Abstract
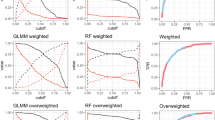
Information on ecological systems often comes from diverse sources with varied levels of complexity, bias, and uncertainty. Accordingly, analytical techniques continue to evolve that address these challenges to reveal the characteristics of ecological systems and inform conservation actions. We applied multiple statistical learning algorithms (i.e., machine learning) with a range of information sources including fish tracking data, environmental data, and visual surveys to identify potential spawning aggregation sites for a marine fish species, permit (Trachinotus falcatus), in the Florida Keys. Recognizing the potential complementarity and some level of uncertainty in each information source, we applied supervised (classic and conditional random forests; RF) and unsupervised (fuzzy k-means; FKM) algorithms. The two RF models had similar predictive performance, but generated different predictor variable importance structures and spawning site predictions. Unsupervised clustering using FKM identified unique site groupings that were similar to the likely spawning sites identified with RF. The conservation of aggregate spawning fish species depends heavily on the protection of key spawning sites; many of these potential sites were identified here for permit in the Florida Keys, which consisted of relatively deep-water natural and artificial reefs with high mean permit residency periods. The application of multiple machine learning algorithms enabled the integration of diverse information sources to develop models of an ecological system. Faced with increasingly complex and diverse data sources, ecologists, and conservation practitioners should find increasing value in machine learning algorithms, which we discuss here and provide resources to increase accessibility.






Similar content being viewed by others
References
Adams AJ, Blewett DA (2004) Spatial patterns of estuarine habitat type use and temporal patterns in abundance of juvenile permit, trachinotus falcatus, in Charlotte Harbor, Florida. Gulf Caribb Res 16:129–139. https://doi.org/10.18785/gcr.1602.01
Adams AJ, Cooke SJ (2015) Advancing the science and management of flats fisheries for bonefish, tarpon, and permit. Environ Biol Fishes 98:2123–2131. https://doi.org/10.1007/s10641-015-0446-9
Adams AJ, Wolfe RK, Kellison GT, Victor BC (2006) Patterns of juvenile habitat use and seasonality of settlement by permit, Trachinotus falcatus. Environ Biol Fishes 75:209–217. https://doi.org/10.1007/s10641-006-0013-5
Adams AJ, Shenker JM, Jud ZR et al (2019) Identifying pre-spawning aggregation sites for bonefish (Albula vulpes) in the Bahamas to inform habitat protection and species conservation. Environ Biol Fishes 102:159–173. https://doi.org/10.1007/s10641-018-0802-7
Aguilar-Perera A (2006) Disappearance of a Nassau grouper spawning aggregation off the southern Mexican Caribbean coast. Mar Ecol Prog Ser 327:289–296. https://doi.org/10.3354/meps327289
Archer SK, Allgeier JE, Semmens BX et al (2015) Hot moments in spawning aggregations: implications for ecosystem-scale nutrient cycling. Coral Reefs 34:19–23. https://doi.org/10.1007/s00338-014-1208-4
Ascough JC, Maier HR, Ravalico JK, Strudley MW (2008) Future research challenges for incorporation of uncertainty in environmental and ecological decision-making. Ecol Modell 219:383–399. https://doi.org/10.1016/j.ecolmodel.2008.07.015
Binder TR, Farha SA, Thompson HT et al (2018) Fine-scale acoustic telemetry reveals unexpected lake trout, Salvelinus namaycush, spawning habitats in northern Lake Huron, North America. Ecol Freshw Fish 27:594–605. https://doi.org/10.1111/eff.12373
Breiman L (2001) Random forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Breiman L, Friedman J, Stone CJ, Olshen RA (1984) Classification algorithms and regression trees. Classification and regression tree. Wadsworth International Group, Belmont, California, pp 246–280
Brook RK, McLachlan SM (2008) Trends and prospects for local knowledge in ecological and conservation research and monitoring. Biodivers Conserv 17:3501–3512. https://doi.org/10.1007/s10531-008-9445-x
