Abstract
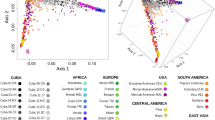
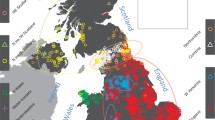
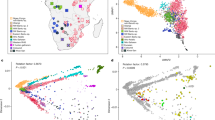
Unlike other European countries, the human population genetics and demographic history of Metropolitan France is surprisingly understudied. In this work, we combined newly genotyped samples from various zones in France with publicly available data and applied both allele frequency and haplotype-based methods to describe the internal structure of this country, using genome-wide single nucleotide polymorphism (SNP) array genotypes. We found out that French Basques, already known for their linguistic uniqueness, are genetically distinct from all other groups and that the populations from southwest France (namely the Gascony region) share a large proportion of their ancestry with Basques. Otherwise, the genetic makeup of the French population is relatively homogeneous and mostly related to Southern and Central European groups. However, a fine-grained, haplotype-based analysis revealed that Bretons slightly separated from the rest of the groups, due mostly to gene flow from the British Isles in a time frame that coincides both historically attested Celtic population movements to this area between the 3th and the ninth centuries CE, but also with a more ancient genetic continuity between Brittany and the British Isles related to the shared drift with hunter-gatherer populations. Haplotype-based methods also unveiled subtle internal structures and connections with the surrounding modern populations, particularly in the periphery of the country.






Similar content being viewed by others
References
Alexander DH, Novembre J (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664. https://doi.org/10.1101/gr.094052.109.vidual
Arias P (1999) The origins of the Neolithic along the Atlantic coast of Continental Europe: a survey. J World Prehist 13:403–464
Auton A, Brooks LD, 1000 Genomes Project Consortium et al (2015) A global reference for human genetic variation. Nature 526:68–74. https://doi.org/10.1038/nature15393
Behr AA, Liu KZ, Liu-Fang G et al (2016) pong: fast analysis and visualization of latent clusters in population genetic data. Bioinformatics 32:2817–2823. https://doi.org/10.1093/bioinformatics/btw327
Biagini SA, Solé-Morata N, Matisoo-Smith E et al (2019) People from Ibiza: an unexpected isolate in the Western Mediterranean. Eur J Hum Genet 27:941–951. https://doi.org/10.1038/s41431-019-0361-1
Blumenthal D (2012) Alsace-Lorraine: a study of the relations of the two provinces to France and to Germany, and a presentation of the just claims of their people (Classic Reprint). Forgotten Books, London
Bycroft C, Fernandez-Rozadilla C, Ruiz-Ponte C et al (2019) Patterns of genetic differentiation and the footprints of historical migrations in the Iberian Peninsula. Nat Commun 10:551. https://doi.org/10.1038/s41467-018-08272-w
Calafell F, Bertranpetit J (1993) A simulation of the genetic history of the Iberian Peninsula. Curr Anthropol 34:735–745
Calafell F, Bertranpetit J (1994) Principal component analysis of gene frequencies and the origin of Basques. Am J Phys Anthropol 93:201–215. https://doi.org/10.1002/ajpa.1330930205
Cambon-Thomsen A, Ohayon E (1988) Practical application of population genetics: the genetic survey “Provinces Françaises.” In: Mayr WR (ed) Advances in forensic haemogenetics. Advances in forensic haemogenetics, vol 2. pp 535–553
Delaneau O, Marchini J, McVeanh GA et al (2014) Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat Commun 5:1–9. https://doi.org/10.1038/ncomms4934
Dray S, Dufour A-B (2007) The ade4 Package: Implementing the duality diagram for ecologists. J Stat Softw. https://doi.org/10.18637/jss.v022.i04
