Abstract
Key message
The Neotropical tree Parkia panurensis shows a spatial genetic structure from the seed to the adult stage that is most likely the outcome of the seed dispersal provided by primates.
Abstract
Seed dispersal and pollination determine the gene flow within plant populations. In addition, seed dispersal creates the template for subsequent stages of plant recruitment. Therefore, the question arises whether and how seed dispersal affects the spatial genetic structure (SGS) of plant populations. In this study, we used microsatellites to analyse the SGS of the Neotropical tree Parkia panurensis (Fabaceae). This plant species is a major food resource for primates and its seeds are mainly dispersed by primates. Seeds were collected during behavioural observations of a tamarin mixed-species troop in north-eastern Peru. Additionally, leaf samples of juveniles and of adults trees of this species were collected throughout the home range of the tamarin troop. A significant SGS for embryos (located within the dispersed seeds) and for non-reproductive plants are found up to a distance of 300 m. This matches the distance within which most seeds are dispersed. In the adult stage, the scale of a significant SGS is reduced to 100 m. While we cannot explain this scale reduction, our study provides the first evidence that primate seed dispersal does influence the SGS of a tropical tree species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The seed shadow created through primary seed dispersal provides the spatial template for subsequent processes like secondary seed dispersal, seedling establishment, etc., that take place during the “seed dispersal loop” (Wang and Smith 2002) and thus may influence the population dynamics of a plant species (Nathan and Muller-Landau 2000; Jordano and Godoy 2002). For zoochorously dispersed seeds, the behaviour and particularly the movement patterns of frugivores influence the shape and dimensions of the seed shadow (Jordano et al. 2007; Karubian et al. 2012; Russo et al. 2006). This template “defines what is possible for the seedling recruitment in a given area” (Howe and Miriti 2004, p. 654), but the question arises whether and how this initial template also contributes to the recruitment of new reproductive adults into the population (Schupp and Jordano 2011). Differential survival of seedlings and growing trees can stabilise, weaken or erase the initial spatial pattern, depending on the selective pressures acting during the different phases of regeneration (Jordano and Herrera 1995). On the genetic level, this turns into the question of whether and how the spatial genetic structure (SGS) of the seed shadow transfers into an SGS in subsequent life stages.
Some studies have provided evidence that seed dispersal by frugivores may affect the SGS of plants. A small-scale SGS in adults of the bird-dispersed Psychotria officinalis and seedlings of the ant-dispersed Globba lancangensis has been related to the short dispersal distances created by the respective dispersal agents (Loiselle et al. 1995; Zhou et al. 2007). “Destination-based” dispersal by umbrellabirds of seeds of Oenocarpus bataua into lek arenas results in a lower SGS amongst seedlings compared to areas where seeds are dispersed by other agents (Karubian et al. 2010). A multi-stage comparison of the SGS of Attalea phalerata showed a strong SGS amongst seedlings, but a much lower SGS at two juvenile stages and at the adult stage (Choo et al. 2012), suggesting that the initially strong effect of seed dispersal does not continue into later stages.
Ideally, analyses of SGS should also include the embryo stage, since the seed shadow provides the initial template. However, quantification of the seed shadow is an inherently difficult task. It is usually impossible or very difficult to observationally follow the fate of individual seeds between removal from the source tree by a frugivore till deposition via defecation or regurgitation. Molecular genetic methods allow overcoming these difficulties by assigning seeds to source trees through comparing the DNA from potential mother trees and from seed coats of dispersed seeds (Godoy and Jordano 2001; Ziegenhagen et al. 2003). In a previous study, we quantified the seed shadow of the Neotropical legume Parkia panurensis created by two sympatrically living small New World primates, black-fronted saddle-back tamarins, Saguinus nigrifrons (previously Saguinus fuscicollis nigrifrons), and moustached tamarins, Saguinus mystax, through DNA fingerprinting and observational data (Heymann et al. 2012). We focussed on P. panurensis as the pod gum of this species a major food resource of these primates (Knogge and Heymann 2003; see also Peres 2000 for Parkia pod gums as a key resource for mammals and birds in Amazonian forests), and because the tamarins are the only known seed dispersers of P. panurensis at our study site in Peruvian Amazonia. This provides the opportunity to study the effect of seed dispersal on the SGS in a simplified system. Therefore, in this study we analysed the SGS of P. panurensis at three life stages (embryos, juveniles, adults) and ask whether this is affected by tamarin seed dispersal.