Brown DD, Kays R, Wikelski M et al (2013) Observing the unwatchable through acceleration logging of animal behavior. Anim Biotelemetry 1:20. https://doi.org/10.1186/2050-3385-1-20
Brownscombe JW, Adams AJ, Young N et al (2019a) Bridging the knowledge-action gap: a case of research rapidly impacting recreational fisheries policy. Mar Policy 104:210–215
Brownscombe JW, Danylchuk AJ, Adams AJ et al (2019b) Bonefish in South Florida: status, threats and research needs. Fish Res 102:329–348
Brownscombe JW, Griffin LP, Morley D et al (2020) Seasonal occupancy and connectivity amongst nearshore flats and reef habitats by permit (Trachinotus falcatus): considerations for fisheries management. J Fish Biol 96:469–479
Bryan DR, Luo J, Ault JS et al (2015) Transport and connectivity modeling of larval permit from an observed spawning aggregation in the Dry Tortugas, Florida. Environ Biol Fishes 98:2263–2276. https://doi.org/10.1007/s10641-015-0445-x
Cagnacci F, Boitani L, Powell RA, Boyce MS (2010) Animal ecology meets GPS-based radiotelemetry: a perfect storm of opportunities and challenges. Philos Trans R Soc B Biol Sci 375:2157–2162. https://doi.org/10.1098/rstb.2010.0107
Chon TS, Park YS, Park JH (2000) Determining temporal pattern of community dynamics by using unsupervised learning algorithms. Ecol Modell 132:151–166. https://doi.org/10.1016/S0304-3800(00)00312-4
Christin S, Hervet É, Lecomte N (2019) Applications for deep learning in ecology. Methods Ecol Evol 10:1632–1644. https://doi.org/10.1111/2041-210X.13256
Cooke SJ, Suski CD, Arlinghaus R, Danylchuk AJ (2013) Voluntary institutions and behaviours as alternatives to formal regulations in recreational fisheries management. Fish Fish 14:439–457. https://doi.org/10.1111/j.1467-2979.2012.00477.x
Crabtree RE, Hood PB, Snodgrass D (2002) Age, growth, and reproduction of permit (Trachinotus falcatus) in Florida waters. Fish Bull 100:26–34
Crossin GT, Heupel MR, Holbrook CM et al (2017) Acoustic telemetry and fisheries management. Ecol Appl 27:1031–1049. https://doi.org/10.1002/eap.1533
Cutler DR, Edwards TC, Beard KH et al (2007) Random forests for classification in ecology. Ecology 88:2783–2792. https://doi.org/10.1890/07-0539.1
Danylchuk AJ, Cooke SJ, Goldberg TL et al (2011) Aggregations and offshore movements as indicators of spawning activity of bonefish (Albula vulpes) in The Bahamas. Mar Biol 158:1981–1999. https://doi.org/10.1007/s00227-011-1707-6
De’Ath G, Fabricius KE (2000) Classification and regression trees: a powerful yet simple technique for ecological data analysis. Ecology 81:3178–3192. https://doi.org/10.1890/0012-9658(2000)081[3178:CARTAP]2.0.CO;2
De Mitcheson YS, Colin PL (2012) Reef fish spawning aggregations: biology, research and management. Springer, New York
Durden JM, Luo JY, Alexander H et al (2017) Integrating “big data” into aquatic ecology: challenges and opportunities. Limnol Oceanogr Bull 26:101–108. https://doi.org/10.1002/lob.10213
Elith J, Leathwick JR, Hastie T (2008) A working guide to boosted regression trees. J Anim Ecol 77:802–813. https://doi.org/10.1111/j.1365-2656.2008.01390.x
Equihua M (1990) Fuzzy clustering of ecological data. J Ecol. https://doi.org/10.2307/2261127
Erisman BE, Allen LG, Claisse JT et al (2011) The illusion of plenty: hyperstability masks collapses in two recreational fisheries that target fish spawning aggregations. Can J Fish Aquat Sci 68:1705–1716. https://doi.org/10.1139/f2011-090
Erisman B, Heyman W, Kobara S et al (2017) Fish spawning aggregations: where well-placed management actions can yield big benefits for fisheries and conservation. Fish Fish 18:128–144. https://doi.org/10.1111/faf.12132
Ferraro MB, Giordani P (2015) A toolbox for fuzzy clustering using the R programming language. Fuzzy Sets Syst 279:1–16. https://doi.org/10.1016/j.fss.2015.05.001
Fulton EA, Blanchard JL, Melbourne-Thomas J et al (2019) Where the ecological gaps remain, a modelers’ perspective. Front Ecol Evol 7:424. https://doi.org/10.3389/fevo.2019.00424
Graham RT, Castellanos DW (2005) Courtship and spawning behaviors of carangid species in Belize. Fish Bull 103:426–432
