Abstract
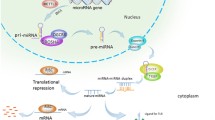
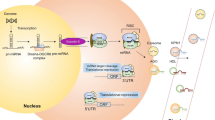
As an important class of non-coding regulatory RNAs, microRNAs (miRNAs) play a key role in a range of biological processes. These molecules serve as post-transcriptional regulators of gene expression and their regulatory activity has been implicated in disease pathophysiology and pharmacological traits. We sought to investigate the impact of miRNAs on cellular proliferation to gain insight into the molecular basis of complex traits that depend on cellular growth, including, most prominently, cancer. We examined the relationship between miRNA expression and intrinsic cellular growth (iGrowth) in the HapMap lymphoblastoid cell lines derived from individuals of different ethnic backgrounds. We found a substantial enrichment for miRNAs (53 miRNAs, FDR < 0.05) correlated with cellular proliferation in pooled CEU (Caucasian of northern and western European descent) and YRI (individuals from Ibadan, Nigeria) samples. Specifically, 119 miRNAs (59 %) were significantly correlated with iGrowth in YRI; of these miRNAs, 18 were correlated with iGrowth in CEU. To gain further insight into the effect of miRNAs on cellular proliferation in cancer, we showed that over-expression of miR-22, one of the top iGrowth-associated miRNAs, leads to growth inhibition in an ovarian cancer cell line (SKOV3). Furthermore, over-expression of miR-22 down-regulates the expression of its target genes (MXI1 and SLC25A37) in this ovarian cancer cell line, highlighting an miRNA-mediated regulatory network potentially important for cellular proliferation. Importantly, our study identified miRNAs that can be used as molecular targets in cancer therapy.




Similar content being viewed by others
Abbreviations
- miRNA:
-
microRNA
- iGrowth:
-
Intrinsic cellular growth
- LCLs:
-
Lymphoblastoid cell lines
- CEU:
-
Centre d’Etude du Polymorphisme Humain (CEPH) people from Utah, USA
- YRI:
-
Yoruba people from Ibadan, Nigeria
- MEM:
-
Mixed effects model averaging
- FDR:
-
False discovery rate
- DAVID:
-
Database for Annotation, Visualization and Integrated Discovery
- GO:
-
Gene ontology
- DOHH:
-
Deoxyhypusine hydroxylase
References
Bar N, Dikstein R (2010) miR-22 forms a regulatory loop in PTEN/AKT pathway and modulates signaling kinetics. PLoS One 5:e10859
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Chen W, Paradkar PN, Li L, Pierce EL, Langer NB, Takahashi-Makise N, Hyde BB, Shirihai OS, Ward DM, Kaplan J et al (2009) Abcb10 physically interacts with mitoferrin-1 (Slc25a37) to enhance its stability and function in the erythroid mitochondria. Proc Natl Acad Sci USA 106:16263–16268
Dang Y, Luo D, Rong M, Chen G (2013) Underexpression of miR-34a in hepatocellular carcinoma and its contribution towards enhancement of proliferating inhibitory effects of agents targeting c-MET. PLoS One 8:e61054
Duan S, Huang RS, Zhang W, Bleibel WK, Roe CA, Clark TA, Chen TX, Schweitzer AC, Blume JE, Cox NJ et al (2008) Genetic architecture of transcript-level variation in humans. Am J Hum Genet 82:1101–1113
Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA, Belmont JW, Boudreau A, Hardenbol P, Leal SM et al (2007) A second generation **human haplotype map of over 3.1 million SNPs. Nature 449:851–861