Methods
Study site and study species
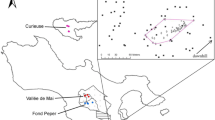
The study was carried out at the Estación Biológica Quebrada Blanco (EBQB), located at 4°21′S 73°09′W in the Amazon lowlands of north-eastern Peru. The site is characterized by rainforest of the bosque de altura (terra firme) type. For further details of the site see Heymann (1995).
Parkia panurensis is member of a pantropical tree genus of the family Fabaceae. Distributed in central and western Amazonia, it is mainly growing in terra firme forests to heights of up to 35 m (Hopkins 1986). Wood maybe used for lumber (Duke and Vasquez 1994) and the bark exudate has medical uses (Corrêa 1909 quoted in Hopkins 1986). At our study site, fruiting has been recorded variably between May and October. Fruits are falcate pods (length up to 35 cm) with 16–23 seeds (Hopkins 1986; E.W. Heymann, personal observation). These seeds (13–18.5 mm long, 6–12 mm wide, 5–8.5 mm high; Hopkins 1986) are surrounded by an edible gum. Seeds of Neotropical Parkia are mainly dispersed by primates (Ateles sp., Lagothrix sp., Saguinus sp.) and terrestrial rodents (Culot 2009; Dew 2008; Hopkins and Hopkins 1983; Knogge and Heymann 2003; Peres 2000; van Roosmalen 1985). Bruchid beetles, ants, large parrots, primates (Cebus apella, Pithecia albicans, Cacajao calvus ucayalii), terrestrial ungulates and rodents are known predators of Parkia seeds (Bowler and Bodmer 2011; Feldmann et al. 2000; Hopkins and Hopkins 1983; E.W. Heymann, personal observation). At our study site, S. nigrifrons and S. mystax are the only frugivores that have been observed to disperse Parkia seeds during focal tree observations. These two primate species live in groups of 3–9 individuals and form stable mixed-species troops (Heymann and Buchanan-Smith 2000). They spent most of their time together (Heymann 1990) and jointly move through home ranges that vary in size between ca. 30–60 ha. Their diet is mainly composed of fruit pulp, insects and exudates, complemented with nectar and small vertebrates (Garber 1988; Knogge and Heymann 2003; Peres 1993). Tamarins disperse the seeds of a large diversity of plant species (Culot 2009; Garber 1986; Knogge and Heymann 2003) to distances of up to 700 m (Heymann et al. unpubl. data). The two species neither vary in the spectrum of dispersed plant species (Culot 2009; Knogge and Heymann 2003) nor in seed dispersal curves nor mean dispersal distances (Heymann et al. unpubl. data). While the number of seeds in tamarin defecations ranges from 1 to >4,800 (in the case of very small seeds like, e.g. from Cecropia, Ficus and Marcgravia), the mean is 1.2 seeds per defecation (Knogge and Heymann 2003). Dispersed seeds remain viable after passage through the tamarins’ guts (Knogge et al. 2003). Tamarins disperse seeds from primary forest into anthropogenic caused secondary forest and thus contribute to natural regeneration (Culot et al. 2010).
At EBQB, tamarin mixed-species troops are routinely monitored on a monthly base or observed in more detail for 5–10 days per month during specific research projects. Therefore, information on home-range location and size (generated through GPS recordings and calculated as minimum convex polygons) is constantly updated.
Sample collection
For genetic analyses, we collected samples from three different life stages of P. panurensis. These stages are embryos (embedded in the seeds), juveniles, and adult. Juveniles were defined as individuals with a height >1.3 m and a dbh <20 cm, adults as individuals with a dbh >20 cm. This criterion is based on the long-term observation that tamarins never feed in smaller P. panurensis trees, suggesting these do not produce fruits. Seeds were collected immediately after dispersal and stored in a saturated NaCl solution until DNA extraction. Intact P. panurensis seeds are generally defecated singly with little or no faecal matter adhering.
