Abstract
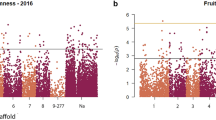
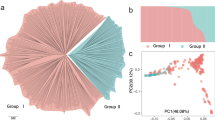
The Japanese apricot (Prunus mume) is a popular fruit tree in Japan. However, the genetic factors associated with fruit trait variations are poorly understood. In this study, we investigated nine fruit-associated traits, including harvesting time, fruit diameter, fruit shape, fruit weight, stone (endocarp) weight, ratio of stone weight to fruit weight, and rate of fruit gumming, using 110 Japanese apricot accessions over four years. A genome-wide association study (GWAS) was performed for these traits and strong signals were detected on chromosome 6 for harvesting time and fruit diameters. These peaks were shown to undergo strong artificial selection during the differentiation of small-fruit cultivars. The genomic region defined by the GWAS and XP-nSL analyses harbored several candidate genes associated with plant hormone regulation. Furthermore, the alleles of small-fruit cultivars in this region were shown to have genetic proximity to some Chinese cultivars of P. mume. These results indicate that the small-fruit trait originated in China; after being introduced into Japan, it was preferred and selected by the Japanese people, resulting in the differentiation of small-fruit cultivars.




Similar content being viewed by others
Data availability
Raw FASTQ reads for Prunus mume accessions sequenced in this study were deposited in the sequence read archive (SRA) under DRA accession number DRA016343.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664. https://doi.org/10.1101/gr.094052.109
Auwera GA, O’Connor BD (2020) Genomics in the cloud : using docker, GATK, and WDL in terra. O’Reilly Media
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bortiri E, Oh S-H, Jiang J et al (2001) Phylogeny and systematics of Prunus (Rosaceae) as determined by sequence analysis of ITS and the chloroplast trnL-trnF spacer DNA. Syst Bot 26:797–807
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Browning BL, Zhou Y, Browning SR (2018) A one-penny imputed genome from next-generation reference panels. Am J Hum Genet 103:338–348. https://doi.org/10.1016/j.ajhg.2018.07.015
Bu H, Yu W, Yuan H et al (2020) Endogenous auxin content contributes to larger size of apple fruit. Front Plant Sci 11:592540. https://doi.org/10.3389/fpls.2020.592540
Calle A, Wünsch A (2020) Multiple-population QTL mapping of maturity and fruit-quality traits reveals LG4 region as a breeding target in sweet cherry (Prunus avium L.). Hortic Res 7:1–13. https://doi.org/10.1038/s41438-020-00349-2
Chen J (2017) China mei flower (Prunus mume) cultivars in colour. China Forestry Publishing House, Beijing
Cingolani P, Platts A, Wang LL et al (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly (austin) 6:80–92. https://doi.org/10.4161/fly.19695
Cirilli M, Bassi D, Ciacciulli A (2016) Sugars in peach fruit: a breeding perspective. Hortic Res 3:15067. https://doi.org/10.1038/hortres.2015.67
Cirilli M, Baccichet I, Chiozzotto R et al (2021) Genetic and phenotypic analyses reveal major quantitative loci associated to fruit size and shape traits in a non-flat peach collection (P. persica L. Batsch). Hortic Res 8:1–17. https://doi.org/10.1038/s41438-021-00661-5
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Dare AP, Yauk Y-K, Tomes S et al (2017) Silencing a phloretin-specific glycosyltransferase perturbs both general phenylpropanoid biosynthesis and plant development. Plant J 91:237–250. https://doi.org/10.1111/tpj.13559
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321. https://doi.org/10.1016/j.cell.2006.12.006
Duan N, Bai Y, Sun H et al (2017) Genome re-sequencing reveals the history of apple and supports a two-stage model for fruit enlargement. Nat Commun 8:249. https://doi.org/10.1038/s41467-017-00336-7
