Abstract
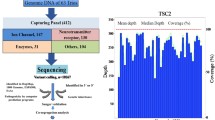
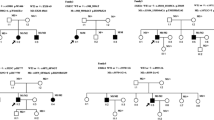
Hereditary neurological disorders (HNDs) are a clinically and genetically heterogeneous group of disorders. These disorders arise from the impaired function of the central or peripheral nervous system due to aberrant electrical impulses. More than 600 various neurological disorders, exhibiting a wide spectrum of overlapping clinical presentations depending on the organ(s) involved, have been documented. Owing to this clinical heterogeneity, diagnosing these disorders has been a challenge for both clinicians and geneticists and a large number of patients are either misdiagnosed or remain entirely undiagnosed. Contribution of genetics to neurological disorders has been recognized since long; however, the complete picture of the underlying molecular bases are under-explored. The aim of this study was to accurately diagnose 11 unrelated Pakistani families with various HNDs deploying NGS as a first step approach. Using exome sequencing and gene panel sequencing, we successfully identified disease-causing genomic variants these families. We report four novel variants, one each in, ECEL1, NALCN, TBR1 and PIGP in four of the pedigrees. In the rest of the seven families, we found five previously reported pathogenic variants in POGZ, FA2H, PLA2G6 and CYP27A1. Of these, three families segregate a homozygous 18 bp in-frame deletion of FA2H, indicating a likely founder mutation segregating in Pakistani population. Genotyping for this mutation can help low-cost population wide screening in the corresponding regions of the country. Our findings not only expand the existing repertoire of mutational spectrum underlying neurological disorders but will also help in genetic testing of individuals with HNDs in other populations.

Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abbas S, Brugger B, Zubair M, Gul S, Blatterer J, Wenninger J, Rehman K, Tatrai B, Khan MA, Windpassinger C (2021) Exome sequencing of a Pakistani family with spastic paraplegia identified an 18 bp deletion in the cytochrome B5 domain of FA2H. Neurol Res 43(2):133–140. https://doi.org/10.1080/01616412.2020.1831329
Al-Dewik N, Mohd H, Al-Mureikhi M, Ali R, Al-Mesaifri F, Mahmoud L, Shahbeck N, El-Akouri K, Almulla M, Al Sulaiman R, Musa S, Al-Marri AA, Richard G, Juusola J, Solomon BD, Alkuraya FS, Ben-Omran T (2019) Clinical exome sequencing in 509 Middle Eastern families with suspected Mendelian diseases: the Qatari experience. Am J Med Genet A 179(6):927–935. https://doi.org/10.1002/ajmg.a.61126
Angius A, Cossu S, Uva P, Oppo M, Onano S, Persico I, Fotia G, Atzeni R, Cuccuru G, Asunis M, Cucca F, Pruna D, Crisponi L (2018) Novel NALCN biallelic truncating mutations in siblings with IHPRF1 syndrome. Clin Genet 93(6):1245–1247. https://doi.org/10.1111/cge.13162
Baig SM, Sabih D, Rahim MK, Azhar A, Tariq M, Sajid Hussain M, Saqlan Naqvi SM, Raja GK, Khan TN, Jameel M, Iram Z, Noor S, Baig UR, Qureshi JA, Baig SA, Bakhtiar SM (2012) Beta-Thalassemia in Pakistan: a pilot program on prenatal diagnosis in Multan. J Pediatr Hematol Oncol 34(2):90–92. https://doi.org/10.1097/MPH.0b013e31823752f3