To sample juveniles, we randomly overlaid a 50 m × 50 m grid over a map of the home range of our tamarin troop. Intersections of this grid were taken as the points of origin for the point quarter method (Krebs 1999); from these points we searched for juveniles within each quarter at a maximum distance of 25 m. For the adult stage, we completed a full inventory within the home range of the tamarin troop. We collected leave samples from juveniles and adults. These were dried and stored in plastic bags on silica gel until DNA extraction.
Finally, the geographical position of all sampled P. panurensis individuals and all sites of seed collection from tamarin defecations were recorded with a Garmin GPSMap® 76CSx.
Molecular genetic methods and analyses of genetic variation
We used nine highly polymorphic nuclear microsatellite loci for genotyping (Luettmann et al. 2010). PCR and genotyping methods are reported in detail in Heymann et al. (2012).
Standard genetic diversity parameters were assessed such as A, A e , H o , H e , F IS . The calculations were performed with the computer programs GenAlEx 6.4 (Peakall and Smouse 2006) and FSTAT (Goudet 2001). In addition, the average number of private alleles was measured. To correct for variation of sample size, the average allelic richness (A R31 ) over all loci was calculated, using the rarefaction method described by El Mousadik and Petit (1996).
In order to detect genetic differentiation among the different stages of P. panurensis, global and pairwise D est (Chao et al. 2008) were measured using SMOGD (Crawford 2010). For calculating confidence limits we used the R-package “diveRsity” (Keenan et al. 2013).
Parentage and paternity analysis were conducted using the software CERVUS version 3.03 (Kalinowski et al. 2007). In all analyses the proportion of fathers sampled was set to 0.35.
Spatial genetic analyses
Our analyses are based on 90 embryos taken from dispersed seeds, 90 juveniles, and 31 adults. To test for significant autocorrelation, we calculated the multivariate Mantel correlogram for each stage using the method described by Oden and Sokal (1986). This procedure was first applied to spatial genetic structures by Matesanz et al. (2011). We determined genetic distances using the inter-individual distance from the R-package “gstudio” and the function ‘mpmcorrelogram’ from the package “mpmcorrelogram”. For identifying statistical significance we employed a progressive Bonferroni correction. Correlation between matrices was calculated using the Spearman rank correlation coefficient. Since the genetic distance matrix increases its value with genetic distance, negative values in the Mantel correlogram correspond to a positive autocorrelation (Legendre and Legendre 2012).
Results
Genetic variation of Parkia panurensis and parentage analyses
Genetic diversity as measured by the absolute number of alleles measured over all loci (A), the average number of effective alleles measured over all loci (A e ) and allelic richness (A R31 ) is similarly high for juveniles as well as for adults, but much lower in embryos (Table 1).With regard to the average and absolute number of private alleles, embryos and adults were similar, but non-reproductive trees had much higher numbers.
Global D est among the different P. panurensis age classes was not significant (D est = 0.0278). Embryos differed slightly from juveniles and adults (0.047 and 0.042), but the latter two were not different from each other (−0.013; Table 2).
The 90 embryos originate from five different mothers. 60 out of 90 embryos could be assigned to eight different putative fathers. Because P. panurensis has bisexual flowers and is monoecious, five trees are assigned both as fathers and as mothers. From the possible combinations of parents only 14 different male–female pairs have been found. There was no case of selfing.
Both parents could be identified for 19 out of 90 juveniles. They descend from 13 different reproductive adults and 12 male–female pairs. In only one case selfing is the most probable combination.
Spatial genetic structure in Parkia panurensis
In all three stages, a significant positive SGS was detected. For embryos and juveniles, the SGS extends up to 300 m, for adults up to 100 m (Fig. 1). Additionally, there is a significant negative SGS in embryos at 400 and 500 m.
Multivariate Mantel correlogram of the three different Parkia panurensis life stages. a embryos, b juveniles, c adults. Significant values are indicated by filled squares. Values of the x-axis correspond to the upper limit of the respective distance class. Note that negative values correspond to a positive spatial genetic structure (Legendre and Legendre 2012)
Discussion
Our study differs from most other studies that examined the SGS of tropical trees by including the embryo stage. The rationale for including the embryo stage is obvious: their spatial distribution, created through seed dispersal, is the template on which all steps of the “seed dispersal loop” build upon. Starting with the seedling stage ignores this initial template and requires inference about the seed dispersal process and its influence on SGS (Choo et al. 2012; Hardesty et al. 2005). The fact that we could follow the seed dispersal vectors—the two tamarin species—at close range and collect their defecation with dispersed seeds allowed for the inclusion of the embryo stage.