Forlani S, Mizzotti C, Masiero S (2021) The NAC side of the fruit: tuning of fruit development and maturation. BMC Plant Biol 21:238. https://doi.org/10.1186/s12870-021-03029-y
Gel B, Serra E (2017) karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33:3088–3090. https://doi.org/10.1093/bioinformatics/btx346
Horiuchi S, Yoshida M, Kariya H et al (1996) Nihonnoume Sekainoume. Yokendo, Tokyo (Japanese)
Huang X, Ni Z, Shi T et al (2022) Novel insights into the dissemination route of Japanese apricot (Prunus mume Sieb. et Zucc.) based on genomics. Plant J 110:1182–1197. https://doi.org/10.1111/tpj.15731
Kalyaanamoorthy S, Minh BQ, Wong TKF et al (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Kitamura Y, Habu T, Yamane H et al (2018) Identification of QTLs controlling chilling and heat requirements for dormancy release and bud break in Japanese apricot (Prunus mume). Tree Genet Genomes 14:33. https://doi.org/10.1007/s11295-018-1243-3
Korte A, Farlow A (2013) The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9:29. https://doi.org/10.1186/1746-4811-9-29
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li Y-L, Weng J-C, Hsiao C-C et al (2015) PEAT: an intelligent and efficient paired-end sequencing adapter trimming algorithm. BMC Bioinform 16:S2. https://doi.org/10.1186/1471-2105-16-S1-S2
Li M, Sang M, Wen Z et al (2022) Mapping floral genetic architecture in Prunus mume, and ornamental woody plant. Front Plant Sci 13:828579. https://doi.org/10.3389/fpls.2022.828579
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:13033997 [q-bio]
Mega K, Tomita E, Kitamura S et al (1988) The grand dictionary of horticulture. Shogakukan, Tokyo (Japanese)
Migicovsky Z, Gardner KM, Money D et al (2016) Genome to phenome mapping in apple using historical data. Plant Genome. https://doi.org/10.3835/plantgenome2015.11.0113
Minamikawa MF, Nonaka K, Kaminuma E et al (2017) Genome-wide association study and genomic prediction in citrus: potential of genomics-assisted breeding for fruit quality traits. Sci Rep 7:4721. https://doi.org/10.1038/s41598-017-05100-x
Minamikawa MF, Takada N, Terakami S et al (2018) Genome-wide association study and genomic prediction using parental and breeding populations of Japanese pear (Pyrus pyrifolia Nakai). Sci Rep 8:11994. https://doi.org/10.1038/s41598-018-30154-w
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195. https://doi.org/10.1093/molbev/mst024
Minh BQ, Schmidt HA, Chernomor O et al (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534. https://doi.org/10.1093/molbev/msaa015
Ministry of Agriculture, Forestry and Fisheries Japan (2023) THE 96th STATISTICAL YEARBOOK OF MINISTRY OF AGRICULTURE, FORESTRY AND FISHERIES : MAFF. https://www.maff.go.jp/e/data/stat/96th/index.html. Accessed 5 May 2023
Numaguchi K, Ishio S, Kitamura Y et al (2019) Microsatellite marker development and population structure analysis in Japanese apricot (Prunus mume Sieb. et Zucc.). Hort J 88:222–231. https://doi.org/10.2503/hortj.UTD-013
Numaguchi K, Akagi T, Kitamura Y et al (2020) Interspecific introgression and natural selection in the evolution of Japanese apricot (Prunus mume). Plant J 104:1551–1567. https://doi.org/10.1111/tpj.15020
Numaguchi K, Kashiwamoto T, Ishikawa R et al (2023) Genome-wide association study detects loci involved in scab susceptibility in Japanese apricot. Horticulturae 9:872. https://doi.org/10.3390/horticulturae9080872
Nuñez-Lillo G, Cifuentes-Esquivel A, Troggio M et al (2015) Identification of candidate genes associated with mealiness and maturity date in peach [Prunus persica (L.) Batsch] using QTL analysis and deep sequencing. Tree Genet Genomes 11:86. https://doi.org/10.1007/s11295-015-0911-9
Ono E, Homma Y, Horikawa M et al (2010) Functional differentiation of the glycosyltransferases that contribute to the chemical diversity of bioactive flavonol glycosides in grapevines (Vitis vinifera). Plant Cell 22:2856–2871. https://doi.org/10.1105/tpc.110.074625
