Abstract
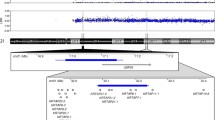
The human and chimpanzee genomes are distinguishable in terms of ten gross karyotypic differences including nine pericentric inversions and a chromosomal fusion. Seven of these large pericentric inversions are chimpanzee-specific whereas two of them, involving human chromosomes 1 and 18, were fixed in the human lineage after the divergence of humans and chimpanzees. We have performed detailed molecular and computational characterization of the breakpoint regions of the human-specific inversion of chromosome 1. FISH analysis and sequence comparisons together revealed that the pericentromeric region of HSA 1 contains numerous segmental duplications that display a high degree of sequence similarity between both chromosomal arms. Detailed analysis of these regions has allowed us to refine the p-arm breakpoint region to a 154.2 kb interval at 1p11.2 and the q-arm breakpoint region to a 562.6 kb interval at 1q21.1. Both breakpoint regions contain human-specific segmental duplications arranged in inverted orientation. We therefore propose that the pericentric inversion of HSA 1 was mediated by intra-chromosomal non-homologous recombination between these highly homologous segmental duplications that had themselves arisen only recently in the human lineage by duplicative transposition.





Similar content being viewed by others
References
Bailey JA, Yavor AM, Massa HF, Trask BJ, Eichler EE (2001) Segmental duplications: organization and impact within the current human genome project assembly. Genome Res 11:1005–1017
Bailey JA, Gu Z, Clark RA, Reinert K, Samonte RV, Schwartz S, Adams MD, Myers EW, Li PW, Eichler EE (2002) Recent segmental duplications in the human genome. Science 297:1003–1007
Barbi G, Spaich Ch, Adolph S, Rossier E, Kehrer-Sawatzki H (2005) Supernumerary der(1) marker chromosome derived from a ring chromosome 1 which has retained the original centromere and euchromatin from 1q21.1→q21.3 with substantial loss of 1q12 heterochromatin in a female with dysmorphic features and psychomotoric developmental delay. Am J Med Genet A 132:419–424
Buckland RA (1992) A primate transfer RNA gene cluster and the evolution of human chromosome 1. Cytogenet Cell Genet 61:1–4
Cheng Z, Ventura M, She X, Khaitovich P, Graves T, Osoegawa K, Church D, DeJong P, Wilson RK, Pääbo S, Rocchi M, Eichler EE (2005) Genome-wide comparison of recent chimpanzee and human segmental duplications. Nature 437:88–93
Cheung J, Estivill X, Khaja R, MacDonald JR, Lau K, Tsui LC, Scherer SW (2003) Genome-wide detection of segmental duplications and potential assembly errors in the human genome sequence. Genome Biol 4:R25
Choudhuri JV, Schleiermacher C, Kurtz S, Giegerich R (2004) GenAlyzer: interactive visualization of sequence similarities between entire genomes. Bioinformatics 20:1964–1965
Conrad DF, Andrews TD, Carter NP, Hurles ME, Pritchard JK (2006) A high-resolution survey of deletion polymorphism in the human genome. Nat Genet 38:75–81
Dennehey BK, Gutches DG, McConkey EH, Krauter KS (2004) Inversion, duplication, and changes in gene context are associated with human chromosome 18 evolution. Genomics 83:493–501
Dutrillaux B (1979) Chromosomal evolution in primates: tentative phylogeny from Microcebus murinus (Prosimian) to man. Hum Genet 48:251–314
Faas BH, Mieloo H, Van Es-Van Gaal JW, Van Ravenswaaij C (2003) A new case of dup(1)(q21.2q12) in an individual with mild mental retardation. Genet Couns 14:407–411
Fan Y, Linardopoulou E, Friedman C, Williams E, Trask BJ (2002) Genomic structure and evolution of the ancestral chromosome fusion site in 2q13–2q14.1 and paralogous regions on other human chromosomes. Genome Res 12:1651–1662
Fortna A, Kim Y, MacLaren E, Marshall K, Hahn G, Meltesen L, Brenton M, Hink R, Burgers S, Hernandez-Boussard T, Karimpour-Fard A, Glueck D, McGavran L, Berry R, Pollack J, Sikela JM (2004) Lineage-specific gene duplication and loss in human and great ape evolution. PLoS Biol 2:E207
Goidts V, Szamalek JM, Hameister H, Kehrer-Sawatzki H (2004) Segmental duplication associated with the human-specific inversion of chromosome 18: a further example of the impact of segmental duplications on karyotype and genome evolution in primates. Hum Genet 115:116–122
Goidts V, Szamalek JM, de Jong PJ, Cooper DN, Chuzhanova N, Hameister H, Kehrer-Sawatzki H (2005) Independent intrachromosomal recombination events underlie the pericentric inversions of chimpanzee and gorilla chromosomes homologous to human chromosome 16. Genome Res 15:1232–1242