Graham RT, Carcamo R, Rhodes KL et al (2008) Historical and contemporary evidence of a mutton snapper (Lutjanus analis Cuvier, 1828) spawning aggregation fishery in decline. Coral Reefs 27:311–319. https://doi.org/10.1007/s00338-007-0329-4
Green J, Willis K, Hughes E et al (2007) Generating best evidence from qualitative research: the role of data analysis. Aust N Z J Public Health 31:545–550. https://doi.org/10.1111/j.1753-6405.2007.00141.x
Guilford T, Meade J, Willis J et al (2009) Migration and stopover in a small pelagic seabird, the Manx shearwater Puffinus puffinus: insights from machine learning. Proc R Soc B Biol Sci 276:1215–1223. https://doi.org/10.1098/rspb.2008.1577
Halsey LG (2019) The reign of the p -value is over: what alternative analyses could we employ to fill the power vacuum? Biol Lett 15:20190174. https://doi.org/10.1098/rsbl.2019.0174
Hamilton R, De Mitcheson YS, Aguilar-Perera A (2012) The role of local ecological knowledge in the conservation and management of reef fish spawning aggregations. In: Sadovy de Mitcheson Y, Colin P (eds) Reef fish spawning aggregations: biology, research and management. Springer, Dordrecht, pp 331–369
Hanski I, Gaggiotti O (2004) Ecology, genetics and evolution of metapopulations. Academic Press, Cambridge
Holder PE, Griffin LP, Adams AJ et al (2020) Stress, predators, and survival: exploring permit (Trachinotus falcatus) catch-and-release fishing mortality in the Florida Keys. J Exp Mar Bio Ecol 524:151289
Hoppner F, Klawonn F, Kruse R, Runkler T (2000) Fuzzy cluster analysis: methods for classification, data analysis and image recognition. Wiley, West Sussex
Hothorn T, Zeileis A (2015) Partykit: A modular toolkit for recursive partytioning in R. J Mach Learn Res 16(1):3905–3909
Hothorn T, Hornik K, Zeileis A (2006) Unbiased recursive partitioning: a conditional inference framework. J Comput Graph Stat 15:651–674. https://doi.org/10.1198/106186006X133933
Hussey NE, Kessel ST, Aarestrup K et al (2015) Aquatic animal telemetry: a panoramic window into the underwater world. Science 348:1255642. https://doi.org/10.1126/science.1255642
Johnson DH (1981) The use and misuse of statistics in wildlife habitat studies. Use Multivar Stat Stud Wildl Habitat Gen Tech Rep Stn. https://doi.org/10.5962/bhl.title.99662
Kobara S, Heyman WD (2008) Geomorphometric patterns of Nassau grouper (Epinephelus striatus) spawning aggregation sites in the Cayman Islands. Mar Geod 31:231–245. https://doi.org/10.1080/01490410802466397
Kobara S, Heyman WD (2010) Sea bottom geomorphology of multi-species spawning aggregation sites in Belize. Mar Ecol Prog Ser 405:243–254. https://doi.org/10.3354/meps08512
Kuhn M, Wing J, Weston S et al (2019) caret: Classification and regression training. R package version 6.0-84. https://CRAN.Rproject.org/package=caret
Leis JM, Carson-Ewart BM, Hay AC, Cato DH (2003) Coral-reef sounds enable nocturnal navigation by some reef-fish larvae in some places and at some times. J Fish Biol 63:724–737. https://doi.org/10.1046/j.1095-8649.2003.00182.x
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2:18–22. https://doi.org/10.1177/154405910408300516
Lowerre-Barbieri SK, Walters S, Bickford J et al (2013) Site fidelity and reproductive timing at a spotted seatrout spawning aggregation site: individual versus population scale behavior. Mar Ecol Prog Ser 481:181–197. https://doi.org/10.3354/meps10224
Lowerre-Barbieri SK, Walters Burnsed SL, Bickford JW (2016) Assessing reproductive behavior important to fisheries management: a case study with red drum, Sciaenops ocellatus. Ecol Appl 26:979–995. https://doi.org/10.1890/15-0497
Lowerre-Barbieri S, DeCelles G, Pepin P et al (2017) Reproductive resilience: a paradigm shift in understanding spawner–recruit systems in exploited marine fish. Fish Fish 18:285–312. https://doi.org/10.1111/faf.12180
Ludwig D, Hilborn R, Walters C (1993) Uncertainty, resource exploitation, and conservation: lessons from history. Science. https://doi.org/10.1126/science.260.5104.17
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197. https://doi.org/10.1016/S0169-5347(03)00008-9
