Abstract
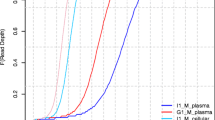
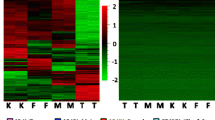
Chromosome stability is a key point in genome evolution, particularly that of the Y chromosome. The Y chromosome loss in blood and tumor cells is well established. Through processes that are common to other chromosomes too, the Y chromosome undergoes degradation and fragmentation in the blood stream before elimination. This process gives rise to circulating DNA (cirDNA) fragments, whose examination may provide potential insight into the role of DNA fragmentation in blood for the Y chromosome elimination. In this study, we employed shallow whole genome sequencing (sWGS) to comprehensively assess the total cirDNA and the individual chromosome fragment size profiles in the plasma of healthy male individuals. Here, we show that (i) the fragment size profiles of total circulating DNA (cirDNA) and DNA fragments originating from autosomes and the X chromosome in blood plasma are homogeneous, and have a remarkably low variability (mean CV = 7%) among healthy individuals, (ii) the Y chromosome has a distinct fragment size profile with the accumulation of the fragment < 145 bp and depletion of the dinucleosome-associated fragments (290–390 bp), and its fragment fraction in blood decreases with age. These results indicate a higher fragmentation of the Y chromosome compared to other chromosomes and this in turn might be due to its increased susceptibility to degradation. Our findings pave the way for an elucidation of the impact of chromosomal origin on DNA degradation and the Y chromosome biology.


Similar content being viewed by others
Data availability
Sequence data has been deposited at the European Genome-phenome Archive (EGA), which is hosted by the EBI and the CRG, under accession number EGAS00001006830.
References
An Y, Zhao X, Zhang Z et al (2023) DNA methylation analysis explores the molecular basis of plasma cell-free DNA fragmentation. Nat Commun 14:287. https://doi.org/10.1038/s41467-023-35959-6
Bianchi NO (2009) Y chromosome structural and functional changes in human malignant diseases. Mutation Res/rev Mutation Res 682:21–27. https://doi.org/10.1016/j.mrrev.2009.02.001
Chan KCA, Zhang J, Hui ABY et al (2004) Size distributions of maternal and fetal DNA in maternal plasma. Clin Chem 50:88–92. https://doi.org/10.1373/clinchem.2003.024893
Chandrananda D, Thorne NP, Bahlo M (2015) High-resolution characterization of sequence signatures due to non-random cleavage of cell-free DNA. BMC Med Genomics 8:29. https://doi.org/10.1186/s12920-015-0107-z
Duijf PHG, Schultz N, Benezra R (2013) Cancer cells preferentially lose small chromosomes. Int J Cancer 132:2316–2326. https://doi.org/10.1002/ijc.27924
Forsberg LA (2017) Loss of chromosome Y (LOY) in blood cells is associated with increased risk for disease and mortality in aging men. Hum Genet 136:657–663. https://doi.org/10.1007/s00439-017-1799-2
Guizard S, Piégu B, Bigot Y (2016) DensityMap: a genome viewer for illustrating the densities of features. BMC Bioinform 17:204. https://doi.org/10.1186/s12859-016-1055-0
Han DSC, Ni M, Chan RWY et al (2020) The biology of cell-free DNA fragmentation and the roles of DNASE1, DNASE1L3, and DFFB. Am J Hum Genet 106:202–214. https://doi.org/10.1016/j.ajhg.2020.01.008
Hauer MH, Seeber A, Singh V et al (2017) Histone degradation in response to DNA damage enhances chromatin dynamics and recombination rates. Nat Struct Mol Biol 24:99–107. https://doi.org/10.1038/nsmb.3347
Hunter S, Gramlich T, Abbott K, Varma V (1993) Y chromosome loss in esophageal carcinoma: an in situ hybridization study. Genes Chromosom Cancer 8:172–177. https://doi.org/10.1002/gcc.2870080306
Jacobs PA, Brunton M, Court Brown WM et al (1963) Change of human chromosome count distribution with age: evidence for a sex differences. Nature 197:1080–1081. https://doi.org/10.1038/1971080a0
Jahr S, Hentze H, Englisch S et al (2001) DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res 61:1659–1665
Koyama R, Arai T, Kijima M et al (2016) DNase γ, DNase I and caspase-activated DNase cooperate to degrade dead cells. Genes Cells 21:1150–1163. https://doi.org/10.1111/gtc.12433
Kumar D, Upadhya D, Uppangala S et al (2013) Nuclear DNA fragmentation negatively affects zona binding competence of Y bearing mouse spermatozoa. J Assist Reprod Genet 30:1611–1615. https://doi.org/10.1007/s10815-013-0123-x
Li G, Wang C, Guan X, et al (2022) Age-related DNA methylation on Y chromosome and their associations with total mortality among Chinese males. Aging Cell 21:e13563. https://doi.org/10.1111/acel.13563