Goidts V, Armengol L, Schempp W, Conroy J, Nowak N, Muller S, Cooper DN, Estivill X, Enard W, Szamalek JM, Hameister H, Kehrer-Sawatzki H (2006) Identification of large-scale human-specific copy number differences by inter-species array comparative genomic hybridization. Hum Genet 119:185–198
Gross M, Starke H, Trifonov V, Claussen U, Liehr T, Weise A (2006) A molecular cytogenetic study of chromosome evolution in chimpanzee. Cytogenet Genome Res 112:67–75
Haig D (2005) The complex history of distal human chromosome 1q. Genomics 86:767–770
Horvath JE, Schwartz S, Eichler EE (2000) The mosaic structure of human pericentromeric DNA: a strategy for characterizing complex regions of the human genome. Genome Res 10:839–852
Kehrer-Sawatzki H, Schreiner B, Tanzer S, Platzer M, Muller S, Hameister H (2002) Molecular characterization of the pericentric inversion that causes differences between chimpanzee chromosome 19 and human chromosome 17. Am J Hum Genet 71:375–388
Kehrer-Sawatzki H, Sandig C, Chuzhanova N, Goidts V, Szamalek JM, Tanzer S, Muller S, Platzer M, Cooper DN, Hameister H (2005a) Breakpoint analysis of the pericentric inversion distinguishing human chromosome 4 from the homologous chromosome in the chimpanzee (Pan troglodytes). Hum Mutat 25:45–55
Kehrer-Sawatzki H, Sandig CA, Goidts V, Hameister H (2005b) Breakpoint analysis of the pericentric inversion between chimpanzee chromosome 10 and the homologous chromosome 12 in humans. Cytogenet Genome Res 108:91–97
Kehrer-Sawatzki H, Szamalek JM, Tanzer S, Platzer M, Hameister H (2005c) Molecular characterization of the pericentric inversion of chimpanzee chromosome 11 homologous to human chromosome 9. Genomics 85:542–550
Linardopoulou EV, Williams EM, Fan Y, Friedman C, Young JM, Trask BJ (2005) Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 437:94–100
Locke DP, Archidiacono N, Misceo D, Cardone MF, Deschamps S, Roe B, Rocchi M, Eichler EE (2003) Refinement of a chimpanzee pericentric inversion breakpoint to a segmental duplication cluster. Genome Biol 4:R50
Martin CL, Wong A, Gross A, Chung J, Fantes JA, Ledbetter DH (2002) The evolutionary origin of human subtelomeric homologies—or where the ends begin. Am J Hum Genet 70:972–984
Maresco DL, Blue LE, Culley LL, Kimberly RP, Anderson CL, Theil KS (1998) Localization of FCGR1 encoding Fcgamma receptor class I in primates: molecular evidence for two pericentric inversions during the evolution of human chromosome 1. Cytogenet Cell Genet 82:71–74
Marzella R, Viggiano L, Miolla V, Storlazzi CT, Ricco A, Gentile E, Roberto R, Surace C, Fratello A, Mancini M, Archidiacono N, Rocchi M (2000) Molecular cytogenetic resources for chromosome 4 and comparative analysis of phylogenetic chromosome IV in great apes. Genomics 63:307–313
McCarroll SA, Hadnott TN, Perry GH, Sabeti PC, Zody MC, Barrett JC, Dallaire S, Gabriel SB, Lee C, Daly MJ, Altshuler DM, International HapMap Consortium (2005) Common deletion polymorphisms in the human genome. Nat Genet 38:86–92
McConkey EH (1997) The origin of human chromosome 18 from a human/ape ancestor. Cytogenet Cell Genet 76:189–191
Mikkelsen TS, Hillier LW, Eichler EE, Zody MC, Jaffe DB, Yang S-P, Enard W, Hellmann I, Lindblad-Toh K, Altheide TK, Archidiacono N, Bork P, Butler J, Chang JL, Cheng Z, Chinwalla AT, deJong P, Delehaunty KD, Fronick CC, Fulton LL, Gilad Y, Glusman G, Gnerre S, Graves TA, Hayakawa T, Hayden KE, Huang X, Ji H, Kent WJ, King M-C, Kulbokas EJ, Lee MK, Liu G, Lopez-Otin C, Makova KD, Man O, Mardis ER, Mauceli E, Miner TL, Nash WE, Nelson JO, Pääbo S, Patterson NJ, Poh CS, Pollard KS, Prüfer K, Puente XS, Reich D, Rocchi M, Rosenbloom K, Ruvolo M, Richter DJ, Schaffner SF, Smit AFA, Smith SM, Suyama M, Taylor J, Torrents D, Tuzun E, Varki A, Velasco G, Ventura M, Wallis JW, Wend MC, Wilson RK, Lander ES, Waterston RJ (2005) Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437:69–87
Mitelman F, Johansson B and Mertens F (2006) Mitelman Database of Chromosome Aberrations in Cancer. http://www.cgap.nci.nih.gov/Chromosomes/Mitelman
Monfouilloux S, Avet-Loiseau H, Amarger V, Balazs I, Pourcel C, Vergnaud G (1998) Recent human-specific spreading of a subtelomeric domain. Genomics 51:165–176
Müller S, Wienberg J (2001) “Bar-coding” primate chromosomes: molecular cytogenetic screening for the ancestral hominoid karyotype. Hum Genet 109:85–94