Dubut V, Chollet L, Murail P et al (2004) mtDNA polymorphisms in five French groups: importance of regional sampling. Eur J Hum Genet 12:293–300. https://doi.org/10.1038/sj.ejhg.5201145
Fine JVA (1985) The ancient greeks: a critical history. Belknap Press: an Imprint of Harvard University Press
Forstenzer TR (2016) French Provincial Police and the Fall of the Second Republic: Social Fear and Counterrevolution. Princeton University Press, Princeton
Forster P, Toth A (2003) Toward a phylogenetic chronology of ancient Gaulish, Celtic, and Indo-European. Proc Natl Acad Sci 100:9079–9084. https://doi.org/10.1073/pnas.1331158100
Günther T, Valdiosera C, Malmström H et al (2015) Ancient genomes link early farmers from Atapuerca in Spain to modern-day Basques. Proc Natl Acad Sci USA 112:11917–11922. https://doi.org/10.1073/pnas.1509851112
Haine WS (2000) The history of France. Greenwood Press, Santa Barbara
Hellenthal G (2015) Instruction Manual for “GLOBETROTTER: a program for identifying, dating and describing admixture events in population data” for GLOBETROTTER, pp 1–24
Hellenthal G, Busby GBJ, Band G et al (2014) A genetic atlas of human admixture history. Science (80-) 343:747–751. https://doi.org/10.1126/science.1243518
House GL, Hahn MW (2018) Evaluating methods to visualize patterns of genetic differentiation on a landscape. Mol Ecol Resour 18:448–460. https://doi.org/10.1111/1755-0998.12747
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Kamvar ZN, Brooks JC, Grünwald NJ (2015) Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front Genet. https://doi.org/10.3389/fgene.2015.00208
Karakachoff M, Duforet-Frebourg N, Simonet F et al (2015) Fine-scale human genetic structure in Western France. Eur J Hum Genet 23:831–836. https://doi.org/10.1038/ejhg.2014.175
Kherumian R, Moullec J, Nguyen VC (1967) Groupes sanguins érythrocytaires A2, A1, BO, MN, Rh (CcDE) et sériques, Hp, Tf, Gm dans quatre régions militaires françaises. Bull Mem Soc Anthropol Paris 1:377–384. https://doi.org/10.3406/bmsap.1967.1396
Koch J (2005) Breton Migrations. Celtic culture : a historical encyclopedia. ABC-CLIO, Santa Barbara, pp 275–277
Lao O, Lu TT, Nothnagel M et al (2008) Correlation between genetic and geographic structure in Europe. Curr Biol 18:1241–1248
Lawson DJ, Hellenthal G, Myers S, Falush D (2012) Inference of population structure using dense haplotype data. PLoS Genet 8:e1002453. https://doi.org/10.1371/journal.pgen.1002453
Lazaridis I, Patterson N, Mittnik A et al (2014) Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513:409–413. https://doi.org/10.1038/nature13673
Lazaridis I, Nadel D, Rollefson G et al (2016) Genomic insights into the origin of farming in the ancient Near East. Nature 536:419–424. https://doi.org/10.1038/nature19310
Leslie S, Winney B, Hellenthal G et al (2015) The fine-scale genetic structure of the British population. Nature 519:309–314. https://doi.org/10.1038/nature14230
MacMaster N (1997) Colonial Migrants and Racism Algerians in France, 1900–62. Palgrave Macmillan, London
Martínez-Cruz B, Harmant C, Platt DE et al (2012) Evidence of pre-Roman tribal genetic structure in Basques from uniparentally inherited markers. Mol Biol Evol 29:2211–2222. https://doi.org/10.1093/molbev/mss091
Monnier J (1997) Chapitre 6: L’immigration bretonne en Armorique. In: Monnier J, Cassard J (eds) Toute l’histoire de Bretagne. Skol Vreizh, Morlaix, pp 97–106
Novembre J, Johnson T, Bryc K et al (2008) Genes mirror geography within Europe. Nature 456:98–101
O’Connell J, Gurdasani D, Delaneau O et al (2014) A general approach for haplotype phasing across the full spectrum of relatedness. PLoS Genet 10:e1004234. https://doi.org/10.1371/journal.pgen.1004234