We lumped juveniles of different size and trees with height >130 cm and dbh <20 cm into a single stage. This leads to the inclusion of individuals of different ages who emerged from seeds dispersed in different reproductive seasons. Depending on inter-individual variation in fertility (i.e. whether or not the same adults dominate seed input over subsequent seasons), this may or may not influence the strength of the SGS. This argument would also hold true for the adult stage which includes individuals of different ages (and thus of different reproductive seasons). However, we are not aware that any study on the SGS of tropical trees has controlled for the age of adults (which is unlikely to be known in most cases).
Our analyses revealed a significant SGS at the three different life stages, but the spatial scale of the SGS as measured by the significance of the Mantel correlation coefficients is much smaller in adults (100 m) than in embryos and juveniles (300 m). Tamarins disperse Parkia seeds up to 700 m (Heymann et al. 2012), which corresponds to the diameter of a typical home range. However, the majority of seeds are dispersed within a distance of 300 m from the source tree (85 % of observationally determined and 88 % of genetically determined dispersal events within 300 m; see Fig. 1 in Heymann et al. 2012). This is a consistent pattern: in independent studies at EBQB in 1994/1995 and 2006, 89 and 85 %, respectively, of dispersal events were within 300 m (Heymann et al. unpubl. data).
Due to density-dependent mortality which is likely to be highest in the embryo stage, one could expect a demographic thinning from the embryo to the juvenile stage. Consequently, one could also expect a reduction of the scale of SGS from embryos to juveniles. However, we do not find such a reduction and envision the following scenario to account for this: Our juvenile stage includes individuals of a wide range of sizes. Thus, they are most likely the result of dispersal events in different years. Tamarins use the same feeding trees over decades (a Parkia tree first seen to be used in 1985 was still in use in 2011; E.W. Heymann, personal observation). They also use the same resting and sleeping trees over prolonged periods. Resting and sleeping trees are places of increased defecation, and consequently, areas around these trees become foci of seed dispersal and seedling recruitment (Muñoz Lazo et al. 2011; Heymann et al. unpubl. data). However, tamarins rarely disperse more than two Parkia seeds in a single defecation, and tamarin groups are small (5–6 individuals on average). Thus, even the repeated use of the same resting and sleeping sites within a short period is unlikely to produce seed and seedling densities that could result in high density-dependent mortality. Over prolonged periods, repeated seed dispersal to areas around the same resting and sleeping sites would lead to the accumulation of juveniles of different ages that stem from a limited pool of source trees. However, this scenario does not explain why the scale of the significant SGS is reduced from the juvenile stage to the adult stage. Demographic thinning (Hardesty et al. 2005) is likely to continue throughout the juvenile stage into the adult stage. Fitted distributions (Weibull, Gamma) of dispersal distances—both genetically and observationally determined–indicate maximum seed dispersal in a radius of around 100 m around source trees (Heymann et al. 2012). Therefore, even after demographic thinning throughout different stages, enough individuals may survive into the adult stage to create a significant SGS at this scale.
The reduction in scale of the SGS is in line with findings from other studies. The scale, but not the strength of the SGS declines from the smallest to the largest size class in the zoochorously dispersed Simarouba amara (Simaroubaceae) (Hardesty et al. 2005). Similarly, in the autochorous Alseis blackiana (Rubiaceae) and the anemochorous Platypodium elegans (Fabaceae), a positive SGS exists in juveniles up to 30 and 100 m, respectively, but is lost in adults, presumably due to random mortality during recruitment, respectively (Hamrick et al. 1993). Both the scale and the strength of SGS decline from seedlings to the adult stage in Attalea phalerata (Arecaceae) (Choo et al. 2012). However, it is also obvious from other studies that long-term variation and stochastic effects may influence the findings on SGS. While Degen et al. (2001) reported a significant SGS up to 300 m in adult Carapa procera (Meliaceae), an earlier study at the same site did find an SGS neither in adults nor in seeds of this species (Doligez and Joly 1997). Loiselle et al. (1995, p. 1424) attributed the “anomalous finding” of a significant positive SGS at 90 m distance and a significant negative at 105 m to chance.