Baig SM, Fatima A, Tariq M, Khan TN, Ali Z, Faheem M, Mahmood H, Killela P, Waitkus M, He Y, Zhao F, Wang S, Jiao Y, Yan H (2019) Hereditary brain tumor with a homozygous germline mutation in PMS2: pedigree analysis and prenatal screening in a family with constitutional mismatch repair deficiency (CMMRD) syndrome. Fam Cancer 18(2):261–265. https://doi.org/10.1007/s10689-018-0112-4
Bohlega SA, Al-Mubarak BR, Alyemni EA, Abouelhoda M, Monies D, Mustafa AE, Khalil DS, Al Haibi S, Abou Al-Shaar H, Faquih T, El-Kalioby M, Tahir AI, Al Tassan NA (2016) Clinical heterogeneity of PLA2G6-related Parkinsonism: analysis of two Saudi families. BMC Res Notes 9:295. https://doi.org/10.1186/s13104-016-2102-7
Bower MA, Bushara K, Dempsey MA, Das S, Tuite PJ (2011) Novel mutations in siblings with later-onset PLA2G6-associated neurodegeneration (PLAN). Mov Disord 26(9):1768–1769. https://doi.org/10.1002/mds.23617
Cali JJ, Hsieh CL, Francke U, Russell DW (1991) Mutations in the bile acid biosynthetic enzyme sterol 27-hydroxylase underlie cerebrotendinous xanthomatosis. J Biol Chem 266(12):7779–7783. https://www.ncbi.nlm.nih.gov/pubmed/2019602
Chen YJ, Chen YC, Dong HL, Li LX, Ni W, Li HF, Wu ZY (2018) Novel PLA2G6 mutations and clinical heterogeneity in Chinese cases with phospholipase A2-associated neurodegeneration. Parkinsonism Relat Disord 49:88–94. https://doi.org/10.1016/j.parkreldis.2018.02.010
Choi DK, Suzuki Y, Yoshimura S, Togashi T, Hida M, Taylor TD, Wang Y, Sugano S, Hattori M, Sakaki Y (2001) Molecular cloning and characterization of a gene expressed in mouse developing tongue, mDscr5 gene, a homolog of human DSCR5 (Down syndrome Critical Region gene 5). Mamm Genome 12(5):347–351. https://doi.org/10.1007/s003350010283
Cochet-Bissuel M, Lory P, Monteil A (2014) The sodium leak channel, NALCN, in health and disease. Front Cell Neurosci 8:132. https://doi.org/10.3389/fncel.2014.00132
Daida K, Nishioka K, Li Y, Yoshino H, Shimada T, Dougu N, Nakatsuji Y, Ohara S, Hashimoto, T, Okiyama, R, Yokochi F, Suzuki C, Tomiyama M, Kimura K, Ueda N, Tanaka F, Yamada H, Fujioka S, Tsuboi Y, Hattori N (2021) PLA2G6 variants associated with the number of affected alleles in Parkinson's disease in Japan. Neurobiol Aging, 97, 147 e141–147 e149. https://doi.org/10.1016/j.neurobiolaging.2020.07.004
Deriziotis P, O’Roak BJ, Graham SA, Estruch SB, Dimitropoulou D, Bernier RA, Gerdts J, Shendure J, Eichler EE, Fisher SE (2014) De novo TBR1 mutations in sporadic autism disrupt protein functions. Nat Commun 5:4954. https://doi.org/10.1038/ncomms5954
Desvignes JP, Bartoli M, Delague V, Krahn M, Miltgen M, Beroud C, Salgado D (2018) VarAFT: a variant annotation and filtration system for human next generation sequencing data. Nucleic Acids Res 46(W1):W545–W553. https://doi.org/10.1093/nar/gky471
Dick KJ, Al-Mjeni R, Baskir W, Koul R, Simpson MA, Patton MA, Raeburn S, Crosby AH (2008) A novel locus for an autosomal recessive hereditary spastic paraplegia (SPG35) maps to 16q21-q23. Neurology 71(4):248–252. https://doi.org/10.1212/01.wnl.0000319610.29522.8a
Dick KJ, Eckhardt M, Paisan-Ruiz C, Alshehhi AA, Proukakis C, Sibtain NA, Maier H, Sharifi R, Patton MA, Bashir W, Koul R, Raeburn S, Gieselmann V, Houlden H, Crosby AH (2010) Mutation of FA2H underlies a complicated form of hereditary spastic paraplegia (SPG35). Hum Mutat 31(4):E1251-1260. https://doi.org/10.1002/humu.21205