Menzies CR (2006) Traditional ecological knowledge and natural resource management. U of Nebraska Press, Lincoln
Michener R, Lajtha K (2008) Stable isotopes in ecology and environmental science, 2nd edn. Wiley, Hoboken
Michener WK, Jones MB (2012) Ecoinformatics: supporting ecology as a data-intensive science. Trends Ecol Evol 27:85–93. https://doi.org/10.1016/j.tree.2011.11.016
Molnar C (2019) Interpretable machine learning. a guide for making black box models explainable. Lean Publishing
Mourier J, Maynard J, Parravicini V et al (2016) Extreme inverted trophic pyramid of reef sharks supported by spawning groupers. Curr Biol 26:2011–2016. https://doi.org/10.1016/j.cub.2016.05.058
Nguyen VM, Young N, Cooke SJ (2017) A roadmap for knowledge exchange and mobilization research in conservation and natural resource management. Conserv Biol 31:789–798. https://doi.org/10.1111/cobi.12857
Nguyen VM, Young N, Cooke S (2018) Applying a knowledge-action framework for navigating barriers to incorporating telemetry science into fisheries management and conservation: a qualitative study. Can J Fish Aquat Sci 75:1733–1743
Nowlin WH, Vanni MJ, Yang LH (2008) Comparing resource pulses in aquatic and terrestrial ecosystems. Ecology 89(3):647–659
Olden JD, Lawler JJ, Poff NL (2008) Machine learning methods without tears: a primer for ecologists. Q Rev Biol 83:171–193. https://doi.org/10.1086/587826
Oppel S, Strobl C, Huettmann F (2009) Alternative methods to quantify variable importance in ecology. Ludwig-Maximilians-Universität, München, pp 1–7
Paris CB, Atema J, Irisson JO et al (2013) Reef odor: a wake up call for navigation in reef fish larvae. PLoS ONE 8:e72808. https://doi.org/10.1371/journal.pone.0072808
Peters DPC, Havstad KM, Cushing J et al (2014) Harnessing the power of big data: Infusing the scientific method with machine learning to transform ecology. Ecosphere 5:1–15. https://doi.org/10.1890/ES13-00359.1
Piironen J, Paasiniemi M, Vehtari A (2020) Projective inference in high-dimensional problems: prediction and feature selection. Electron J Stat. https://doi.org/10.1214/20-ejs1711
Pitcher TJ (2001) Fish schooling. In: Steele JH, Thorpe SA, Turekian KK (eds) Encyclopedia of ocean sciences: marine biology, pp 337–349
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.Rproject.org/
Ribeiro MT, Singh S, Guestrin C (2016) “Why should I trust you?” Explaining the predictions of any classifier. Proc ACM SIGKDD Int Conf Knowl Discov Data Min. https://doi.org/10.1145/2939672.2939778
RStudio Team (2016) RStudio: Integrated Development for R. RStudio Inc., Boston, MA. http://www.rstudio.com/
Sadovy Y, Domeier M (2005) Are aggregation-fisheries sustainable? Reef fish fisheries as a case study. Coral Reefs 24:254–262. https://doi.org/10.1007/s00338-005-0474-6
Sala E, Ballesteros E, Starr RM (2001) Rapid decline of Nassau grouper spawning aggregations in belize: fishery management and conservation needs. Fisheries 26:23–30. https://doi.org/10.1577/1548-8446(2001)026<0023:rdongs>2.0.co;2
Salski A (2007) Fuzzy clustering of fuzzy ecological data. Ecol Inform 2:262–269. https://doi.org/10.1016/j.ecoinf.2007.07.002
Sancho G, Petersen CW, Lobel PS (2000) Predator–prey relations at a spawning aggregation site of coral reef fishes. Mar Ecol Prog Ser. https://doi.org/10.3354/meps203275
Santos RO, Rehage JS, Kroloff EKN et al (2018) Combining data sources to elucidate spatial patterns in recreational catch and effort: fisheries-dependent data and local ecological knowledge applied to the South Florida bonefish fishery. Environ Biol Fishes 201:299–317
Silvano RAM, MacCord PFL, Lima RV, Begossi A (2006) When does this fish spawn? Fishermen’s local knowledge of migration and reproduction of Brazilian coastal fishes. Environ Biol Fishes 76:371–386. https://doi.org/10.1007/s10641-006-9043-2
Simpfendorfer CA, Huveneers C, Steckenreuter A et al (2015) Ghosts in the data: false detections in VEMCO pulse position modulation acoustic telemetry monitoring equipment. Anim Biotelemetry 3:55. https://doi.org/10.1186/s40317-015-0094-z
Soria M, Dagorn L, Potin G, Fréon P (2009) First field-based experiment supporting the meeting point hypothesis for schooling in pelagic fish. Anim Behav 78:1441–1446. https://doi.org/10.1016/j.anbehav.2009.09.025
