Abstract
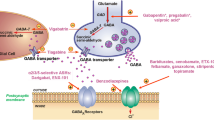
GRIA3 at Xq25 encodes glutamate ionotropic receptor AMPA type 3 (GluA3), a subunit of postsynaptic glutamate-gated ion channels mediating neurotransmission. Hemizygous loss-of-function (LOF) variants in GRIA3 cause a neurodevelopmental disorder (NDD) in male individuals. Here, we report a gain-of-function (GOF) variant at GRIA3 in a male patient. We identified a hemizygous de novo missense variant in GRIA3 in a boy with an NDD: c.1844C > T (p.Ala615Val) using whole-exome sequencing. His neurological signs, such as hypertonia and hyperreflexia, were opposite to those in previous cases having LOF GRIA3 variants. His seizures and hypertonia were ameliorated by carbamazepine, inhibiting glutamate release from presynapses. Patch-clamp recordings showed that the human GluA3 mutant (p.Ala615Val) had slower desensitization and deactivation kinetics. A fly line expressing a human GluA3 mutant possessing our variant and the Lurcher variant, which makes ion channels leaky, showed developmental defects, while one expressing a mutant possessing either of them did not. Collectively, these results suggest that p.Ala615Val has GOF effects. GRIA3 GOF variants may cause an NDD phenotype distinctive from that of LOF variants, and drugs suppressing glutamatergic neurotransmission may ameliorate this phenotype. This study should help in refining the clinical management of GRIA3-related NDDs.




Similar content being viewed by others
References
Abdelsayed M, Sokolov S (2013) Voltage-gated sodium channels: pharmaceutical targets via anticonvulsants to treat epileptic syndromes. Channels (austin) 7:146–152. https://doi.org/10.4161/chan.24380
Adzhubei IA et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249. https://doi.org/10.1038/nmeth0410-248
Ahrens-Nicklas RC et al (2017) Precision therapy for a new disorder of AMPA receptor recycling due to mutations in ATAD1. Neurol Genet 3:e130. https://doi.org/10.1212/nxg.0000000000000130
Allen NM et al (2016) Unexplained early onset epileptic encephalopathy: exome screening and phenotype expansion. Epilepsia 57:e12-17. https://doi.org/10.1111/epi.13250
Aspromonte MC et al (2020) Characterization of intellectual disability and autism comorbidity through gene panel sequencing. Hum Mutat 41:1183. https://doi.org/10.1002/humu.24012
Bischof J, Maeda RK, Hediger M, Karch F, Basler K (2007) An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc Natl Acad Sci USA 104:3312–3317. https://doi.org/10.1073/pnas.0611511104
Bliss TV, Collingridge GL (1993) A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361:31–39. https://doi.org/10.1038/361031a0
Bonnet C, Leheup B, Beri M, Philippe C, Gregoire MJ, Jonveaux P (2009) Aberrant GRIA3 transcripts with multi-exon duplications in a family with X-linked mental retardation. Am J Med Genet Part A 149a:1280–1289. https://doi.org/10.1002/ajmg.a.32858
Bredt DS, Nicoll RA (2003) AMPA receptor trafficking at excitatory synapses. Neuron 40:361–379. https://doi.org/10.1016/s0896-6273(03)00640-8
Chérot E et al (2018) Using medical exome sequencing to identify the causes of neurodevelopmental disorders: experience of 2 clinical units and 216 patients. Clin Genet 93:567–576. https://doi.org/10.1111/cge.13102
Chiyonobu T et al (2007) Partial tandem duplication of GRIA3 in a male with mental retardation. Am J Med Genet Part A 143a:1448–1455. https://doi.org/10.1002/ajmg.a.31798
Davies B et al (2017) A point mutation in the ion conduction pore of AMPA receptor GRIA3 causes dramatically perturbed sleep patterns as well as intellectual disability. Hum Mol Genet 26:3869–3882. https://doi.org/10.1093/hmg/ddx270
Enjoji M, Yanai N (1961) Analytic test for development in infancy and childhood. Pediatr Int 4:2–6. https://doi.org/10.1111/j.1442-200X.1961.tb01032.x
Fernández-Marmiesse A et al (2019) Rare variants in 48 genes account for 42% of cases of epilepsy with or without neurodevelopmental delay in 246 pediatric patients. Front Neurosci 13:1135. https://doi.org/10.3389/fnins.2019.01135
Garcia-Boronat M, Diez-Rivero CM, Reinherz EL, Reche PA (2008) PVS: a web server for protein sequence variability analysis tuned to facilitate conserved epitope discovery. Nucleic Acids Res 36:W35-41. https://doi.org/10.1093/nar/gkn211
Gauthier AC, Mattson RH (2015) Clobazam: a safe efficacious, and newly rediscovered therapeutic for epilepsy. CNS Neurosci Ther 21:543–548. https://doi.org/10.1111/cns.12399
Gecz J et al (1999) Characterization of the human glutamate receptor subunit 3 gene (GRIA3), a candidate for bipolar disorder and nonspecific X-linked mental retardation. Genomics 62:356–368. https://doi.org/10.1006/geno.1999.6032