Dieterich K, Quijano-Roy S, Monnier N, Zhou J, Faure J, Smirnow DA, Carlier R, Laroche C, Marcorelles P, Mercier S, Megarbane A, Odent S, Romero N, Sternberg D, Marty I, Estournet B, Jouk PS, Melki J, Lunardi J (2013) The neuronal endopeptidase ECEL1 is associated with a distinct form of recessive distal arthrogryposis. Hum Mol Genet 22(8):1483–1492. https://doi.org/10.1093/hmg/dds514
Edvardson S, Hama H, Shaag A, Gomori JM, Berger I, Soffer D, Korman SH, Taustein I, Saada A, Elpeleg O (2008) Mutations in the fatty acid 2-hydroxylase gene are associated with leukodystrophy with spastic paraparesis and dystonia. Am J Hum Genet 83(5):643–648. https://doi.org/10.1016/j.ajhg.2008.10.010
Federico A, Dotti MT, Lore F, Nuti R (1993) Cerebrotendinous xanthomatosis: pathophysiological study on bone metabolism. J Neurol Sci 115(1):67–70. https://doi.org/10.1016/0022-510x(93)90068-a
Ferretti A, Barresi S, Trivisano M, Ciolfi A, Dentici ML, Radio FC, Vigevano F, Tartaglia M, Specchio N (2019) POGZ-related epilepsy: Case report and review of the literature. Am J Med Genet A 179(8):1631–1636. https://doi.org/10.1002/ajmg.a.61206
Gilissen C, Hehir-Kwa JY, Thung DT, van de Vorst M, van Bon BW, Willemsen MH, Kwint M, Janssen IM, Hoischen A, Schenck A, Leach R, Klein R, Tearle R, Bo T, Pfundt R, Yntema HG, de Vries BB, Kleefstra T, Brunner HG, Veltman JA (2014) Genome sequencing identifies major causes of severe intellectual disability. Nature 511(7509):344–347. https://doi.org/10.1038/nature13394
Gregory A, Polster BJ, Hayflick SJ (2009) Clinical and genetic delineation of neurodegeneration with brain iron accumulation. J Med Genet 46(2):73–80. https://doi.org/10.1136/jmg.2008.061929
Grimberg J, Nawoschik S, Belluscio L, McKee R, Turck A, Eisenberg A (1989) A simple and efficient non-organic procedure for the isolation of genomic DNA from blood. Nucleic Acids Res 17(20):8390. https://doi.org/10.1093/nar/17.20.8390
Grunseich C, Sarkar N, Lu J, Owen M, Schindler A, Calabresi PA, Sumner CJ, Roda RH, Chaudhry V, Lloyd TE, Crawford TO, Subramony SH, Oh SJ, Richardson P, Tanji K, Kwan JY, Fischbeck KH, Mankodi A (2021) Improving the efficacy of exome sequencing at a quaternary care referral centre: novel mutations, clinical presentations and diagnostic challenges in rare neurogenetic diseases. J Neurol Neurosurg Psychiatry 92(11):1186–1196. https://doi.org/10.1136/jnnp-2020-325437
Hayflick SJ, Kurian MA, Hogarth P (2018) Neurodegeneration with brain iron accumulation. Handb Clin Neurol 147:293–305. https://doi.org/10.1016/B978-0-444-63233-3.00019-1
Hevner RF, Shi L, Justice N, Hsueh Y, Sheng M, Smiga S, Bulfone A, Goffinet AM, Campagnoni AT, Rubenstein JL (2001) Tbr1 regulates differentiation of the preplate and layer 6. Neuron 29(2):353–366. https://doi.org/10.1016/s0896-6273(01)00211-2
Huang TN, Chuang HC, Chou WH, Chen CY, Wang HF, Chou SJ, Hsueh YP (2014) Tbr1 haploinsufficiency impairs amygdalar axonal projections and results in cognitive abnormality. Nat Neurosci 17(2):240–247. https://doi.org/10.1038/nn.3626