Murphy WJ, Fronicke L, O’Brien SJ, Stanyon R (2003) The origin of human chromosome 1 and its homologs in placental mammals. Genome Res 13:1880–1888
Newman TL, Tuzun E, Morrison VA, Hayden KE, Ventura M, McGrath SD, Rocchi M, Eichler EE (2005) A genome-wide survey of structural variation between human and chimpanzee. Genome Res 15:1344–1356
Nickerson E, Nelson DL (1998) Molecular definition of pericentric inversion breakpoints occurring during the evolution of humans and chimpanzees. Genomics 50:368–372
Parsons JD (1995) Miropeats: graphical DNA sequence comparisons. Comput Appl Biosci. 11:615–619
Ruiz-Herrera A, Garcia F, Azzalin C, Giulotto E, Egozcue J, Ponsa M, Garcia M (2002) Distribution of intrachromosomal telomeric sequences (ITS) on Macaca fascicularis (Primates) chromosomes and their implication for chromosome evolution. Hum Genet 110:578–586
Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, Maner S, Massa H, Walker M, Chi M, Navin N, Lucito R, Healy J, Hicks J, Ye K, Reiner A, Gilliam TC, Trask B, Patterson N, Zetterberg A, Wigler M (2004) Large-scale copy number polymorphism in the human genome. Science 305:525–528
She X, Jiang Z, Clark RA, Liu G, Cheng Z, Tuzun E, Church DM, Sutton G, Halpern AL, Eichler EE (2004) Shotgun sequence assembly and recent segmental duplications within the human genome. Nature 431:927–930
Shimada MK, Kim CG, Kitano T, Ferrell RE, Kohara Y, Saitou N (2005) Nucleotide sequence comparison of a chromosome rearrangement on human chromosome 12 and the corresponding ape chromosomes. Cytogenet Genome Res 108:83–90
Szamalek JM, Goidts V, Chuzhanova N, Hameister H, Cooper DN, Kehrer-Sawatzki H (2005) Molecular characterisation of the pericentric inversion that distinguishes human chromosome 5 from the homologous chimpanzee chromosome. Hum Genet 117:168–176
Szamalek JM, Goidts V, Searle JB, Cooper DN, Hameister H, Kehrer-Sawatzki H (2006a) The chimpanzee-specific pericentric inversions that distinguish humans and chimpanzees have identical breakpoints in Pan troglodytes and Pan paniscus. Genomics 87:39–45
Szamalek JM, Cooper DN, Schempp W, Minich P, Kohn M, Hoegel J, Goidts V, Hameister H, Kehrer-Sawatzki H (2006b) Polymorphic micro-inversions contribute to the genomic variability of humans and chimpanzees. Hum Genet 119:103–112
Trask BJ, Friedman C, Martin-Gallardo A, Rowen L, Akinbami C, Blankenship J, Collins C, Giorgi D, Iadonato S, Johnson F, Kuo WL, Massa H, Morrish T, Naylor S, Nguyen OT, Rouquier S, Smith T, Wong DJ, Youngblom J, van den Engh G (1998) Members of the olfactory receptor gene family are contained in large blocks of DNA duplicated polymorphically near the ends of human chromosomes. Hum Mol Genet 7:13-26
Tuzun E, Sharp AJ, Bailey JA, Kaul R, Morrison VA, Pertz LM, Haugen E, Hayden H, Albertson D, Pinkel D, Olson MV, Eichler EE (2005) Fine-scale structural variation of the human genome. Nat Genet 37:727–732
Weise A, Starke H, Mrasek K, Claussen U, Liehr T (2005) New insights into the evolution of chromosome 1. Cytogenet Genome Res 108:217–222
White PS, Matise TC (2003) Chromosome 1. In: Cooper DN (ed) Nature encyclopedia of the human genome, vol 1. Nature Publishing Group, London, pp 550–559
Wilson GM, Flibotte S, Missirlis PI, Marra MA, Jones S, Thornton K, Clark AG, Holt RA (2006) Identification by full-coverage array CGH of human DNA copy number increases relative to chimpanzee and gorilla. Genome Res 16:173–181
Won YJ, Hey J (2005) Divergence population genetics of chimpanzees. Mol Biol Evol 22:297–307
Yoder AD, Yang Z (2000) Estimation of primate speciation dates using local molecular clocks. Mol Biol Evol 17:1081–1090
Yunis JJ, Prakash O (1982) The origin of man: a chromosomal pictorial legacy. Science 215:1525–1530
Acknowledgment
This research was funded by the Deutsche Forschungsgemeinschaft (DFG KE 724/2-1).
Author information
Authors and Affiliations
Corresponding author
Additional information
Justyna M. Szamalek and Violaine Goidts are contributed equally to the paper.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Szamalek, J.M., Goidts, V., Cooper, D.N. et al. Characterization of the human lineage-specific pericentric inversion that distinguishes human chromosome 1 from the homologous chromosomes of the great apes. Hum Genet 120, 126–138 (2006). https://doi.org/10.1007/s00439-006-0209-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-006-0209-y