Fonseca-Sanchéz MA, Pérez-Plasencia C, Fernández-Retana J, Arechaga-Ocampo E, Marchat LA, Rodríguez-Cuevas S, Bautista-Piña V, Arellano-Anaya ZE, Flores-Pérez A, Diaz-Chávez J et al (2013) microRNA-18b is upregulated in breast cancer and modulates genes involved in cell migration. Oncol Rep 30:2399–2410
Gamazon ER, Ziliak D, Im HK, LaCroix B, Park DS, Cox NJ, Huang RS (2012) Genetic architecture of microRNA expression: implications for the transcriptome and complex traits. Am J Hum Genet 90:1046–1063
González-Gugel E, Villa-Morales M, Santos J, Bueno MJ, Malumbres M, Rodríguez-Pinilla SM, Piris MÁ, Fernández-Piqueras J (2013) Down-regulation of specific miRNAs enhances the expression of the gene smoothened and contributes to T-cell lymphoblastic lymphoma development. Carcinogenesis 34(4):902–908
He J, Wu J, Xu N, Xie W, Li M, Li J, Jiang Y, Yang BB, Zhang Y (2012) MiR-210 disturbs mitotic progression through regulating a group of mitosis-related genes. Nucleic Acids Res 41(1):498–508. doi:10.1093/nar/gks995
Hsiao C-P, Wang D, Kaushal A, Saligan L (2013) Mitochondria-related gene expression changes are associated with fatigue in patients with nonmetastatic prostate cancer receiving external beam radiation therapy. Cancer Nurs 36(3):189–197
Huang RS, Kistner EO, Bleibel WK, Shukla SJ, Dolan ME (2007) Effect of population and gender on chemotherapeutic agent-induced cytotoxicity. Mol Cancer Ther 6:31–36
Hurlin PJ, Dezfouli S (2004) Functions of myc:max in the control of cell proliferation and tumorigenesis. Int Rev Cytol 238:183–226
Huang Z-P, Chen J, Seok HY, Zhang Z, Kataoka M, Hu X, Wang D-Z (2013) MicroRNA-22 regulates cardiac hypertrophy and remodeling in response to stress. Circ Res 112:1234–1243
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Hwang H-W, Mendell JT (2006) MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 94:776–780
Im HK, Gamazon ER, Stark AL, Huang RS, Cox NJ, Dolan ME (2012) Mixed effects modeling of proliferation rates in cell-based models: consequence for pharmacogenomics and cancer. PLoS Genet 8:e1002525
Imam JS, Buddavarapu K, Lee-Chang JS, Ganapathy S, Camosy C, Chen Y, Rao MK (2010) MicroRNA-185 suppresses tumor growth and progression by targeting the Six1 oncogene in human cancers. Oncogene 29:4971–4979
Kashat M, Azzouz L, Sarkar SH, Kong D, Li Y, Sarkar FH (2012) Inactivation of AR and Notch-1 signaling by miR-34a attenuates prostate cancer aggressiveness. Am J Transl Res 4:432–442
Kaur K, Pandey AK, Srivastava S, Srivastava AK, Datta M (2011) Comprehensive miRNome and in silico analyses identify the Wnt signaling pathway to be altered in the diabetic liver. Mol Biosyst 7:3234–3244
Lal A, Thomas MP, Altschuler G, Navarro F, O’Day E, Li XL, Concepcion C, Han Y-C, Thiery J, Rajani DK et al (2011) Capture of microRNA-bound mRNAs identifies the tumor suppressor miR-34a as a regulator of growth factor signaling. PLoS Genet 7:e1002363
Li X-F, Yan P-J, Shao Z-M (2009) Downregulation of miR-193b contributes to enhance urokinase-type plasminogen activator (uPA) expression and tumor progression and invasion in human breast cancer. Oncogene 28:3937–3948
Li J, Liang S, Yu H, Zhang J, Ma D, Lu X (2010) An inhibitory effect of miR-22 on cell migration and invasion in ovarian cancer. Gynecol Oncol 119:543–548