OECD (2017) OECD multi-level governance studies multi-level governance reforms overview of OECD Country Experiences
Olalde I, Mallick S, Patterson N et al (2019) The genomic history of the Iberian Peninsula over the past 8000 years. Science 363:1230–1234. https://doi.org/10.1126/science.aav4040
Patterson N, Price AL, Reich D (2006) Population structure and eigen analysis. PLoS Genet 2:2074–2093. https://doi.org/10.1371/journal.pgen.0020190
Patterson N, Moorjani P, Luo Y et al (2012) Ancient admixture in human history. Genetics 192:1065–1093. https://doi.org/10.1534/genetics.112.145037
Pereira L, Richards M, Goios A et al (2005) High-resolution mtDNA evidence for the late-glacial resettlement of Europe from an Iberian refugium. Genome Res 15:19–24. https://doi.org/10.1101/gr.3182305
Petkova D, Novembre J, Stephens M (2016) Visualizing spatial population structure with estimated effective migration surfaces. Nat Genet 48:94–100. https://doi.org/10.1038/ng.3464
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ramos-Luis E, Blanco-Verea A, Brión M et al (2009) Phylogeography of French male lineages. Forensic Sci Int Genet Suppl Ser 2:439–441
Ramos-Luis E, Blanco-Verea A, Brión M et al (2014) Y-chromosomal DNA analysis in French male lineages. Forensic Sci Int Genet 9:162–168. https://doi.org/10.1016/j.fsigen.2013.12.008
Raveane A, Aneli S, Montinaro F et al (2019) Population structure of modern-day Italians reveals patterns of ancient and archaic ancestries in Southern Europe. Sci Adv 5:1–13. https://doi.org/10.1126/sciadv.aaw3492
Richard C, Richard C, Pennarun E et al (2007) An mtDNA perspective of French genetic variation. Ann Hum Biol 34:68–79. https://doi.org/10.1080/03014460601076098
Rodríguez-Ezpeleta N, Álvarez-Busto J, Imaz L et al (2010) High-density SNP genotyping detects homogeneity of Spanish and French Basques, and confirms their genomic distinctiveness from other European populations. Hum Genet 128:113–117. https://doi.org/10.1007/s00439-010-0833-4
RStudio Team (2015) RStudio Tea: integrated development for R
Sowerwine C (2009) France since 1870: Culture, Society and the Making of the Republic, 2nd edn. Palgrave Macmillan, London
Tian C, Kosoy R, Nassir R et al (2009) European population genetic substructure: further definition of ancestry informative markers for distinguishing among diverse European ethnic groups. Mol Med 15:371–383. https://doi.org/10.2119/molmed.2009.00094
Wickham H (2007) Reshaping data with the reshape Package. J Stat Softw. https://doi.org/10.18637/jss.v021.i12
Wickham H (2009) ggplot2. Springer, New York, New York
Zuazo K (2010) El euskera y sus dialectos. Alberdania, Zarautz
Acknowledgments
We thank Dr. Véronique Van Huffel, from the “Unité de Polymorphisme Génétique” of the “Institut National de la Transfusion Sanguine” in Paris for collecting most of the samples used in this paper, and Inés Quintela and the National Genotyping Center (CEGEN – USC) for their assistance in genotyping the samples. Of course, this work would not have been possible without the kind collaboration of all the sample donors. Funding was provided by the Agencia Estatal de Investigación and Fondo Europeo de Desarollo Regional (FEDER) (grant CGL2016-75389-P), Agència de Gestió d’Ajuts Universitaris i de la Recerca (Generalitat de Catalunya) Grant 2014SGR2866, and “Unidad de Excelencia María de Maeztu”, funded by the MINECO (ref: MDM-2014–0370). SAB was supported by the Agencia Estatal de Investigación FPI grant BES-2014–069224.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Biagini, S.A., Ramos-Luis, E., Comas, D. et al. The place of metropolitan France in the European genomic landscape. Hum Genet 139, 1091–1105 (2020). https://doi.org/10.1007/s00439-020-02158-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-020-02158-y