, respectively), and the eastern area where distance to nearest market was not the most influential variable.
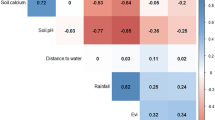
All three models produced R 2 values between the fitted function and observed rate of spread values in excess of 0.8 (p < 0.01) indicating good fits. The best predictors identified closely matched the GWR results: distance to nearest market was found to be the most influential variable in both the northern (Fig. 7a) and southern areas (Fig. 7b), being positively correlated with spread in the north and negatively correlated in the south, while it was not found to be the most influential variable in the eastern area (Fig. 7c).
Plots of the relationship between the BRT model predictions and actual data for the most influential variable from three separate BRT models: a model for the northern area where distance to nearest market was the most influential variable and was positively correlated with spread in the GWR, b model for the southern area where distance to nearest market was the most influential variable and was negatively correlated with spread in the GWR, and c model for the eastern area where distance to nearest market was not the most influential variable in the GWR. Percentages represent the relative importance of each variable in explaining the rate of spread of bTB within the respective models
4 Discussion
Bovine tuberculosis is a complicated multifactorial disease, and the factors associated with the disease in Great Britain have been explored in a number of studies (Reilly and Courtenay 2007; Carrique-Mas et al. 2008; Ramírez-Villaescusa et al. 2010; Johnston et al. 2011; Vial et al. 2011). These studies have focused on the association between risk factors and disease occurrence at the herd level, but many of the factors associated with bTB vary geographically and the contribution of regional differences in risk factors to the epidemiology of bTB is not well understood. The move to more regional bTB control policies in England and Wales in the last few years has created a need for more tailored interventions to fit the local disease situation. This approach is likely to improve the effectiveness of bTB controls, but requires policy makers to have an understanding of local drivers of disease. We have demonstrated that there is geographical variation in the predictors of endemic bTB and shown the utility of GWR in characterising this variation.
Many of the traditionally accepted predictors of bTB risk such as estimates of wildlife density have not been retained in the local level models developed here. The European badger (Meles meles) has long been known to be a wildlife reservoir of infection for bTB in cattle (Cheeseman et al. 1989). A number of variables could be considered proxies for badger suitability (e.g. elevation, soil type); although they could be equally proxies for cattle suitability. Predictors of the rate of spread of endemic bTB may be different to predictors of bTB persistence or incidence which are traditionally used as outcomes in risk factor investigations. This may imply that while badgers may be associated with the incidence or persistence of bTB in an area (Reilly and Courtenay 2007; Johnston et al. 2011), they may not be driving the spread of endemic bTB into new areas. It is also possible that causal factors at the edge of the endemic area will be different to those operating in core endemic areas, which may have previously been investigated in more detail.
An association that was consistently identified by both GWR and OLS linear regression was an increase in risk of flooding and an increase in the rate of spread. This may simply reflect the fact that many flood plains support grazing suitable for cattle and so be a focus of animal numbers. There may also be more mechanistic explanations. Seasonally wet soils on farms have been shown to protect against bTB in England and Wales (Johnston et al. 2011), though if flooding leads to enforced contact between herds there may be increased transmission and prevalence as observed in Tanzania (Cleaveland et al. 2005). Flooding might exacerbate environmental contamination of grazing pastures or drinking water with M. bovis and might also reduce badger food sources, forcing them to visit farm feed stores, so increasing contact between badgers and cattle (Garnett et al. 2002). Additionally, areas prone to flooding may harbour the helminth Fasciola hepatica (liver fluke), and concurrent infection of cattle with this parasite may reduce the sensitivity of the tuberculin skin test (Flynn et al. 2009; Ezenwa et al. 2010; Claridge et al. 2012).
The number of new OTF-W incidents in a hexagon in the year prior to the spread of endemic bTB was found to be negatively associated with the rate of spread in the OLS linear regression model. This variable was also identified in the GWR analysis. While an increase in the occurrence of OTF-W incidents suggests a greater infection pressure in the area which could be expected to exacerbate the spread of the disease, it may be that the observed reduction in the rate of spread in the national model is a result of the control measures put in place on these herds to limit the movement of animals, and possibly through farmers engaging in more protective measures as a result of having an incident.