Despite a significant SGS, values of A, A e and F IS indicate that there is no inbreeding. Therefore, gene flow from outside the sampled population takes place, most likely through pollen dispersal. The fact that most fathers seem to stand outside the sampled population supports this contention. Most Neotropical Parkia species are pollinated by bats, mainly phyllostomids (Hopkins 1984). Species of this bat family may range over distances that are larger than the diameter of our tamarin study troop’s home range (Meyer et al. 2005) and can therefore transfer pollen between individuals from a larger population than our sample.
The majority of primates are highly or partially frugivorous and contribute substantially to seed dispersal in tropical habitats (Chapman and Russo 2007; Lambert and Garber 1998; Peres and van Roosmalen 2002). Despite a large number of studies on primate seed dispersal, the question of whether and how this affects the SGS of dispersed plant species has hardly been addressed. Only de Moraes et al. (2004) attributed a significant SGS for adult Cryptocarya moschata (Lauraceae) up to 150 m to seed dispersal by muriquis, Brachyteles arachnoides. However, they did not provide information on the patterns and distances of seed dispersal by these primates. Our study is therefore the first to provide evidence that seed dispersal by two primate species actually does influence the SGS of one of their major food resources.
References
Bowler M, Bodmer RE (2011) Diet and food choice in Peruvian red uakaris (Cacajao calvus ucayalii): selective or opportunistic seed predation? Int J Primatol 32:1109–1122
Chao A, Jost L, Chiang SC, Jiang Y, Chazdon RL (2008) A two-stage probabilistic approach to multiple-community similarity indices. Biometrics 64:1178–1186
Chapman CA, Russo SE (2007) Primate seed dispersal. Linking behavioral ecology with forest community structure. In: Campbell CJ, Fuentes A, Mackinnon KC, Panger M, Bearder SK (eds) Primates in perspective. Oxford University Press, Oxford, pp 510–525
Choo J, Juenger TE, Simpson BB (2012) Consequences of frugivore-mediated seed dispersal for the spatial and genetic structures of a neotropical palm. Mol Ecol 21:1019–1031
Crawford NG (2010) SMOGD: software for the measurement of genetic diversity. Mol Ecol Resour 10:556–557
Culot L (2009) Primary seed dispersal by two sympatric species of tamarins, Saguinus fuscicollis and Saguinus mystax, and post-dispersal seed fate. Doctoral dissertation, Université de Liège, Belgium
Culot L, Muñoz Lazo FJJ, Poncin P, Huynen MC, Heymann EW (2010) Seasonal variation in seed dispersal by tamarins alters seed rain in a secondary rainforest. Int J Primatol 31:553–569
de Moraes PLR, Monteiro RV, Vencovsky R (2004) Estructura genética intrapopulacional em Cryptocarya moschata Nees (Lauraceae). Rev bras Bot 27:475–487
Degen B, Caron H, Bandou E, Maggia L, Chevallier MH, Leveau A, Kremer A (2001) Fine scale spatial genetic structure of eight tropical tree species as analysed by RAPDs. Heredity 87:497–507
Dew JL (2008) Spider monkeys as seed dispersers. In: Campbell CJ (ed) Spider monkeys. Behavior, ecology and evolution of the genus Ateles. Cambridge University Press, Cambridge, pp 155–182
Doligez A, Joly HI (1997) Genetic diversity and spatial structure within a natural stand of a tropical forest tree species, Carapa procera (Meliaceae), in French Guiana. Heredity 79:72–82
Duke JA, Vasquez R (1994) Amazonian ethnobotanical dictionary. CRC Press, Boca Raton
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (L.) Skeels) endemic to Morocco. Theor Appl Genet 92:832–839
Feldmann M, Verhaagh M, Heymann EW (2000) Sericomyrmex ants as seed predators. Ecotropica 6:207–209
Garber PA (1986) The ecology of seed dispersal in two species of callitrichid primates (Saguinus mystax and Saguinus fuscicollis). Am J Primatol 10:155–170