Geisheker MR et al (2017) Hotspots of missense mutation identify neurodevelopmental disorder genes and functional domains. Nat Neurosci 20:1043–1051. https://doi.org/10.1038/nn.4589
Gregory JM, Livesey MR, McDade K, Selvaraj BT, Barton SK, Chandran S, Smith C (2020) Dysregulation of AMPA receptor subunit expression in sporadic ALS post-mortem brain. J Pathol 250:67–78. https://doi.org/10.1002/path.5351
Guerois R, Nielsen JE, Serrano L (2002) Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J Mol Biol 320:369–387. https://doi.org/10.1016/s0022-2836(02)00442-4
Hazelett DJ, Bourouis M, Walldorf U, Treisman JE (1998) decapentaplegic and wingless are regulated by eyes absent and eyegone and interact to direct the pattern of retinal differentiation in the eye disc. Development (cambridge, England) 125:3741–3751
Henley JM, Wilkinson KA (2016) Synaptic AMPA receptor composition in development, plasticity and disease. Nat Rev Neurosci 17:337–350. https://doi.org/10.1038/nrn.2016.37
Higasa K et al (2016) Human genetic variation database, a reference database of genetic variations in the Japanese population. J Hum Genet 61:547–553. https://doi.org/10.1038/jhg.2016.12
Karczewski KJ et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581:434–443. https://doi.org/10.1038/s41586-020-2308-7
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006. https://doi.org/10.1101/gr.229102
Kondo S, Ueda R (2013) Highly improved gene targeting by germline-specific Cas9 expression in Drosophila. Genetics 195:715–721. https://doi.org/10.1534/genetics.113.156737
Kumar P, Henikoff S, Ng PC (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4:1073–1081. https://doi.org/10.1038/nprot.2009.86
Lal D et al (2020) Gene family information facilitates variant interpretation and identification of disease-associated genes in neurodevelopmental disorders. Genome Med 12:28. https://doi.org/10.1186/s13073-020-00725-6
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, Maglott DR (2014) ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42:D980-985. https://doi.org/10.1093/nar/gkt1113
Maren S, Tocco G, Standley S, Baudry M, Thompson RF (1993) Postsynaptic factors in the expression of long-term potentiation (LTP): increased glutamate receptor binding following LTP induction in vivo. Proc Natl Acad Sci USA 90:9654–9658. https://doi.org/10.1073/pnas.90.20.9654
Markstein M, Pitsouli C, Villalta C, Celniker SE, Perrimon N (2008) Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nat Genet 40:476–483. https://doi.org/10.1038/ng.101
Nagasaki M et al (2015) Rare variant discovery by deep whole-genome sequencing of 1070 Japanese individuals. Nat Commun 6:8018. https://doi.org/10.1038/ncomms9018
Pérez-Palma E et al (2020) Identification of pathogenic variant enriched regions across genes and gene families. Genome Res 30:62–71. https://doi.org/10.1101/gr.252601.119
Philippe A et al (2013) Xq25 duplications encompassing GRIA3 and STAG2 genes in two families convey recognizable X-linked intellectual disability with distinctive facial appearance. Am J Med Genet Part A 161a:1370–1375. https://doi.org/10.1002/ajmg.a.35307
Philips AK et al (2014) X-exome sequencing in Finnish families with intellectual disability-four novel mutations and two novel syndromic phenotypes. Orphanet J Rare Dis 9:49. https://doi.org/10.1186/1750-1172-9-49
Platt SR (2007) The role of glutamate in central nervous system health and disease—a review. Vet J (london England: 1997) 173:278–286. https://doi.org/10.1016/j.tvjl.2005.11.007
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M (2019) CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res 47:D886-d894. https://doi.org/10.1093/nar/gky1016
Richards S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med off J Am Coll Med Genet 17:405–424. https://doi.org/10.1038/gim.2015.30
Sakamoto M et al (2021) Novel EXOSC9 variants cause pontocerebellar hypoplasia type 1D with spinal motor neuronopathy and cerebellar atrophy. J Hum Genet 66:401–407. https://doi.org/10.1038/s10038-020-00853-2
Schwarz JM, Rödelsperger C, Schuelke M, Seelow D (2010) MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 7:575–576. https://doi.org/10.1038/nmeth0810-575
Schwarz N et al (2016) Mutations in the sodium channel gene SCN2A cause neonatal epilepsy with late-onset episodic ataxia. J Neurol 263:334–343. https://doi.org/10.1007/s00415-015-7984-0
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L (2005) The FoldX web server: an online force field. Nucleic Acids Res 33:W382-388. https://doi.org/10.1093/nar/gki387