Liu L, Jiang Y, Zhang H, Greenlee AR, Yu R, Yang Q (2010) miR-22 functions as a micro-oncogene in transformed human bronchial epithelial cells induced by anti-benzo[a]pyrene-7,8-diol-9,10-epoxide. Toxicol In Vitr 24:1168–1175
Morris KV, Chan SW-L, Jacobsen SE, Looney DJ (2004) Small interfering RNA-induced transcriptional gene silencing in human cells. Science 305:1289–1292
Pandey DP, Picard D (2009) miR-22 inhibits estrogen signaling by directly targeting the estrogen receptor α mRNA. Mol Cell Biol 29:3783–3790
Pickrell JK, Marioni JC, Pai AA, Degner JF, Engelhardt BE, Nkadori E, Veyrieras J-B, Stephens M, Gilad Y, Pritchard JK (2010) Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464:768–772
Puisségur M-P, Mazure NM, Bertero T, Pradelli L, Grosso S, Robbe-Sermesant K, Maurin T, Lebrigand K, Cardinaud B, Hofman V et al (2011) miR-210 is overexpressed in late stages of lung cancer and mediates mitochondrial alterations associated with modulation of HIF-1 activity. Cell Death Differ 18:465–478
Qi J, Rice SJ, Salzberg AC, Runkle EA, Liao J, Zander DS, Mu D (2012) MiR-365 regulates lung cancer and developmental gene thyroid transcription factor 1. Cell Cycle 11:177–186
Ramberg H, Alshbib A, Berge V, Svindland A, Taskén KA (2011) Regulation of PBX3 expression by androgen and Let-7d in prostate cancer. Mol Cancer 10:50
Schafer KA (1998) The cell cycle: a review. Vet Pathol 35:461–478
Song Y-X, Yue Z-Y, Wang Z-N, Xu Y-Y, Luo Y, Xu H-M, Zhang X, Jiang L, Xing C-Z, Zhang Y (2011a) MicroRNA-148b is frequently down-regulated in gastric cancer and acts as a tumor suppressor by inhibiting cell proliferation. Mol Cancer 10:1
Song Y, Xu Y, Wang Z, Chen Y, Yue Z, Gao P, Xing C, Xu H (2011b) MicroRNA-148b suppresses cell growth by targeting cholecystokinin-2 receptor in colorectal cancer. Int J Cancer 1051:1042–1051
Stranger BE, Nica AC, Forrest MS, Dimas A, Bird CP, Beazley C, Ingle CE, Dunning M, Flicek P, Koller D et al (2007) Population genomics of human gene expression. Nat Genet 39:1217–1224
Stark AL, Zhang W, Mi S, Duan S, O’Donnell PH, Huang RS, Dolan ME (2010) Heritable and non-genetic factors as variables of pharmacologic phenotypes in lymphoblastoid cell lines. Pharmacogenomics J 10:505–512
Storey JD, Tibshirani R (2003) Statistical significance for genome-wide studies. Proc Natl Acad Sci USA 100:9440–9445
Suzuki HI, Miyazono K (2011) Emerging complexity of microRNA generation cascades. J Biochem 149:15–25
Takata A, Otsuka M, Kojima K, Yoshikawa T, Kishikawa T, Yoshida H, Koike K (2011) MicroRNA-22 and microRNA-140 suppress NF-κB activity by regulating the expression of NF-κB coactivators. Biochem Biophys Res Commun 411:826–831
The 1000 Genomes Project Consortium (2010) A map of human genome variation from population-scale sequencing. Nature 467:1061–1073
Ting Y, Medina DJ, Strair RK, Schaar DG (2010) Differentiation-associated miR-22 represses Max expression and inhibits cell cycle progression. Biochem Biophys Res Commun 394:606–611
Tsoi M, Laguë M-N, Boyer A, Paquet M, Nadeau M-È, Boerboom D (2013) Anti-VEGFA therapy reduces tumor growth and extends survival in a murine model of ovarian granulosa cell tumor. Transl Oncol 6:226–233
Tsuchiya N, Izumiya M, Ogata-Kawata H, Okamoto K, Fujiwara Y, Nakai M, Okabe A, Schetter AJ, Bowman ED, Midorikawa Y et al (2011) Tumor suppressor miR-22 determines p53-dependent cellular fate through post-transcriptional regulation of p21. Cancer Res 71:4628–4639
