Abstract
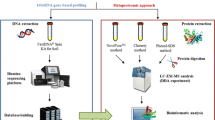
Mass spectrometry-based proteomics was applied to analyze proteins isolated from dissolved organic matter (DOM). The focal question was to identify the type and biological origin of proteins in DOM, and to describe diversity of protein origin at the level of higher taxonomic units, as well as to detect extracellular enzymes possibly important in the carbon cycle. Identified proteins were classified according to their phylogenetic origin and metabolic function using the National Center for Biotechnology Information (NCBI) protein and taxonomy database. Seventy-eight percent of the proteins in DOM from the lake but less than 50% in forest soil DOM originated from bacteria. In a deciduous forest, the number of identified proteins decreased from 75 to 28 with increasing soil depth and decreasing total soil organic carbon content. The number of identified proteins and taxonomic groups was 50% higher in winter than in summer. In spruce forest, number of proteins and taxonomic groups decreased by 50% on a plot where trees had been girdled a year before and carbohydrate transport to roots was terminated. After girdling, proteins from four taxonomic groups remained as compared to nine taxonomic groups in healthy forest. Enzymes involved in degradation of organic matter were not identified in free soil DOM. However, cellulases and laccases were found among proteins extracted from soil particles, indicating that degradation of soil organic matter takes place in biofilms on particle surfaces. These results demonstrate a novel application of proteomics to obtain a “proteomic fingerprint” of presence and activity of organisms in an ecosystem.






Similar content being viewed by others
References
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422:198–207
Almendros G, Frund R, Gonzalez-Vila FJ, Haider KM, Knicker H, Ludemann HD (1991) Analysis of 13C and 15N CPMAS NMR-spectra of soil organic matter and composts. FEBS Lett 282:119–121
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Arnold RJ, Reilly JP (1998) Fingerprint matching of E. coli strains with matrix-assisted laser desorption/ionization time-of-flight mass spectrometry of whole cells using a modified correlation approach. Rapid Commun Mass Spectrom 12:630–636
Báldi A (2003) Using higher taxa as surrogates of species richness: a study based on 3700 Coleoptera, Diptera and Acri in Central-Hungarian reserves. Basic Appl Ecol 4:589–593
Battin TJ, Kaplan LA, Newbold JD, Hansen CME (2003) Contributions of microbial biofilms to ecosystem processes in stream mesocosms. Nature 426:439–440
Christensen BT (1992) Physical fractionation of soil and organic matter in primary particle size and density separates. Adv Soil Sci 20:1–90
Dickinson DN, La Duc MT, Haskins WE, Gornushkin I, Winefordner JD, Powell DH, Venkateswaran K (2004a) Species differentiation of a diverse suite of bacillus spores by mass spectrometry-based protein profiling. Appl Environ Microbiol 70:472–482
Dickinson DN, La Duc MT, Satomi M, Wienfordner JD, Powell DH, Venkateswaran K (2004b) MALDI-TOFMS compared with other polyphasic taxonomy approaches for the identification and classification of Bacillus pumilus spores. J Microbiol Methods 58:1–12
Gleixner G, Czimczik C, Kramer C, Lühker B, Schmidt MWI (2001) Plant compounds and their turnover and stability as soil organic matter. In: Schulze ED et al (eds) Global biogeochemical cycles in the climate system. Academic, San Diego, pp 201–216
Graham IA, Baker CJ, Leaver CJ (1994) Carbon catabolite repression regulates glyoxylate cycle gene expression in cucumber. Plant Cell 6:761–772
Guggenberger G, Kaiser K (2003) Dissolved organic matter in soil: challenging the paradigm of sorptive preservation. Geoderma 113:293–310
Habermann B, Oegerma J, Sunyaev S, Shevchenko A (2004) The power and the limitations of cross-species protein identification by mass spectrometry-driven sequence similarity searches. Mol Cell Proteomics 3:238–249
Hahn V (2003) Determination and modelling of 14C ages of soil respiration. Max-Planck Institute of Biogeochemistry, Jena
Harris SA, Robinson JP, Juniper BE (2002) Genetic clues to the origin of the apple. Trends Gene 18:426–430
Heywood VH, Watson RT (1995) Global biodiversity assessment. Cambridge University Press, London
Ishihama Y, Rappsilber J, Andersen JS, Mann M (2002) Microcolumns with self-assembled particle frits for proteomics. J Chromatogr A 979:233–239
Kaiser K, Guggenberger G, Haumaer L, Zech W (2001) Seasonal variations in the chemical composition of dissolved organic matter in organic forest floor layer leachates of old-growth Scots pine (Pinus sylvestris L.) and European beech (Fags sylvatica L.) stands in northeastern Bavaria, Germany. Biogeochemistry 55:103–143
Kjöller A, Miller M, Struwe S, Wolters V, Pflug A (2000) Diversity and role of microorganisms. In: Schulze ED (ed) Carbon and nitrogen cycling in European forest ecosystems, vol 142. Springer, Berlin Heidelberg New York, pp 382–404
Knohl A, Schulze ED, Kolle O, Buchmann N (2003) Large carbon uptake by an unmanaged 250-year-old deciduous forest in central Germany. Agr Forest Meteorol 118:151–167
Kracht O, Gleixner G (2000) Isotope analysis of pyrolysis products from Sphagnum peat and dissolved organic matter from bog water. Org Geochem 31:645–654
Lipson DA, Schadt CW, Schmidt SK (2002) Changes in soil microbial community structure and function in an alpine dry meadow following spring snow melt. Microb Ecol 43:307–314
Loreau M, Hector A, Inchausti P (2002) Biodiversity and ecosystem functioning. Oxford University Press, Oxford
Luis P, Walther G, Kellner H, Martin F, Buscot F (2004) Diversity of laccase genes from basidiomycetes in a forest soil. Soil Biol Biochem (in press)
Michalzik B, Matzner E (1999) Dynamics of dissolved organic nitrogen and carbon in a Central European Norway spruce ecosystem. Eur J Soil Sci 50:579–590
Perkins DN, Pappin DJC, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20:3551–3567
Purvis A, Hector A (2000) Getting the measure of biodiversity. Nature 405:212–219
Rappsilber J, Mann M (2002) What does it mean to identify a protein in proteomics. Trends Biochem Sci 27:74–78
Rappsilber J, Ryder U, Lamon AI, Mann M (2002) Large-scale proteomic analysis of the human spliceosome. Genome Res 12:1231–1245
Rappsilber J, Ishihama Y, Mann M (2003) Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal Chem 75:663–670
Schulten HR, Gleixner G (1999) Analytical pyrolysis of humic substances and dissolved organic matter in aquatic systems: structure and origin. Water Res 33:2489–2498
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem 68:850–858
Stemmer M, Gerzabek MH, Kandeler E (1998) Organic matter and enzyme activity in particle-size fractions of soils obtained after low-energy sonication. Soil Biol Biochem 30:9–17
Tyers M, Mann M (2003) From genomics to proteomics. Nature 422:193–197
Tyson GW et al. (2004) Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428:37–43
Venter JC et al. (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science 304:66–74
Zelles L (1997) Phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere 35:275–294
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schulze, W.X., Gleixner, G., Kaiser, K. et al. A proteomic fingerprint of dissolved organic carbon and of soil particles. Oecologia 142, 335–343 (2005). https://doi.org/10.1007/s00442-004-1698-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00442-004-1698-9