Johnstone DL, Nguyen TT, Murakami Y, Kernohan KD, Tetreault M, Goldsmith C, Doja A, Wagner JD, Huang L, Hartley T, St-Denis A, Le Deist F, Majewski J, Bulman DE, Care4Rare Canada C, Kinoshita T, Dyment DA, Boycott KM, Campeau PM (2017) Compound heterozygous mutations in the gene PIGP are associated with early infantile epileptic encephalopathy. Hum Mol Genet 26(9):1706–1715. https://doi.org/10.1093/hmg/ddx077
Kim YJ, Lyoo CH, Hong S, Kim NY, Lee MS (2015) Neuroimaging studies and whole exome sequencing of PLA2G6-associated neurodegeneration in a family with intrafamilial phenotypic heterogeneity. Parkinsonism Relat Disord 21(4):402–406. https://doi.org/10.1016/j.parkreldis.2015.01.010
Kim MJ, Yum MS, Seo GH, Lee Y, Jang HN, Ko TS, Lee BH (2020) Clinical application of whole exome sequencing to identify rare but remediable neurologic disorders. J Clin Med 9(11). https://doi.org/10.3390/jcm9113724
Koh K, Ichinose Y, Ishiura H, Nan H, Mitsui J, Takahashi J, Sato W, Itoh Y, Hoshino K, Tsuji S, Takiyama Y, Japan Spastic Paraplegia Research C (2019a) Correction: PLA2G6-associated neurodegeneration presenting as a complicated form of hereditary spastic paraplegia. J Hum Genet 64(1):61–63. https://doi.org/10.1038/s10038-018-0533-9
Koh K, Ichinose Y, Ishiura H, Nan H, Mitsui J, Takahashi J, Sato W, Itoh Y, Hoshino K, Tsuji S, Takiyama Y, Japan Spastic Paraplegia Research C (2019b) PLA2G6-associated neurodegeneration presenting as a complicated form of hereditary spastic paraplegia. J Hum Genet 64(1):55–59. https://doi.org/10.1038/s10038-018-0519-7
Krenn M, Knaus A, Westphal DS, Wortmann SB, Polster T, Woermann FG, Karenfort M, Mayatepek E, Meitinger T, Wagner M, Distelmaier F (2019) Biallelic mutations in PIGP cause developmental and epileptic encephalopathy. Ann Clin Transl Neurol 6(5):968–973. https://doi.org/10.1002/acn3.768
Mari F, Berti B, Romano A, Baldacci J, Rizzi R, Grazia Alessandri M, Tessa A, Procopio E, Rubegni A, Lourenco CM, Simonati A, Guerrini R, Santorelli FM (2018) Clinical and neuroimaging features of autosomal recessive spastic paraplegia 35 (SPG35): case reports, new mutations, and brief literature review. Neurogenetics 19(2):123–130. https://doi.org/10.1007/s10048-018-0538-8
Matilla-Duenas A, Corral-Juan M, Rodriguez-Palmero Seuma A, Vilas D, Ispierto L, Morais S, Sequeiros J, Alonso I, Volpini V, Serrano-Munuera C, Pintos-Morell G, Alvarez R, Sanchez I (2017) Rare neurodegenerative diseases: clinical and genetic update. Adv Exp Med Biol 1031:443–496. https://doi.org/10.1007/978-3-319-67144-4_25
McDermott JH, Study DDD, Clayton-Smith J, Briggs TA (2018) The TBR1-related autistic-spectrum-disorder phenotype and its clinical spectrum. Eur J Med Genet 61(5):253–256. https://doi.org/10.1016/j.ejmg.2017.12.009
McMillin MJ, Below JE, Shively KM, Beck AE, Gildersleeve HI, Pinner J, Gogola GR, Hecht JT, Grange DK, Harris DJ, Earl DL, Jagadeesh S, Mehta SG, Robertson SP, Swanson JM, Faustman EM, Mefford HC, Shendure J, Nickerson DA, University of Washington Center for Mendelian G (2013) Mutations in ECEL1 cause distal arthrogryposis type 5D. Am J Hum Genet 92(1):150–156. https://doi.org/10.1016/j.ajhg.2012.11.014
Morgan NV, Westaway SK, Morton JE, Gregory A, Gissen P, Sonek S, Cangul H, Coryell J, Canham N, Nardocci N, Zorzi G, Pasha S, Rodriguez D, Desguerre I, Mubaidin A, Bertini E, Trembath RC, Simonati A, Schanen C, Hayflick SJ (2006) PLA2G6, encoding a phospholipase A2, is mutated in neurodegenerative disorders with high brain iron. Nat Genet 38(7):752–754. https://doi.org/10.1038/ng1826