Another factor identified by both models was the presence of multiple genotypes in a hexagon and its neighbours. Though this is likely to be indicative of multiple incidents, potentially from multiple sources, and thus the level of infection in an area, it could also be a reflection of the convergence of two or more genotype home ranges (Smith et al. 2006). However, any interpretation of genotype data should consider that the data is restricted to a single isolate per incident, so the true extent of the genotypes involved in an incident may not be known.
A significant association was observed between the rate of spread and lengths of boundaries of fragments of land shared with land from a different farm in the GWR model where it was the fourth most frequent “winning” variable. In the linear regression model a negative association was observed. This variable gives an indication of farm fragmentation and contiguity between farms so it was expected that this would be positively correlated with the rate of spread (Johnston et al. 2011).
Fitting a global model to bTB spread data is a complicated process, not least because of the spatial nature of many of the variables of interest, but also because spread into new areas can only be detected when testing takes place. There is also the problem that many of the variables are related in some way. The variable selection and modelling approaches used here were deliberately stringent in order to reduce collinearity and identify the most important variables, but it is likely that there are less important associations between variables which were not identified. For example, in the linear regression model there may have been associations between variables in different thematic models, e.g. rainfall in the climate data set and the risk of flooding in the landscape characteristics data set. Because of the large number of variables under consideration, the individual associations between all variables were not examined in great detail except where strong correlations were observed. Instead, more importance was placed on the contribution each variable made to the fit of the model rather than the magnitude of its effect.
Applying a traditional multivariable linear regression approach to a complex disease such as bTB is likely to lead to important spatial differences in relationships between variables being overlooked. Inclusion of the SAC term considerably increased the R squared value of the linear regression model, indicating that accounting for the spatial autocorrelation was important to explaining the variation in the rate of spread. The simpler SAC approach of using an autocorrelation term based on the dependant variable was selected for this study as multiple models were being developed, but it would be of interest to see what effect using a residual based SAC term (RAC) has on the final model. Crase et al. (2012) found the RAC approach, which only represents the portion of spatial structure in the dependant variable that is not explained by the explanatory variables, improved the accuracy of parameter estimates and identification of statistically significant variables. An alternative approach is that taken by Pioz et al. (2012) when modelling the spread of bluetongue virus in France. They used a simultaneous autoregressive model to account for spatial dependence.
The GW-PCA analysis proved useful when choosing between collinear variables during the model selection stage. The combination of PCA to reduce collinearity and subsequent GWR has been described for estimating crop water requirements in China (Wang et al. 2013). In that study, the principal components were used to create integrated independent variables for the GWR. In this study the PCA was used to guide the selection of variables for inclusion in the modelling rather than creating new variables. This enabled easier interpretation of the model parameters which was important as this was an exploratory analysis of variables rather than a purely predictive modelling exercise.
Even though GWR was used as a more robust method for accounting for spatial dependence, the final model explained a relatively small amount of the variation. The relationships identified were complex with regression coefficients switching between negative and positive values in different locations, which indicates that while focusing interventions on a particular risk factor might be beneficial in one area, it may actually be detrimental in another area. As with any regression method, they cannot prove causality, and other drivers may not have been retained as a result of the deliberately stringent method of covariate selection. The fact that the most important variables occur in blocks greater than half the GWR bandwidth rather than random scattering suggests the regional heterogeneity is real, with demonstrably different factors associated with the spread of bTB in different areas where spread has occurred, and it is this that is perhaps the most important finding of this study.
A preliminary BRT analysis was performed to see if it would produce comparable outputs to the GWR analysis. BRT produced statistically reliable area wide models, and identified similar regional relationships as GWR. While GWR worked well for identifying the most important variables per hexagon, the BRT produced better overall models and may be better suited than GWR to predicting the rate of spread beyond the study area if such analysis was desired.