Lo YM, Corbetta N, Chamberlain PF et al (1997) Presence of fetal DNA in maternal plasma and serum. Lancet 350:485–487. https://doi.org/10.1016/S0140-6736(97)02174-0
Lui YYN, Chik K-W, Chiu RWK et al (2002) Predominant hematopoietic origin of cell-free DNA in plasma and serum after sex-mismatched bone marrow transplantation. Clin Chem 48:421–427
Meddeb R, Pisareva E, Thierry AR (2019) Guidelines for the preanalytical conditions for analyzing circulating cell-free DNA. Clin Chem 65:623–633. https://doi.org/10.1373/clinchem.2018.298323
Moss J, Magenheim J, Neiman D et al (2018) Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease. Nat Commun. https://doi.org/10.1038/s41467-018-07466-6
Mouliere F, Robert B, Arnau Peyrotte E et al (2011) High fragmentation characterizes tumour-derived circulating DNA. PLoS ONE 6:e23418. https://doi.org/10.1371/journal.pone.0023418
Park S-J, Jeong S-Y, Kim HJ (2006) Y chromosome loss and other genomic alterations in hepatocellular carcinoma cell lines analyzed by CGH and CGH array. Cancer Genet Cytogenet 166:56–64. https://doi.org/10.1016/j.cancergencyto.2005.08.022
Pisareva E, Mihalovičová L, Pastor B et al (2022) Neutrophil extracellular traps have auto-catabolic activity and produce mononucleosome-associated circulating DNA. Genome Medicine 14:135. https://doi.org/10.1186/s13073-022-01125-8
Reinhold K, Engqvist L (2013) The variability is in the sex chromosomes. Evolution 67:3662–3668. https://doi.org/10.1111/evo.12224
Sanchez C, Snyder MW, Tanos R et al (2018) New insights into structural features and optimal detection of circulating tumor DNA determined by single-strand DNA analysis. NPJ Genom Med 3:31. https://doi.org/10.1038/s41525-018-0069-0
Sanchez C, Roch B, Mazard T et al (2021) Circulating nuclear DNA structural features, origins, and complete size profile revealed by fragmentomics. JCI Insight 6:144561. https://doi.org/10.1172/jci.insight.144561
Skaletsky H, Kuroda-Kawaguchi T, Minx PJ et al (2003) The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423:825–837. https://doi.org/10.1038/nature01722
Snyder MW, Kircher M, Hill AJ et al (2016) Cell-free DNA comprises an in vivo nucleosome footprint that informs its tissues-of-origin. Cell 164:57–68. https://doi.org/10.1016/j.cell.2015.11.050
Sun K, Jiang P, Chan KCA et al (2015) Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments. Proc Natl Acad Sci USA 112:E5503-5512. https://doi.org/10.1073/pnas.1508736112
Thierry AR (2023) Circulating DNA fragmentomics and cancer screening. Cell Genomics 3:100242. https://doi.org/10.1016/j.xgen.2022.100242
Thierry AR, El Messaoudi S, Gahan PB et al (2016) Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev 35:347–376. https://doi.org/10.1007/s10555-016-9629-x
Xue Y, Zhao G, Qiao L et al (2020) Sequencing shorter cfDNA fragments decreases the false negative rate of non-invasive prenatal testing. Front Genet 11:280. https://doi.org/10.3389/fgene.2020.00280
Yates AD, Achuthan P, Akanni W et al (2020) Ensembl 2020. Nucleic Acids Res 48:D682–D688. https://doi.org/10.1093/nar/gkz966
You Y-A, Kwon W-S, Saidur Rahman M et al (2017) Sex chromosome-dependent differential viability of human spermatozoa during prolonged incubation. Hum Reprod 32:1183–1191. https://doi.org/10.1093/humrep/dex080
Zheng YWL, Chan KCA, Sun H et al (2012) Nonhematopoietically derived DNA is shorter than hematopoietically derived DNA in plasma: a transplantation model. Clin Chem 58:549–558. https://doi.org/10.1373/clinchem.2011.169318
Acknowledgements
The authors thank the excellent technical assistance of F. Frayssinoux (IRCM, Institut de Recherche en Cancérologie de Montpellier, INSERM U1194, Université de Montpellier, Institut régional du Cancer de Montpellier, Montpellier, F-34134298, France) and Cormac Mc Carthy (Mc Carthy Consultant, Montpellier) for English editing (financial compensation). We thank the healthy donors who participated in this study.
Funding
This work was partially supported by SIRIC Montpellier Cancer Grant INCa_Inserm_DGOS_12553.
Author information
Authors and Affiliations
Contributions
Conceptualization: AT, EP. Methodology: AT, EP. Investigation: EP, CS. Visualization: AT, EP, CS. Supervision: AT, EP. Writing—original draft: AT, EP. Writing—review and editing: AT, EP.
Corresponding authors
Ethics declarations
Conflict of interest
Authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thierry, A.R., Sanchez, C., Colinge, J. et al. Circulating DNA reveals a specific and higher fragmentation of the Y chromosome. Hum. Genet. 142, 1603–1609 (2023). https://doi.org/10.1007/s00439-023-02600-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-023-02600-x