Nagy D, Verheyen S, Wigby KM, Borovikov A, Sharkov A, Slegesky V, Larson A, Fagerberg C, Brasch-Andersen C, Kibaek M, Bader I, Hernan, R, High FA, Chung WK, Schieving JH, Behunova J, Smogavec M, Laccone F, Witsch-Baumgartner M, Weis D (2022) Genotype-phenotype comparison in POGZ-related neurodevelopmental disorders by using clinical scoring. Genes (Basel). https://doi.org/10.3390/genes13010154
Ope O, Bhoj EJ, Nelson B, Li D, Hakonarson H, Sobering AK (2020) A homozygous truncating NALCN variant in two Afro-Caribbean siblings with hypotonia and dolichocephaly. Am J Med Genet A 182(8):1877–1880. https://doi.org/10.1002/ajmg.a.61744
Paisan-Ruiz C, Bhatia KP, Li A, Hernandez D, Davis M, Wood NW, Hardy J, Houlden H, Singleton A, Schneider SA (2009) Characterization of PLA2G6 as a locus for dystonia-parkinsonism. Ann Neurol 65(1):19–23. https://doi.org/10.1002/ana.21415
Reuter MS, Tawamie H, Buchert R, Hosny Gebril O, Froukh T, Thiel C, Uebe S, Ekici AB, Krumbiegel M, Zweier C, Hoyer J, Eberlein K, Bauer J, Scheller U, Strom TM, Hoffjan S, Abdelraouf ER, Meguid NA, Abboud A, Abou Jamra R (2017) Diagnostic Yield and Novel Candidate Genes by Exome Sequencing in 152 Consanguineous Families With Neurodevelopmental Disorders. JAMA Psychiat 74(3):293–299. https://doi.org/10.1001/jamapsychiatry.2016.3798
Riazuddin S, Hussain M, Razzaq A, Iqbal Z, Shahzad M, Polla DL, Song Y, van Beusekom E, Khan AA, Tomas-Roca L, Rashid M, Zahoor MY, Wissink-Lindhout WM, Basra MAR, Ansar M, Agha Z, van Heeswijk K, Rasheed F, Van de Vorst M, Riazuddin S (2017) Exome sequencing of Pakistani consanguineous families identifies 30 novel candidate genes for recessive intellectual disability. Mol Psychiatry 22(11):1604–1614. https://doi.org/10.1038/mp.2016.109
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL, Committee ALQA (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Rupps R, Hukin J, Balicki M, Mercimek-Mahmutoglu S, Rolfs A, Dias C (2013) Novel Mutations in FA2H-Associated Neurodegeneration: An Underrecognized Condition? J Child Neurol 28(11):1500–1504. https://doi.org/10.1177/0883073812458538
Sainio MT, Aaltio J, Hyttinen V, Kortelainen M, Ojanen S, Paetau A, Tienari P, Ylikallio E, Auranen M, Tyynismaa H (2022) Effectiveness of clinical exome sequencing in adult patients with difficult-to-diagnose neurological disorders. Acta Neurol Scand 145(1):63–72. https://doi.org/10.1111/ane.13522
Santos-Cortez RLP, Khan V, Khan FS, Mughal ZU, Chakchouk I, Lee K, Rasheed M, Hamza R, Acharya A, Ullah E, Saqib MAN, Abbe I, Ali G, Hassan MJ, Khan S, Azeem Z, Ullah I, Bamshad MJ, Nickerson DA, Leal SM (2018) Novel candidate genes and variants underlying autosomal recessive neurodevelopmental disorders with intellectual disability. Hum Genet 137(9):735–752. https://doi.org/10.1007/s00439-018-1928-6
Sapey-Triomphe LA, Reversat J, Lesca G, Chatron N, Bussa M, Mazoyer S, Schmitz C, Sonie S, Edery P (2020) A de novo frameshift pathogenic variant in TBR1 identified in autism without intellectual disability. Hum Genomics 14(1):32. https://doi.org/10.1186/s40246-020-00281-5