Strobl C, Boulesteix AL, Zeileis A, Hothorn T (2007) Bias in random forest variable importance measures: illustrations, sources and a solution. BMC Bioinform 8:25. https://doi.org/10.1186/1471-2105-8-25
Strobl C, Boulesteix A-L, Kneib T et al (2008) Conditional variable importance for random forests. BMC Bioinform 9:307. https://doi.org/10.1186/1471-2105-9-307
Strobl C, Hothorn T, Zeileis A (2009) Party on! A new, conditional variable importance measure available in the party package. Technical Report Number 050, Department of Statistics, University of Munich
Waterhouse L, Heppell SA, Pattengill-Semmens CV et al (2020) Recovery of critically endangered Nassau grouper (Epinephelus striatus) in the Cayman Islands following targeted conservation actions. Proc Natl Acad Sci USA 117:1587–1595. https://doi.org/10.1073/pnas.1917132117
West JB, Bowen GJ, Cerling TE, Ehleringer JR (2006) Stable isotopes as one of nature’s ecological recorders. Trends Ecol Evol 21:408–414. https://doi.org/10.1016/j.tree.2006.04.002
Young N, Gingras I, Nguyen VM et al (2013) Mobilizing new science into management practice: the challenge of biotelemetry for fisheries management, a case study of Canada’s Fraser River. J Int Wildl Law Policy 16:331–351. https://doi.org/10.1080/13880292.2013.805074
Zeller DC (1998) Spawning aggregations: Patterns of movement of the coral trout Plectropomus leopardus (Serranidae) as determined by ultrasonic telemetry. Mar Ecol Prog Ser 162:253–263. https://doi.org/10.3354/meps162253
Zeng X, Adams A, Roffer M, He R (2018) Potential connectivity among spatially distinct management zones for Bonefish (Albula vulpes) via larval dispersal. Environ Biol Fishes 102:233–252. https://doi.org/10.1007/s10641-018-0826-z
Zuur AF, Ieno EN, Elphick CS (2010) A protocol for data exploration to avoid common statistical problems. Methods Ecol Evol 1:3–14. https://doi.org/10.1111/j.2041-210X.2009.00001.x
Acknowledgements
This project was funded by Bonefish and Tarpon Trust with support from Costa Del Mar, The March Merkin Fishing Tournament, Hell’s Bay Boatworks, and private donations. Additional support was provided by a NASEM Gulf Research Program through a hurricane recovery grant, and the acoustic receiver array was partially supported by a loan from the Ocean Tracking Network. We thank the fishing guides and anglers who assisted with the telemetry array design and Permit tagging for this project including Captains Travis and Bear Holeman, Will Benson, Rob Kramarz, Zack Stells, Brandon, and Jared Cyr, Chris Trosset, Nathaniel Linville, Ian Slater, Augustine Moss, Sandy Horn, Ted Margo, Richard Berlin, and Jeff Rella. We also thank the researchers that shared permit acoustic telemetry detection data with us through integrated Tracking of Animals in the Gulf (iTAG) and Florida Acoustic Telemetry Network (FACT), in particular Harold "Wes" Pratt of Mote Marine Laboratory, funded by The Shark Foundation/Hai Stiftung, and Mike McAllister. Brownscombe is supported by a Banting Postdoctoral Fellowship, Dalhousie University, Carleton University, and Bonefish and Tarpon Trust. This research was conducted with permission of the Florida Keys National Marine Sanctuary under permit # FKNMS-2013-040-A2, and the Florida Fish and Wildlife Conservation Commission under permit # SAL-16-1205.
Author information
Authors and Affiliations
Contributions
JWB designed and conducted data collection, analysis, and manuscript writing. LPG, DM, AA, JH, SKL-B, AJA, AJD, SJC contributed to data collection and manuscript preparation.
Corresponding author
Additional information
Communicated by Yannis Papastamatiou.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary file2 (MOV 30821 kb)
Rights and permissions
About this article
Cite this article
Brownscombe, J.W., Griffin, L.P., Morley, D. et al. Application of machine learning algorithms to identify cryptic reproductive habitats using diverse information sources. Oecologia 194, 283–298 (2020). https://doi.org/10.1007/s00442-020-04753-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00442-020-04753-2