Abstract
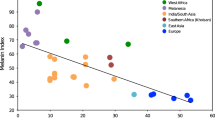
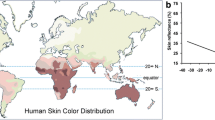
Skin pigmentation is a human phenotype that varies greatly among human populations and it has long been speculated that this variation is adaptive. We therefore expect the genes that contribute to these large differences in phenotype to show large allele frequency differences among populations and to possibly harbor signatures of positive selection. To identify the loci that likely contribute to among-population human skin pigmentation differences, we measured allele frequency differentiation among Europeans, Chinese and Africans for 24 human pigmentation genes from 2 publicly available, large scale SNP data sets. Several skin pigmentation genes show unusually large allele frequency differences among these populations. To determine whether these allele frequency differences might be due to selection, we employed a within-population test based on long-range haplotype structure and identified several outliers that have not been previously identified as putatively adaptive. Most notably, we identify the DCT gene as a candidate for recent positive selection in the Chinese. Moreover, our analyses suggest that it is likely that different genes are responsible for the lighter skin pigmentation found in different non-African populations.


Similar content being viewed by others
References
Beaumont MA, Balding DJ (2004) Identifying adaptive genetic divergence among populations from genome scans. Mol Ecol 13:969–980
Bersaglieri T, Sabeti PC, Patterson N, Vanderploeg T, Schaffner SF, Drake JA, Rhodes M, Reich DE, Hirschhorn JN (2004) Genetic signatures of strong recent positive selection at the lactase gene. Am J Hum Genet 74:1111–1120
Costin GE, Valencia JC, Wakamatsu K, Ito S, Solano F, Milac AL, Vieira WD, Yamaguchi Y, Rouzaud F, Petrescu AJ, Lamoreux ML, Hearing VJ (2005) Mutations in dopachrome tautomerase (Dct) affect eumelanin/pheomelanin synthesis, but do not affect intracellular trafficking of the mutant protein. Biochem J 391:249–259
Darwin C (1871) The descent of man. Princeton University Press, Princeton
del Marmol V, Beermann F (1996) Tyrosinase and related proteins in mammalian pigmentation. FEBS Lett 381:165–168
Guyonneau L, Murisier F, Rossier A, Moulin A, Beermann F (2004) Melanocytes and pigmentation are affected in dopachrome tautomerase knockout mice. Mol Cell Biol 24:3396–3403
Harding RM, Healy E, Ray AJ, Ellis NS, Flanagan N, Todd C, Dixon C, Sajantila A, Jackson IJ, Birch-Machin MA, Rees JL (2000) Evidence for variable selective pressures at MC1R. Am J Hum Genet 66:1351–1361
Hinds DA, Stuve LL, Nilsen GB, Halperin E, Eskin E, Ballinger DG, Frazer KA, Cox DR (2005) Whole-genome patterns of common DNA variation in three human populations. Science 307:1072–1079
Izagirre N, Garcia I, Junquera C, de la Rua C, Alonso S (2006) A scan for signatures of positive selection in candidate loci for skin pigmentation in humans. Mol Biol Evol 23:1697–1706
Jablonski NG (2004) The evolution of human skin and skin color. Ann Rev Anthro 33:585–623
Kong A, Gudbjartsson D, Sainz J, Jonsdottir G, Gudjonsson S, Richardsson B, Sigurdardottir S, Barnard J, Hallbeck B, Masson G, Shlien A, Palsson S, Frigge M, Thorgeirsson T, Gulcher J, Stefansson K (2002) A high-resolution recombination map of the human genome. Nat Genet 31:241–247
Lamason RL, Mohideen M-APK, Mest JR, Wong AC, Norton HL, Aros MC, Jurynec MJ, Mao X, Humphreville VR, Humbert JE, Sinha S, Moore JL, Jagadeeswaran P, Zhao W, Ning G, Makalowska I, McKeigue PM, O’Donnell D, Kittles R, Parra EJ, Mangini NJ, Grunwald DJ, Shriver MD, Canfield VA, Cheng KC (2005) SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310:1782–1786