Sugama S, Kimura A, Chen W, Kubota S, Seyama Y, Taira N, Eto Y (2001) Frontal lobe dementia with abnormal cholesterol metabolism and heterozygous mutation in sterol 27-hydroxylase gene (CYP27). J Inherit Metab Dis 24(3):379–392. https://doi.org/10.1023/a:1010564920930
Tekin D, Yan D, Bademci G, Feng Y, Guo S, Foster J 2nd, Blanton S, Tekin M, Liu X (2016) A next-generation sequencing gene panel (MiamiOtoGenes) for comprehensive analysis of deafness genes. Hear Res 333:179–184. https://doi.org/10.1016/j.heares.2016.01.018
Vetro A, Pisano T, Chiaro S, Procopio E, Guerra A, Parrini E, Mei D, Virdo S, Mangone G, Azzari C, Guerrini R (2020) Early infantile epileptic-dyskinetic encephalopathy due to biallelic PIGP mutations. Neurol Genet 6(1):e387. https://doi.org/10.1212/NXG.0000000000000387
White J, Beck CR, Harel T, Posey JE, Jhangiani SN, Tang S, Farwell KD, Powis Z, Mendelsohn NJ, Baker JA, Pollack L, Mason KJ, Wierenga KJ, Arrington DK, Hall M, Psychogios A, Fairbrother L, Walkiewicz M, Person RE, Sutton VR (2016) POGZ truncating alleles cause syndromic intellectual disability. Genome Med 8(1):3. https://doi.org/10.1186/s13073-015-0253-0
Wirth T, Weibel S, Montaut S, Bigaut K, Rudolf G, Chelly J, Tranchant C, Anheim M (2017) Severe early-onset impulsive compulsive behavior and psychosis in PLA2G6-related juvenile Parkinson’s disease. Parkinsonism Relat Disord 41:127–129. https://doi.org/10.1016/j.parkreldis.2017.05.014
Xue Y, Ankala A, Wilcox WR, Hegde MR (2015) Solving the molecular diagnostic testing conundrum for Mendelian disorders in the era of next-generation sequencing: single-gene, gene panel, or exome/genome sequencing. Genet Med 17(6):444–451. https://doi.org/10.1038/gim.2014.122
Yoshino H, Tomiyama H, Tachibana N, Ogaki K, Li Y, Funayama M, Hashimoto T, Takashima S, Hattori N (2010) Phenotypic spectrum of patients with PLA2G6 mutation and PARK14-linked parkinsonism. Neurology 75(15):1356–1361. https://doi.org/10.1212/WNL.0b013e3181f73649
Acknowledgements
The authors are grateful to the patients and their families for taking part in this study.
Funding
This work was supported by grants from Higher Education Commission of Pakistan (NRPU 20-12107 and NRPU 7451), National Natural Science Foundation of China (31625015, 31521003 and 32100480), China Postdoctoral Science Foundation (2020TQ0072), Shanghai Municipal Science and Technology Major Project (2017SHZDZX01), and the 111 Project (B13016). The aforementioned grants covered the costs of materials required for DNA extraction and expenses of whole exome sequencing and Sanger sequencing. AK was supported by HEC’s IRSIP fellowship.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Web resources
1000 Genomes Project, https://www.internationalgenome.org. Genome Aggregation Database (gnomAD), http://gnomad.broadinstitute.org. The Human Gene Mutation Database (HGMD), http://www.hgmd.cf.ac.uk.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, A., Tian, S., Tariq, M. et al. NGS-driven molecular diagnosis of heterogeneous hereditary neurological disorders reveals novel and known variants in disease-causing genes. Mol Genet Genomics 297, 1601–1613 (2022). https://doi.org/10.1007/s00438-022-01945-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01945-8