Abstract
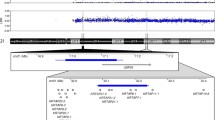
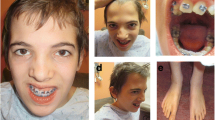
We describe the characterization of an interstitial duplication of 12p, dup(12)(p11.21p13.31), by array-CGH and FISH in a patient with mental retardation and dysmorphic features. The sequence analysis of the breakpoints revealed the presence of homologous low copy repeats (LCRs) flanking the duplication region, thus suggesting that they have mediated the rearrangement. Pip-maker analysis showed that a third cluster of homologous LCRs lie distally to the two mediating the 12p duplication. We hypothesize that this duplication might be a new recurrent rearrangement and that, thanks to the different orientations of the homologous regions lying within each cluster, the three clusters are responsible for at least some of the several 12p aneuploidies reported in the literature such as direct and inverted duplications, deletions and supernumerary analphoid chromosomes. Moreover, we excluded that polymorphic inversions between these three clusters are present in the normal population.




Similar content being viewed by others
References
Allen TL, Brothman AR, Carey JC, Chance PF (1996) Cytogenetic and molecular analysis in trisomy 12p. Am J Med Genet 63:250–256
Amann J, Valentine M, Kidd VJ, Lahti JM (1996) Localization of chi1-related helicase genes to human chromosome regions 12p11 and 12p13: similarity between parts of these genes and conserved human telomeric-associated DNA. Genomics 32:260–265
Amor DJ, Choo KH (2002) Neocentromeres: role in human disease, evolution, and centromere study. Am J Hum Genet 71:695–714
Barbouti A, Stankiewicz P, Nusbaum C, Cuomo C, Cook A, Hoglund M, Johansson B, Hagemeijer A, Park SS, Mitelman F, Lupski JR, Fioretos T (2004) The breakpoint region of the most common isochromosome, i(17q), in human neoplasia is characterized by a complex genomic architecture with large, palindromic, low-copy repeats. Am J Hum Genet 74:1–10
Dufke A, Walczak C, Liehr T, Starke H, Trifonov V, Rubtsov N, Schoning M, Enders H, Eggermann T (2001) Partial tetrasomy 12pter-12p12.3 in a girl with Pallister–Killian syndrome: extraordinary finding of an analphoid, inverted duplicated marker. Eur J Hum Genet 9:572–576
Eichler EE, Sankoff D (2003) Structural dynamics of eukaryotic chromosome evolution. Science 301:793–797
Giglio S, Broman KW, Matsumoto N, Calvari V, Gimelli G, Neumann T, Ohashi H, Voullaire L, Larizza D, Giorda R, Weber JL, Ledbetter DH, Zuffardi O (2001) Olfactory receptor-gene clusters, genomic-inversion polymorphisms, and common chromosome rearrangements. Am J Hum Genet 68:874–883
Giglio S, Calvari V, Gregato G, Gimelli G, Camanini S, Giorda R, Ragusa A, Guerneri S, Selicorni A, Stumm M, Tonnies H, Ventura M, Zollino M, Neri G, Barber J, Wieczorek D, Rocchi M, Zuffardi O (2002) Heterozygous submicroscopic inversions involving olfactory receptor-gene clusters mediate the recurrent t(4;8)(p16;p23) translocation. Am J Hum Genet 71:276–285
Horvath JE, Viggiano L, Loftus BJ, Adams MD, Archidiacono N, Rocchi M, Eichler EE (2000) Molecular structure and evolution of an alpha satellite/non-alpha satellite junction at 16p11. Hum Mol Genet 9:113–123
Jacobs PA, Browne C, Gregson N, Joyce C, White H (1992) Estimates of the frequency of chromosome abnormalities detectable in unselected newborns using moderate levels of banding. J Med Genet 29:103–108
Oldenburg J, Rost S, El-Maarri O, Leuer M, Olek K, Muller CR, Schwaab R (2000) De novo factor VIII gene intron 22 inversion in a female carrier presents as a somatic mosaicism. Blood 96:2905–2906
Osborne LR, Li M, Pober B, Chitayat D, Bodurtha J, Mandel A, Costa T, Grebe T, Cox S, Tsui LC, Scherer SW (2001) A 1.5 million-base pair inversion polymorphism in families with Williams-Beuren syndrome. Nat Genet 29:321–325
Perez Jurado AL (2003) Williams–Beuren syndrome: a model of recurrent genomic mutation. Horm Res 59:106–113
Pramparo T, Giglio S, Gregato G, de Gregori M, Patricelli MG, Ciccone R, Scappaticci S, Mannino G, Lombardi C, Pirola B, Giorda R, Rocchi M, Zuffardi O (2004) Inverted duplications: how many of them are mosaic? Eur J Hum Genet 12:713–717
