Abstract
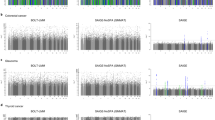
We applied a recently developed multilocus association testing method (localized haplotype clustering) to Wellcome Trust Case Control Consortium data (14,000 cases of seven common diseases and 3,000 shared controls genotyped on the Affymetrix 500 K array). After rigorous data quality filtering, we identified three disease-associated loci with strong statistical support from localized haplotype cluster tests but with only marginal significance in single marker tests. These loci are chromosomes 10p15.1 with type 1 diabetes (p = 5.1 × 10−9), 12q15 with type 2 diabetes (p = 1.9 × 10−7) and 15q26.2 with hypertension (p = 2.8 × 10−8). We also detected the association of chromosome 9p21.3 with type 2 diabetes (p = 2.8 × 10−8), although this locus did not pass our stringent genotype quality filters. The association of 10p15.1 with type 1 diabetes and 9p21.3 with type 2 diabetes have both been replicated in other studies using independent data sets. Overall, localized haplotype cluster analysis had better success detecting disease associated variants than a previous single-marker analysis of imputed HapMap SNPs. We found that stringent application of quality score thresholds to genotype data substantially reduced false-positive results arising from genotype error. In addition, we demonstrate that it is possible to simultaneously phase 16,000 individuals genotyped on genome-wide data (450 K markers) using the Beagle software package.


Similar content being viewed by others
References
Bektas A, Hughes JN, Warram JH, Krolewski AS, Doria A (2001) Type 2 diabetes locus on 12q15. Further mapping and mutation screening of two candidate genes. Diabetes 50:204–208
Browning BL, Browning SR (2007a) Efficient multilocus association testing for whole genome association studies using localized haplotype clustering. Genet Epidemiol 31:365–375
Browning SR (2006) Multilocus association mapping using variable-length Markov chains. Am J Hum Genet 78:903–913
Browning SR, Browning BL (2007b) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81:1084–1097
Clayton D, Leung HT (2007) An R package for analysis of whole-genome association studies. Hum Hered 64:45–51
Clayton DG, Walker NM, Smyth DJ, Pask R, Cooper JD, Maier LM, Smink LJ, Lam AC, Ovington NR, Stevens HE, Nutland S, Howson JM, Faham M, Moorhead M, Jones HB, Falkowski M, Hardenbol P, Willis TD, Todd JA (2005) Population structure, differential bias and genomic control in a large-scale, case–control association study. Nat Genet 37:1243–1246
Ehm MG, Karnoub MC, Sakul H, Gottschalk K, Holt DC, Weber JL, Vaske D, Briley D, Briley L, Kopf J, McMillen P, Nguyen Q, Reisman M, Lai EH, Joslyn G, Shepherd NS, Bell C, Wagner MJ, Burns DK (2000) Genomewide search for type 2 diabetes susceptibility genes in four American populations. Am J Hum Genet 66:1871–1881
Joe B, Letwin NE, Garrett MR, Dhindaw S, Frank B, Sultana R, Verratti K, Rapp JP, Lee NH (2005) Transcriptional profiling with a blood pressure QTL interval-specific oligonucleotide array. Physiol Genomics 23:318–326
Lowe CE, Cooper JD, Brusko T, Walker NM, Smyth DJ, Bailey R, Bourget K, Plagnol V, Field S, Atkinson M, Clayton DG, Wicker LS, Todd JA (2007) Large-scale genetic fine mapping and genotype-phenotype associations implicate polymorphism in the IL2RA region in type 1 diabetes. Nat Genet 39:1074–1082
Marchini J, Howie B, Myers S, McVean G, Donnelly P (2007) A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39:906–913
Matsuzaki H, Dong S, Loi H, Di X, Liu G, Hubbell E, Law J, Berntsen T, Chadha M, Hui H, Yang G, Kennedy GC, Webster TA, Cawley S, Walsh PS, Jones KW, Fodor SP, Mei R (2004) Genotyping over 100,000 SNPs on a pair of oligonucleotide arrays. Nat Methods 1:109–111