This analysis has identified some clustering of potential risk factors that could be explored in more detail, but these do not easily translate into implementable interventions. An obvious area for further investigation would be a more detailed examination of those areas where flooding has been shown to be influential in order to understand the reasons for this association. Only by understanding the mechanism by which flooding may increase the rate of spread of endemic bTB can interventions be developed. Understanding the factors that affect the expansion of the area affected by endemic bTB is necessary to guide policy makers in the implementation of tailored local controls to halt the spread of the disease. The three methods used in this study have demonstrated the importance of accounting for spatial differences in risk factors for bTB, and have shown some consistency in the identification of certain factors. We have demonstrated that GWR is a useful approach for exploring bTB data and improves on least-squares linear regression by identifying regional differences in the factors associated with bTB spread. However, interpretation of these differences is difficult as relationships often varied spatially between negative and positive associations, and the approach does not lend itself to predictive models which are likely to be of more value to policy makers. Methods such as BRT may be more suited to such a task and we have demonstrated that GWR and BRT can produce comparable outputs. Finley (2011) concludes that other methods (such as Bayesian spatially-varying-coefficients (SVC)) may be better at predictive models but that GWR is less computationally intensive and is a useful tool for descriptive and exploratory data analysis, as demonstrated in this study.
In conclusion, this is the first attempt to explore the regional heterogeneity of factors associated with the spread of endemic bTB in England and Wales using GWR. Although a number of variables have been identified as significant in different locations in this study, the key message is that a complex regional pattern emerges which, though largely compatible with that identified from national analyses, should be able to help understand how national policies could be tailored to tackle bTB at a regional level.
References
Breusch TT, Pagan AR (1979) A simple test for heteroscedasticity and random coefficient variation. Econometrica 47:1287–1294
Brunsdon C, Fotheringham S, Charlton M (1998) Geographically weighted regression—modelling spatial non-stationarity. The Statistician 47:431–443
Brunton LA, Nicholson R, Ashton A, Alexander N, Wint W, Enticott G, Ward K, Broughan JM, Goodchild AV (2015) A novel approach to mapping and calculating the rate of spread of endemic bovine tuberculosis in England and Wales. Spatial and Spatio-temporal Epidemiology 13:41–50
Burnham KP, Anderson DR (2002) Model selection and multi-model inference: a practical information-theoretic approach. Springer-Verlag, New-York
Carrique-Mas JJ, Medley GF, Green LE (2008) Risks for bovine tuberculosis in British cattle farms restocked after the foot and mouth disease epidemic of 2001. Prev Vet Med 84:85–93
Cheeseman CL, Wilesmith JW, Stuart FA (1989) Tuberculosis: the disease and its epidemiology in the badger, a review. Epidemiol Infect 103:113–125
Claridge J, Diggle P, McCann CM, Mulcahy G, Flynn R, McNair J, Strain S, Welsh M, Baylis M, Williams DJL (2012) Fasciola hepatica is associated with the failure to detect bovine tuberculosis in dairy cattle. Nat Commun 3:853
Cleaveland S, Mlengeya T, Kazwala RR, Michel A, Kaare MT, Jones SL, Eblate E, Shirima GM, Packer C (2005) Tuberculosis in Tanzanian wildlife. J Wildlife Dis 41:446–453
Crase B, Liedloff AC, Wintle BA (2012) A new method for dealing with residual spatial autocorrelation in species distribution models. Ecography 35:879–888
Defra (2011) Bovine TB Eradication programme for England. https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/69443/pb13601-bovinetb-eradication-programme-110719.pdf. Accessed July 25, 2016
Defra (2014) The strategy for achieving officially bovine tuberculosis free status for England. https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/300447/pb14088-bovine-tb-strategy-140328.pdf. Accessed July 27, 2016
Elith J, Graham CH (2009) Do they? How do they? Why do they differ? On finding reasons for differing performances of species distribution models. Ecography 32:66–77
Ezenwa VO, Etienne RS, Luikart G, Beja-Pereira A, Jolles AE (2010) Hidden consequences of living in a wormy world: nematode-induced immune suppression facilitates tuberculosis invasion in African buffalo. Am Nat 176:613–624