Rauch A, Trautmann U, Pfeiffer RA (1996) Clinical and molecular cytogenetic observations in three cases of “trisomy 12p syndrome”. Am J Med Genet 63:243–249
Saglio G, Storlazzi CT, Giugliano E, Surace C, Anelli L, Rege-Cambrin G, Zagaria A, Jimenez Velasco A, Heiniger A, Scaravaglio P, Torres Gomez A, Roman Gomez J, Archidiacono N, Banfi S, Rocchi M (2002) A 76-kb duplicon maps close to the BCR gene on chromosome 22 and the ABL gene on chromosome 9: possible involvement in the genesis of the Philadelphia chromosome translocation. Proc Natl Acad Sci USA 99:9882–9887
Samonte RV, Eichler EE (2002) Segmental duplications and the evolution of the primate genome. Nat Rev Genet 3:65–72
Schinzel A (2001) Catalogue of unbalanced chromosome aberrations in man. In: Albert Schinzel (ed) 2nd, edn. Berlin
Schwartz S, Zhang Z, Frazer KA, Smit A, Riemer C, Bouck J, Gibbs R, Hardison R, Miller W (2000) PipMaker—a web server for aligning two genomic DNA sequences. Genome Res 1:577–586
Shaw CJ, Lupski JR (2004) Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet 13:57–64
Shaw CJ, Lupski JR (2005) Non-recurrent 17p11.2 deletions are generated by homologous and non-homologous mechanisms. Hum Genet 116:1–7
Shaw-Smith C, Redon R, Rickman L, Rio M, Willatt L, Fiegler H, Firth H, Sanlaville D, Winter R, Colleaux L, Bobrow M, Carter NP (2004) Microarray based comparative genomic hybridisation (array-CGH) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. Med Genet 41:241–248
Stankiewicz P, Lupski JR (2002) Genome architecture, rearrangements and genomic disorders. Trends Genet 18:74–82
Stankiewicz P, Shaw CJ, Dapper JD, Wakui K, Shaffer LG, Withers M, Elizondo L, Park SS, Lupski JR (2003) Genome architecture catalyzes nonrecurrent chromosomal rearrangements. Am J Hum Genet 72:1101–1116
Stefansson H, Helgason A, Thorleifsson G, Steinthorsdottir V, Masson G, Barnard J, Baker A, Jonasdottir A, Ingason A, Gudnadottir VG, Desnica N, Hicks A, Gylfason A, Gudbjartsson DF, Jonsdottir GM, Sainz J, Agnarsson K, Birgisdottir B, Ghosh S, Olafsdottir A, Cazier JB, Kristjansson K, Frigge ML, Thorgeirsson TE, Gulcher JR, Kong A, Stefansson K (2005) A common inversion under selection in Europeans. Nat Genet 37:129–137
Stengel-Rutkowski S, Albert A, Murken JD, Zahn-Messow K, Rodewald A, Zankl M, Saule H, Stene J (1981) New chromosomal dysmorphic syndromes. 4. Trisomy 12p. Eur J Pediatr 136:249–262
Uchida IA, Lin CC (1973) Identification of partial 12 trisomy by quinacrine fluorescence. J Pediatr 82:269–272
Venturin M, Gervasini C, Orzan F, Bentivegna A, Corrado L, Colapietro P, Friso A, Tenconi R, Upadhyaya M, Larizza L, Riva P (2004) Evidence for non-homologous end joining and non-allelic homologous recombination in atypical NF1 microdeletions. Hum Genet 115:69–80
Vissers LE, de Vries BB, Osoegawa K, Janssen IM, Feuth T, Choy CO, Straatman H, van der Vliet W, Huys EH, van Rijk A, Smeets D, van Ravenswaaij-Arts CM, Knoers NV, van der Burgt I, de Jong PJ, Brunner HG, van Kessel AG, Schoenmakers EF, Veltman JA (2003) Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 73:1261–1270
Warburton PE (2004) Chromosomal dynamics of human neocentromere formation. Chromosome Res 12:617–626
Zumkeller W, Volleth M, Muschke P, Tonnies H, Heller A, Liehr T, Wieacker P, Stumm M (2004) Genotype/phenotype analysis in a patient with pure and complete trisomy 12p. Am J Med Genet 129A:261–264
Acknowledgements
We are grateful to the YAC Screening Centre of San Raffaele Biomedical Science Park (Milan) for providing BAC clones. This work was supported by cofin03-MIUR (to OZ), cofin04-MIUR, the FIRB 2001(to OZ), the Italian Telethon Foundation (GP0247Y01 to OZ) and the Cariplo Foundation (to OZ).
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuela De Gregori, Tiziano Pramparo contributed equally to this paper.
Rights and permissions
About this article
Cite this article
De Gregori, M., Pramparo, T., Memo, L. et al. Direct duplication 12p11.21–p13.31 mediated by segmental duplications: a new recurrent rearrangement?. Hum Genet 118, 207–213 (2005). https://doi.org/10.1007/s00439-005-0008-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-005-0008-x