Whitfield ML, George LK, Grant GD, Perou CM (2006) Common markers of proliferation. Nat Rev Cancer 6:99–106
Xiong J, Yu D, Wei N, Fu H, Cai T, Huang Y, Wu C, Zheng X, Du Q, Lin D et al (2010a) An estrogen receptor alpha suppressor, microRNA-22, is downregulated in estrogen receptor alpha-positive human breast cancer cell lines and clinical samples. FEBS J 277:1684–1694
Xiong J, Du Q, Liang Z (2010b) Tumor-suppressive microRNA-22 inhibits the transcription of E-box-containing c-Myc target genes by silencing c-Myc binding protein. Oncogene 29:4980–4988
Xu D, Takeshita F, Hino Y, Fukunaga S, Kudo Y, Tamaki A, Matsunaga J, Takahashi R-U, Takata T, Shimamoto A et al (2011) miR-22 represses cancer progression by inducing cellular senescence. J Cell Biol 193:409–424
Yamakuchi M, Yagi S, Ito T, Lowenstein CJ (2011) MicroRNA-22 regulates hypoxia signaling in colon cancer cells. PLoS One 6:e20291
Yu T, Wang X, Gong R, Li A, Yang S, Cao Y, Wen Y, Wang C, Yi X (2009) The expression profile of microRNAs in a model of 7,12-dimethyl-benz[a]anthrance-induced oral carcinogenesis in Syrian hamster. J Exp Clin Cancer Res 28:64
Zervos AS, Gyuris J, Brent R (1993) Mxi1, a protein that specifically interacts with Max to bind Myc-Max recognition sites. Cell 72:223–232
Zhao C, Sun G, Ye P, Li S, Shi Y (2013) MicroRNA let-7d regulates the TLX/microRNA-9 cascade to control neural cell fate and neurogenesis. Sci Rep 3:1329
Zhang J, Yang Y, Yang T, Liu Y, Li A, Fu S, Wu M, Pan Z, Zhou W (2010) microRNA-22, downregulated in hepatocellular carcinoma and correlated with prognosis, suppresses cell proliferation and tumourigenicity. Br J Cancer 103:1215–1220
Zhang W, Duan S, Kistner EO, Bleibel WK, Huang RS, Clark TA, Chen TX, Schweitzer AC, Blume JE, Cox NJ et al (2008) Evaluation of genetic variation contributing to differences in gene expression between populations. Am J Hum Genet 82:631–640
Zheng Z-F, Su H-F, Zou Y, Peng Z, Wu S-X (2011) Expression profiles of microRNAs in radioresistant esophageal cell line. Zhonghua Yi Xue Za Zhi 91:639–642
Acknowledgments
We are grateful for the excellent technical support provided by Pharmacogenomics of Anti-cancer Agent Research (PAAR) group cell line core. This study was supported by National Institute of Health/National Cancer Institute grant (R21 CA139278) and by National Institute of Health/National Institute of General Medical Sciences grant (UO1GM61393). RSH also received support from National Institute of Health/National Institute of General Medical Sciences grant (K08GM089941), Circle of Service Foundation Early Career Investigator award, University of Chicago Cancer Center Support Grant (#P30 CA14599), Breast Cancer SPORE Career Development Award (CA125183) a Conquer Cancer Foundation of ASCO Translational Research Professorship award In Memory of Merrill J. Egorin, MD, and pilot grant from National Institute of Health/National Center Advancing Translational Sciences grant (UL1RR024999).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lenkala, D., LaCroix, B., Gamazon, E.R. et al. The impact of microRNA expression on cellular proliferation. Hum Genet 133, 931–938 (2014). https://doi.org/10.1007/s00439-014-1434-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-014-1434-4