Garber PA (1988) Diet, foraging patterns, and resource defense in a mixed-species troop of Saguinus mystax and Saguinus fuscicollis in Amazonian Peru. Behaviour 105:18–34
Godoy JA, Jordano P (2001) Seed dispersal by animals: exact identification of source trees with endocarp DNA microsatellites. Mol Ecol 10:2275–2283
Goudet, J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available from http://www.unil.ch/izea/softwares/fstat.html
Hamrick JL, Murawski DA, Nason JD (1993) The influence of seed dispersal mechanisms on the genetic structure of tropical tree populations. In: Fleming TH, Estrada A (eds) Frugivory and seed dispersal: evolutionary and ecological aspects. Kluwer, Dordrecht, pp 281–297
Hardesty BD, Dick CW, Kremer A, Hubbell S, Bermingham E (2005) Spatial genetic structure of Simarouba amara Aubl. (Simaroubaceae), a dioecious, animal-dispersed Neotropical tree, on Barro Colorado Island. Panama Heredity 95:290–297
Heymann EW (1990) Interspecific relations in a mixed-species troop of moustached tamarins, Saguinus mystax, and saddle-back tamarins, Saguinus fuscicollis (Platyrrhini: Callitrichidae), at the Río Blanco, Peruvian Amazonia. Am J Primatol 21:115–127
Heymann EW (1995) Sleeping habits of tamarins, Saguinus mystax and Saguinus fuscicollis (Mammalia; Primates; Callitrichidae), in north-eastern Peru. J Zool, Lond 237:211–226
Heymann EW, Buchanan-Smith HM (2000) The behavioural ecology of mixed-species troops of callitrichine primates. Biol Rev 75:169–190
Heymann EW, Luettmann K, Michalczyk IM, PinedoSaboya PP, Ziegenhagen B, Bialozyt R (2012) DNA fingerprinting validates seed dispersal curves from observational studies in the Neotropical legume Parkia. PLoS One 7:e35480
Hopkins HC (1984) Floral biology and pollination ecology of the neotropical species of Parkia. J Ecol 72:1–23
Hopkins HC (1986) Parkia (Leguminosae, Mimosoideae). Flora Neotrop 43:1–124
Hopkins HC, Hopkins MJG (1983) Fruit and seed biology of the neotropical species of Parkia. In: Sutton SL, Whitmore TC, Chadwick AC (eds) Tropical rain forest: ecology and management, 2nd edn. Blackwell Scientific Press, Oxford, pp 197–209
Howe HF, Miriti MN (2004) When seed dispersal matters. Bioscience 54:651–660
Jordano P, Godoy JA (2002) Frugivore-generated seed shadows: a landscape view of demographic and genetic effects. In: Levey DJ, Silva WR, Galetti M (eds) Seed dispersal and frugivory: ecology, evolution and conservation. CABI Publishing, Wallingford, pp 305–321
Jordano P, Herrera CM (1995) Shuffling the offspring: uncoupling and spatial discordance of multiple stages in vertebrate seed dispersal. Écoscience 2:230–237
Jordano P, García C, Godoy JA, García-Castaño JL (2007) Differential contributions of frugivores to complex seed dispersal patterns. Proc Nat Acad Sci 104:3278–3282
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
Karubian J, Sork VL, Roorda T, Durães R, Smith TB (2010) Destination-based seed dispersal homogenizes genetic structure of a tropical palm. Mol Ecol 19:1745–1753
Karubian J, Durães R, Storey JL, Smith TB (2012) Mating behavior drives seed dispersal by the long-wattled umbrellabird Cephalopterus penduliger. Biotropica 44:689–698
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodoehl PA (2013) diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788
Knogge C, Heymann EW (2003) Seed dispersal by sympatric tamarins, Saguinus mystax and Saguinus fuscicollis: diversity and characteristics of plant species. Folia Primatol 74:33–47
Knogge C, Tirado Herrera ER, Heymann EW (2003) Effect of passage through tamarin guts on the germination potential of dispersed seeds. Int J Primatol 24:1121–1128
Krebs CJ (1999) Ecological methodology, 2nd edn. Addison Wesley Longman, Menlo Park
Lambert JE, Garber PA (1998) Evolutionary and ecological implications of primate seed dispersal. Am J Primatol 45:9–28
Legendre P, Legendre L (2012) Numerical ecology, 3rd edn. Elsevier, Amsterdam