Qu HQ, Montpetit A, Ge B, Hudson TJ, Polychronakos C (2007) Toward further mapping of the association between the IL2RA locus and type 1 diabetes. Diabetes 56:1174–1176
R Development Core Team (2006) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI, Chen H, Roix JJ, Kathiresan S, Hirschhorn JN, Daly MJ, Hughes TE, Groop L, Altshuler D, Almgren P, Florez JC, Meyer J, Ardlie K, Bengtsson Bostrom K, Isomaa B, Lettre G, Lindblad U, Lyon HN, Melander O, Newton-Cheh C, Nilsson P, Orho-Melander M, Rastam L, Speliotes EK, Taskinen MR, Tuomi T, Guiducci C, Berglund A, Carlson J, Gianniny L, Hackett R, Hall L, Holmkvist J, Laurila E, Sjogren M, Sterner M, Surti A, Svensson M, Svensson M, Tewhey R, Blumenstiel B, Parkin M, Defelice M, Barry R, Brodeur W, Camarata J, Chia N, Fava M, Gibbons J, Handsaker B, Healy C, Nguyen K, Gates C, Sougnez C, Gage D, Nizzari M, Gabriel SB, Chirn GW, Ma Q, Parikh H, Richardson D, Ricke D, Purcell S (2007) Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316:1331–1336
Schaid DJ (2004) Evaluating associations of haplotypes with traits. Genet Epidemiol 27:348–364
Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL, Erdos MR, Stringham HM, Chines PS, Jackson AU, Prokunina-Olsson L, Ding CJ, Swift AJ, Narisu N, Hu T, Pruim R, Xiao R, Li XY, Conneely KN, Riebow NL, Sprau AG, Tong M, White PP, Hetrick KN, Barnhart MW, Bark CW, Goldstein JL, Watkins L, Xiang F, Saramies J, Buchanan TA, Watanabe RM, Valle TT, Kinnunen L, Abecasis GR, Pugh EW, Doheny KF, Bergman RN, Tuomilehto J, Collins FS, Boehnke M (2007) A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316:1341–1345
Servin B, Stephens M (2007) Imputation-based analysis of association studies: candidate regions and quantitative traits. PLoS Genet 3:e114
The International HapMap Consortium (2007) A second generation human haplotype map of over 3.1 million SNPs. Nature 449:851–861
The Wellcome Trust Case Control Consortium (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447:661–678
Vella A, Cooper JD, Lowe CE, Walker N, Nutland S, Widmer B, Jones R, Ring SM, McArdle W, Pembrey ME, Strachan DP, Dunger DB, Twells RC, Clayton DG, Todd JA (2005) Localization of a type 1 diabetes locus in the IL2RA/CD25 region by use of tag single-nucleotide polymorphisms. Am J Hum Genet 76:773–779
Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS, Lango H, Timpson NJ, Perry JR, Rayner NW, Freathy RM, Barrett JC, Shields B, Morris AP, Ellard S, Groves CJ, Harries LW, Marchini JL, Owen KR, Knight B, Cardon LR, Walker M, Hitman GA, Morris AD, Doney AS, McCarthy MI, Hattersley AT (2007) Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316:1336–1341
Acknowledgments
This study makes use of data generated by the Wellcome Trust Case Control Consortium. A full list of the investigators who contributed to the generation of the data is available from http://www.wtccc.org.uk. Funding for the Wellcome Trust Case Control Consortium project was provided by the Wellcome Trust under award 076113. The authors thank three anonymous reviewers for their comments, which helped improve the manuscript, Werner Schmidt for his patient assistance with our demanding computing requirements and Hin-Tak Leung for pointing us to his software for creating genotype cluster plots for the WTCCC data set. This work was supported by a grant from the University of Auckland Research Committee (SRB), and by NIH grant 3R01GM075091-02S1 (SRB and BLB). Nutrigenomics New Zealand is a collaboration between AgResearch Ltd., Crop & Food Research, HortResearch and The University of Auckland, with funding through the Foundation for Research Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Browning, B.L., Browning, S.R. Haplotypic analysis of Wellcome Trust Case Control Consortium data. Hum Genet 123, 273–280 (2008). https://doi.org/10.1007/s00439-008-0472-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-008-0472-1