Finley AO (2011) Comparing spatially-varying coefficients models for analysis of ecological data with non-stationary and anisotropic residual dependence. Methods Ecol Evol 2:143–154
Flynn RJ, Mulcahy G, Welsh M, Cassidy JP, Corbett D, Milligan C, Andersen P, Strain S, McNair J (2009) Co-infection of cattle with Fasciola hepatica and Mycobacterium bovis—immunological consequences. Transbound Emerg Dis 56:269–274
Fotheringham AS, Charlton ME, Brunsdon C (1998) Geographically weighted regression: a natural evolution of the expansion method for spatial data analysis. Environ Plan A 30:1905–1927
Garnett BT, Delahay RJ, Roper TJ (2002) Use of cattle farm resources by badgers (Meles meles) and risk of bovine tuberculosis (Mycobacterium bovis) transmission to cattle. Proc Biol Sci 269:1487–1491
Gollini I, Lu B, Charlton M, Brunsdon C, Harris P (2013) GWmodel: an R package for exploring spatial heterogeneity using geographically weighted models; arXiv:1306.0413v1 [stat.AP] 3 Jun 2013
Jaber SM, Ibbini JH, Hijjawi NS, Amdar NM, Huwail MJ, Al-Aboud K (2013) Exploring recent spatial patterns of cutaneous leishmaniasis and their associations with climate in some countries of the Middle East using geographical information systems. Geospatial Health 8:143–158
Johnston WT, Vial F, Gettinby G, Bourne FJ, Clifton-Hadley RS, Cox DR, Crea P, Donnelly CA, McInerey JP, Mitchell AP, Morrison WI, Woodroffe R (2011) Herd-level risk factors of bovine tuberculosis in England and Wales after the 2001 foot-and-mouth disease epidemic. Int J Infect Dis 15:833–840
Lawes JR, Harris KA, Brouwer A, Broughan JM, Smith NH, Upton PA (2016) Bovine TB surveillance in Great Britain in 2014. Veterinary Record 178:310–315
Lennon JJ (2000) Red-shifts and red herrings in geographical ecology. Ecography 23:101–113
Liao FHF & Wei YHD (2014) Modeling determinants of urban growth in Dongguan, China: a spatial logistic approach. Stoch Environ Res Risk Assess 28:801–816
Moustakas A, Evans MR (2016) Regional and temporal characteristics of bovine tuberculosis in Great Britain. Stoch Environ Res Risk Assess 30:989–1003
Pioz MH, Guis H, Crespin L, Gay E, Calavas D, Durand B (2012) Why did Bluetongue spread the way It did? Environmental factors influencing the velocity of Bluetongue virus serotype 8 epizootic wave in France. PLoS ONE 7:e43360
Ramírez-Villaescusa AM, Medley GF, Mason S, Green LE (2010) Risk factors for herd breakdown with bovine tuberculosis in 148 cattle herds in the south west of England. Prev Vet Med 95:224–230
Reilly LA, Courtenay O (2007) Husbandry practices, badger sett density and habitat composition as risk factors for transient and persistent bovine tuberculosis on UK cattle farms. Prev Vet Med 80:129–142
Smith NH, Gordon SV, de la Rua-Domenech R, Clifton-Hadley RS, Hewinson RG (2006) Bottlenecks and broomsticks: the molecular evolution of Mycobacterium bovis. Nat Rev Micro 4:670–681
Vial F, Johnston WT, Donnelly CA (2011) Local cattle and badger populations affect the risk of confirmed tuberculosis in British cattle herds. PLoS ONE e18058
Wang JL, Kang SZ, Sun JS, Chen ZF (2013) Estimation of crop water requirement based on principal component analysis and geographically weighted regression. Chinese Sci Bull 58:3371–3379
Wang C, Du S, Wen J, Zhang M, Gu H, Shi Y, Xu H (2016) Analyzing explanatory factors of urban pluvial floods in Shanghai using geographcially weighted regression. Stoch Environ Res Risk Assess. doi:10.1007/s00477-016-1242-6
Welsh Government (2016) Intensive action area. http://gov.wales/topics/environmentcountryside/ahw/disease/bovinetuberculosis/intensive-action-area/?lang=en. Accessed July 25, 2016
Acknowledgments
The authors wish to thank the following colleagues who have commented on the analysis or reviewed the final manuscript: Dr Sara Downs, Dr Tony Goodchild, Dr Jessica Parry, Jane Gibbens, Prof. Dirk Pfeiffer and Prof. Glyn Hewinson. The authors are grateful to Rachel Eglin, Jemma Aston and Rosie Sallis for management of the project, and to Rose Nicholson for data collation. This work was funded by the UK Government Department for Environment, Food and Rural Affairs under research project SE3045.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Brunton, L.A., Alexander, N., Wint, W. et al. Using geographically weighted regression to explore the spatially heterogeneous spread of bovine tuberculosis in England and Wales. Stoch Environ Res Risk Assess 31, 339–352 (2017). https://doi.org/10.1007/s00477-016-1320-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00477-016-1320-9