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425
Luettmann K, Michalczyk IM, Mengele C, Ziegenhagen B, Heymann EW, Pinedo Saboya PP, Bialozyt R (2010) Characterization of nuclear microsatellite loci in the Neotropical tree Parkia panurensis (Fabaceae). Am J Bot 97:e34–e36
Matesanz S, Gimeno TE, de la Cruz M, Escudero A, Valladares F (2011) Competition may explain the fine-scale spatial patterns and genetic structure of two co-occurring plant congeners. J Ecol 99:838–848
Meyer CFJ, Weinbeer M, Kalko EKV (2005) Home-range size and spacing patterns of Macrophyllum macrophyllum (Phyllostomidae) foraging over water. J Mamm 86:587–598
Muñoz Lazo FJJ, Culot L, Huynen M-C, Heymann EW (2011) Resting sites and seed dispersal by a mixed-species group of tamarins, Saguinus fuscicollis and Saguinus mystax, in the Amazonian rainforest of Peru. Int J Primatol 32:223–237
Nathan R, Muller-Landau HC (2000) Spatial patterns of seed dispersal, their determinants and consequences for recruitment. Trends Ecol Evol 15:278–285
Oden NL, Sokal RR (1986) Directional autocorrelation: an extension of spatial correlograms to two dimensions. Syst Zool 35:608–617
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Australian National University, Canberra
Peres CA (1993) Diet and feeding ecology of saddle-back (Saguinus fuscicollis) and moustached (S. mystax) tamarins in Amazonian terra firme forest. J Zool Lond 230:567–592
Peres CA (2000) Identifying keystone plant resources in tropical forests: the case of gums from Parkia pods. J Trop Ecol 16:287–317
Peres CA, van Roosmalen M (2002) Primate frugivory in two species-rich neotropical forests: implications for the demography of large-seeded plants in overhunted areas. In: Levey DJ, Silva WR, Galetti M (eds) Seed dispersal and frugivory: ecology, evolution and conservation. CABI Publishing, Wallingford, pp 407–421
Russo SE, Portnoy S, Augspurger CK (2006) Incorporating animal behavior into seed dispersal models: implications for seed shadows. Ecology 87:3160–3174
Schupp EW, Jordano P (2011) The full path of Janzen-Connell effects: genetic tracking of seeds to adult plant recruitment. Mol Ecol 20:3953–3955
van Roosmalen MGM (1985) Fruits of the Guianan flora. Institute of Systematic Botany, Utrecht
Wang BC, Smith TB (2002) Closing the seed dispersal loop. Trends Ecol Evol 17:379–385
Zhou H, Chen J, Chen F (2007) Ant-mediated seed dispersal contributes to the local spatial pattern and genetic structure of Globba lancangensis (Zingiberaceae). J Hered 98:317–324
Ziegenhagen B, Liepelt S, Kuhlenkamp V, Fladung M (2003) Molecular identification of individual oak and fir trees from maternal tissues of their fruits and seeds. Trees 17:345–350
Author contributions statement
EWH, BZ and RB conceived and designed the research project: KL and PPPS performed the field work; KL and IMM performed the genetic analyses; RB, IMM and EWH analysed the data; EWH, RB, IMM and BZ wrote the manuscript.
Acknowledgments
We are grateful to the following people and organizations: Camilo Flores Amasifuén for excellent field assistance; Christina Mengel for intensive lab assistance; the Instituto Nacional de Recursos Naturales (INRENA) in Lima for granting the research permit (no. 1062007-260 INRENA-IFFS-DCB); the Deutsche Forschungsgemeinschaft for funding (grants no. HE 1870/15-1 and HE 1870/15-2); and last but not least, the anonymous reviewer for helpful and constructive comments.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Masaka.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Bialozyt, R., Luettmann, K., Michalczyk, I.M. et al. Primate seed dispersal leaves spatial genetic imprint throughout subsequent life stages of the Neotropical tree Parkia panurensis . Trees 28, 1569–1575 (2014). https://doi.org/10.1007/s00468-014-1061-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-014-1061-7