Abstract
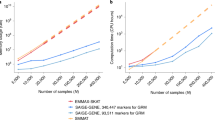
As the ability to measure dense genetic markers approaches the limit of the DNA sequence itself, taking advantage of possible clustering of genetic variants in, and around, a gene would benefit genetic association analyses, and likely provide biological insights. The greatest benefit might be realized when multiple rare variants cluster in a functional region. Several statistical tests have been developed, one of which is based on the popular Kulldorff scan statistic for spatial clustering of disease. We extended another popular spatial clustering method—Tango’s statistic—to genomic sequence data. An advantage of Tango’s method is that it is rapid to compute, and when single test statistic is computed, its distribution is well approximated by a scaled χ 2 distribution, making computation of p values very rapid. We compared the Type-I error rates and power of several clustering statistics, as well as the omnibus sequence kernel association test. Although our version of Tango’s statistic, which we call “Kernel Distance” statistic, took approximately half the time to compute than the Kulldorff scan statistic, it had slightly less power than the scan statistic. Our results showed that the Ionita-Laza version of Kulldorff’s scan statistic had the greatest power over a range of clustering scenarios.





Similar content being viewed by others
References
Asimit J, Zeggini E (2010) Rare variant association analysis methods for complex traits. Annu Rev Genet 44:293–308
Bansal V, Libiger O, Torkamani A, Schork NJ (2010) Statistical analysis strategies for association studies involving rare variants. Nat Rev Genet 11(11):773–785
Basu S, Pan W (2010) Comparison of statistical tests for disease association with rare variants. Genet Epidem 35:606–619
Breslow NE, Day NE (1980) The analysis of case–control studies. Inter Agency Res Cancer, Lyon
Fier H, Won S, Prokopenko D, Alchawa T, Ludwig KU, Fimmers R, Silverman EK, Pagano M, Mangold E, Lange C (2012) ‘Location, location, location’: a spatial approach for rare variant analysis and an application to a study on non-syndromic cleft lip with or without cleft palate. Bioinformatics 28(23):3027–3033
Ionita-Laza I, Makarov V, Buxbaum JD (2012) Scan-statistic approach identifies clusters of rare disease variants in LRP2, a gene linked and associated with autism spectrum disorders, in three datasets. Amer J Hum Genet 90(6):1002–1013
Kulldorff M (2007) A spatial scan statistic. Commun Stat Theo Meth 26:1481–1496
Kwee LC, Liu D, Lin X, Ghosh D, Epstein MP (2008) A powerful and flexible multilocus association test for quantitative traits. Am J Hum Genet 82(2):386–397
Lee S, Emond MJ, Bamshad MJ, Barnes KC, Rieder MJ, Nickerson DA, Christiani DC, Wurfel MM, Lin X (2012a) Optimal unified approach for rare-variant association testing with application to small-sample case–control whole-exome sequencing studies. Am J Hum Genet 91(2):224–237
Lee S, Wu MC, Lin X (2012b) Optimal tests for rare variant effects in sequencing association studies. Biostatistics 13(4):762–775
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Can Res 27:209–220
Ruppert D, Wand P, Carroll R (2005) Semi parametric regression. Cambridge University Press, Cambridge
Tango T (1984) The detection of clustering of disease in time. Biometrics 40:15–26
Tango T (2000) A test for spatial disease clustering adjusted for multiple testing. Stat Med 19(2):191–204
Tango T (2010) Statistical methods for disease clustering. Springer, New York
Wu MC, Lee S, Cai T, Li Y, Boehnke M, Lin X (2011) Rare-variant association testing for sequencing data with the sequence kernel association test. Am J Hum Genet 89(1):82–93
Acknowledgments
This research was supported by the US Public Health Service, National Institutes of Health (NIH), contract Grant Number GM065450 (DJS, JPS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schaid, D.J., Sinnwell, J.P., McDonnell, S.K. et al. Detecting genomic clustering of risk variants from sequence data: cases versus controls. Hum Genet 132, 1301–1309 (2013). https://doi.org/10.1007/s00439-013-1335-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-013-1335-y