Lewontin RC (1972) The apportionment of human diversity. Evol Biol 6:391–398
Makova K, Norton H (2005) Worldwide polymorphism at the MC1R locus and normal pigmentation variation in humans. Peptides 26:1901–1908
Makova KD, Ramsay M, Jenkins T, Li W-H (2001) Human DNA sequence variation in a 6.6-kb region containing the melanocortin 1 receptor promoter. Genetics 158:1253–1268
Neel JV (1962) Diabetes mellitus: a “thrifty” genotype rendered detrimental by “progress”? Bull WHO 77:694–703
Pollinger JP, Bustamante CD, Fledel-Alon A, Schmutz S, Gray MM, Wayne RK (2005) Selective sweep mapping of genes with large phenotypic effects. Genome Res 15:1809–1819
Rana BK, Hewett-Emmett D, Jin L, Chang BHJ, Sambuughin N, Lin M, Watkins S, Bamshad M, Jorde LB, Ramsay M, Jenkins T, Li W-H (1999) High Polymorphism at the human melanocortin 1 receptor locus. Genetics 151:1547–1557
Rees JL (2003) Genetics of hair and skin color. Annu Rev Genet 37:67–90
Relethford JH (2002) Apportionment of global human genetic diversity based on craniometrics and skin color. Am J Phys Anthropol 118:393–398
Sabeti PC, Reich DE, Higgins JM, Levine HZP, Richter DJ, Schaffner SF, Gabriel SB, Platko JV, Patterson NJ, McDonald GJ, Ackerman HC, Campbell SJ, Altshuler D, Cooper R, Kwiatkowski D, Ward R, Lander ES (2002) Detecting recent positive selection in the human genome from haplotype structure. Nature 419:832–837
Sabeti PC, Schaffner SF, Fry B, Lohmueller J, Varilly P, Shamovsky O, Palma A, Mikkelsen TS, Altshuler D, Lander ES (2006) Positive natural selection in the human lineage. Science 312:1614–1620
Soejima M, Tachida H, Ishida T, Sano A, Koda Y (2006) Evidence for recent positive selection at the human AIM1 locus in a European population. Mol Biol Evol 23:179–188
Sturm RA, Teasdale RD, Box NF (2001) Human pigmentation genes: identification, structure and consequences of polymorphic variation. Gene 277:49–62
Tang K, Wong LP, Lee EJD, Chong SS, Lee CGL (2004) Genomic evidence for recent positive selection at the human MDR1 gene locus. Hum Mol Genet 13:783–797
The International HapMap Consortium (2005) A haplotype map of the human genome. Nature 437:1299
Tishkoff SA, Varkonyi R, Cahinhinan N, Abbes S, Argyropoulos G, Destro-Bisol G, Drousiotou A, Dangerfield B, Lefranc G, Loiselet J, Piro A, Stoneking M, Tagarelli A, Tagarelli G, Touma EH, Williams SM, Clark AG (2001) Haplotype diversity and linkage disequilibrium at human G6PD: recent origin of alleles that confer malarial resistance. Science 293:455–462
Tomita Y, Suzuki T (2004) Genetics of pigmentary disorders. Am J Med Genet 131C:75–81
Toyofuku K, Wada I, Spritz RA, Hearing VJ (2001a) The molecular basis of oculocutaneous albinism type 1 (OCA1): sorting failure and degradation of mutant tyrosinases results in a lack of pigmentation. Biochem J 355:259–269
Toyofuku K, Wada I, Valencia JC, Kushimoto T, Ferrans VJ, Hearing VJ (2001b) Oculocutaneous albinism types 1 and 3 are ER retention diseases: mutation of tyrosinase or Tyrp1 can affect the processing of both mutant and wild-type proteins. FASEB J 15:2149–2161
Voight BF, Kudaravalli S, Wen X, Pritchard JK (2006) A map of recent positive selection in the human genome. PLoS Biol 4:e72
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Acknowledgments
We thank Ed Green for technical assistance; and David Hughes, Naim Matasci, Susan Ptak and Michael Lachmann for useful discussion. Supported by the Max Planck Society.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Myles, S., Somel, M., Tang, K. et al. Identifying genes underlying skin pigmentation differences among human populations. Hum Genet 120, 613–621 (2007). https://doi.org/10.1007/s00439-006-0256-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-006-0256-4