Sitges M, Guarneros A, Nekrassov V (2007) Effects of carbamazepine, phenytoin, valproic acid, oxcarbazepine, lamotrigine, topiramate and vinpocetine on the presynaptic Ca2+ channel-mediated release of [3H]glutamate: comparison with the Na+ channel-mediated release. Neuropharmacology 53:854–862. https://doi.org/10.1016/j.neuropharm.2007.08.016
Sitges M, Chiu LM, Reed RC (2016) Effects of levetiracetam, carbamazepine, phenytoin, valproate, lamotrigine, oxcarbazepine, topiramate, vinpocetine and sertraline on presynaptic hippocampal Na(+) and Ca(2+) channels permeability. Neurochem Res 41:758–769. https://doi.org/10.1007/s11064-015-1749-0
Stenson PD et al (2003) Human gene mutation database (HGMD): 2003 update. Hum Mutat 21:577–581. https://doi.org/10.1002/humu.10212
Sun JH et al (2021) X-linked neonatal-onset epileptic encephalopathy associated with a gain-of-function variant p.R660T in GRIA3. PLoS Genet 17:e1009608. https://doi.org/10.1371/journal.pgen.1009608
Twomey EC, Yelshanskaya MV, Grassucci RA, Frank J, Sobolevsky AI (2017a) Channel opening and gating mechanism in AMPA-subtype glutamate receptors. Nature 549:60–65. https://doi.org/10.1038/nature23479
Twomey EC, Yelshanskaya MV, Grassucci RA, Frank J, Sobolevsky AI (2017b) Structural bases of desensitization in AMPA receptor-auxiliary subunit complexes. Neuron 94:569-580.e565. https://doi.org/10.1016/j.neuron.2017.04.025
Wiel L, Baakman C, Gilissen D, Veltman JA, Vriend G, Gilissen C (2019) MetaDome: pathogenicity analysis of genetic variants through aggregation of homologous human protein domains. Hum Mutat 40:1030–1038. https://doi.org/10.1002/humu.23798
Wolff M et al (2017) Genetic and phenotypic heterogeneity suggest therapeutic implications in SCN2A-related disorders. Brain J Neurol 140:1316–1336. https://doi.org/10.1093/brain/awx054
Wu Y et al (2007) Mutations in ionotropic AMPA receptor 3 alter channel properties and are associated with moderate cognitive impairment in humans. Proc Natl Acad Sci USA 104:18163–18168. https://doi.org/10.1073/pnas.0708699104
Yang Y, Hou L, Li Y, Ni J, Liu L (2013) Neuronal necrosis and spreading death in a Drosophila genetic model. Cell Death Dis 4:e723. https://doi.org/10.1038/cddis.2013.232
Yelshanskaya MV, Li M, Sobolevsky AI (2014) Structure of an agonist-bound ionotropic glutamate receptor. Science (new York, NY) 345:1070–1074. https://doi.org/10.1126/science.1256508
Zamanillo D et al (1999) Importance of AMPA receptors for hippocampal synaptic plasticity but not for spatial learning. Science (new York, NY) 284:1805–1811. https://doi.org/10.1126/science.284.5421.1805
Zuo J, De Jager PL, Takahashi KA, Jiang W, Linden DJ, Heintz N (1997) Neurodegeneration in Lurcher mice caused by mutation in delta2 glutamate receptor gene. Nature 388:769–773. https://doi.org/10.1038/42009
Acknowledgements
The authors thank Ms. Kaori Takabe, Mr. Takabumi Miyama, Ms. Nobuko Watanabe, Ms. Mai Sato, and Ms. Sayaka Sugimoto at Department of Human Genetics, Yokohama City University Graduate School of Medicine, and S. Kondo at Tokyo University of Science for their technical assistance, and the Drosophila Genetic Resource Center, Kyoto Institute of Technology, for providing the fly strains. They are also grateful to Edanz (https://jp.edanz.com/ac) for editing the English text of a draft of this manuscript. This work was supported by Japan Agency for Medical Research and Development (AMED) [JP20ek0109486, JP21ek0109549, JP21cm0106503, and JP21ek0109493 to N.M.; and JP19ek0109288 to K. Saito]; Japan Society for the Promotion of Science (JSPS) KAKENHI [JP20K16932 to K. Hamanaka, JP 20K17428 to N.T., JP 21k15097 to Y.U., JP20K17936 to A.F., and JP20K07907 to S.M.]; and an intramural Grant of Yokohama City University (to K. Hamanaka).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
439_2021_2416_MOESM1_ESM.png
Fig. S1 Structural considerations of Gly630Arg variation in GluA3 (a) Close-up views of the region around Gly598 in mouse GluA2 (Gly630 in human GluA3) in the structures calculated by FoldX. The main chain Cα atom of Gly598 and the side chain of Ala583 are shown as spheres, and the side chains of the residues predicted to clash with the variant Arg598 are indicated by transparent spheres. (b) The free energy changes due to the variation of p.Gly598Arg in mouse GluA2 (p.Gly630Arg in human GluA3) are shown as in Figure 2d. (PNG 9539 kb)
Rights and permissions
About this article
Cite this article
Hamanaka, K., Miyoshi, K., Sun, JH. et al. Amelioration of a neurodevelopmental disorder by carbamazepine in a case having a gain-of-function GRIA3 variant. Hum Genet 141, 283–293 (2022). https://doi.org/10.1007/s00439-021-02416-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-021-02416-7