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Rao MJ, Zuo H, Xu Q (2021) Genomic insights into citrus domestication and its important agronomic traits. Plant Commun 2:100138. https://doi.org/10.1016/j.xplc.2020.100138
Shi T, Luo W, Li H et al (2019) Association between blooming time and climatic adaptation in Prunus mume. Ecol Evol 10:292–306. https://doi.org/10.1002/ece3.5894
Shimizu T (2022) Breeding new premium quality cultivars by citrus breeding 2.0 in Japan: an integrative approach suggested by genealogy. Horticulturae 8:559. https://doi.org/10.3390/horticulturae8060559
Soundappan I, Bennett T, Morffy N et al (2015) SMAX1-LIKE/D53 family members enable distinct MAX2-dependent responses to strigolactones and karrikins in Arabidopsis. Plant Cell 27:3143–3159. https://doi.org/10.1105/tpc.15.00562
Szpiech ZA, Hernandez RD (2014) selscan: an efficient multithreaded program to perform EHH-based scans for positive selection. Mol Biol Evol 31:2824–2827. https://doi.org/10.1093/molbev/msu211
Szpiech ZA, Novak TE, Bailey NP, Stevison LS (2021) Application of a novel haplotype-based scan for local adaptation to study high-altitude adaptation in rhesus macaques. Evol Lett 5:408–421. https://doi.org/10.1002/evl3.232
Verde I, Jenkins J, Dondini L et al (2017) The Peach v2.0 release: high-resolution linkage mapping and deep resequencing improve chromosome-scale assembly and contiguity. BMC Genom 18:225. https://doi.org/10.1186/s12864-017-3606-9
Xie Z, Nolan TM, Jiang H, Yin Y (2019) AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci 10:228. https://doi.org/10.3389/fpls.2019.00228
Yaegaki H, Haji T, Nakamura Y et al (2003) Varietal and yearly variations in fruit and endocarp weights and their ratio in Japanese apricot (Prunus mume Sieb. et Zucc.) cultivars. J Japan Soc Hort Sci 72:546–550. https://doi.org/10.2503/jjshs.72.546(Japanese)
Yamaguchi M, Haji T, Yaegaki H (2004) Differences in mesocarp cell number, cell length and occurrence of gumming in fruit of Japanese apricot (Prunus mume Sieb. et Zucc.) cultivars during their development. J Japan Soc Hort Sci 73:200–207. https://doi.org/10.2503/jjshs.73.200
Yoshida M, Yamanishi H (1988) Apricot cultivars in Japan. Acta Hort 209:69–81
Zhang Q, Zhang H, Sun L et al (2018) The genetic architecture of floral traits in the woody plant Prunus mume. Nat Commun 9:1702. https://doi.org/10.1038/s41467-018-04093-z
Zhuo X, Zheng T, Li S et al (2021) Identification of the PmWEEP locus controlling weeping traits in Prunus mume through and integrated genome-wide association study and quantitative trait locus mapping. Hortic Res 8:131. https://doi.org/10.1038/s41438-021-00573-4
Acknowledgements
We are deeply grateful to the researchers and technicians at the Japanese Apricot Laboratory, Wakayama Fruit Tree Experiment Station, for maintaining the Prunus mume accessions. We would like to thank Drs. R. Ishikawa and T. Ishii, Graduate School of Agricultural Science, Kobe University, for investigation support and valuable discussions. This work was supported by JSPS KAKENHI Grant Number JP18K14449 to KN and Wakayama Prefecture.
Author information
Authors and Affiliations
Contributions
KN and YK conceived and designed the study. KN, YK, TK, TM, and TO performed the experiments. KN, YK, and TM analyzed the data. KN, YK, and TM contributed to the bioinformatics analysis. KN and YK prepared the manuscript. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Bing Yang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Numaguchi, K., Kitamura, Y., Kashiwamoto, T. et al. Genomic region and origin for selected traits during differentiation of small-fruit cultivars in Japanese apricot (Prunus mume). Mol Genet Genomics 298, 1365–1375 (2023). https://doi.org/10.1007/s00438-023-02062-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-023-02062-w