Abstract
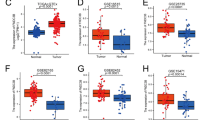
The delayed diagnosis of pancreatic cancer has resulted in rising mortality rate and low survival rate that can be circumvented using potent theranostics biomarkers. The treatment gets complicated with delayed detection resulting in lowered 5-year relative survival rate. In our present study, we employed systems biology approach to identify central genes that play crucial roles in tumor progression. Pancreatic cancer genes collected from various databases were used to construct a statistically significant interactome with 812 genes that was further analysed thoroughly using topological parameters and functional enrichment analysis. The significant genes in the network were then identified based on the maximum degree parameter. The overall survival analysis indicated through hazard ratio [HR] and gene expression [log Fold Change] across pancreatic adenocarcinoma revealed the critical role of FN1 [HR 1.4; log2(FC) 5.748], FGA [HR 0.78; log2(FC) 1.639] FGG [HR 0.9; log2(FC) 1.597], C3 [HR 1.1; log2(FC) 2.637], and QSOX1 [HR 1.4; log2(FC) 2.371]. The functional significance of the identified hub genes signified the enrichment of integrin cell surface interactions and proteoglycan syndecan-mediated cell signaling. The differential expression, low overall survival and functional significance of FN1 gene implied its possible role in controlling metastasis in pancreatic cancer. Furthermore, alternate splice variants of FN1 gene showed 10 protein coding transcripts with conserved cell attachment site and functional domains indicating the variants’ potential role in pancreatic cancer. The strong association of the identified hub-genes can be better directed to design potential theranostics biomarkers for metastasized pancreatic tumor.






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Aggarwal G, Rabe KG, Petersen GM, Chari ST (2012) New-onset diabetes in pancreatic cancer: a study in the primary care setting. Pancreatology 12:156–161. https://doi.org/10.1016/J.PAN.2012.02.003
Ashok G, Miryala SK, Anbarasu A, Ramaiah S (2021) Integrated systems biology approach using gene network analysis to identify the important pathways and new potential drug targets for Neuroblastoma. Gene Rep 23:101101. https://doi.org/10.1016/j.genrep.2021.101101
Bader GD, Hogue CWV (2003) An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4:2. https://doi.org/10.1186/1471-2105-4-2
Baharlou Houreh M, Ghorbani Kalkhajeh P, Niazi A et al (2018) SpliceDetector: a software for detection of alternative splicing events in human and model organisms directly from transcript IDs. Sci Rep 8:5063. https://doi.org/10.1038/s41598-018-23245-1
Barkovskaya A, Buffone A, Žídek M, Weaver VM (2020) Proteoglycans as mediators of cancer tissue mechanics. Front Cell Dev Biol 8:569377
Basu S, Naha A, Veeraraghavan B et al (2021) In silico structure evaluation of BAG3 and elucidating its association with bacterial infections through protein–protein and host-pathogen interaction analysis. J Cell Biochem. https://doi.org/10.1002/jcb.29953
Beauvais DLM, Rapraeger AC (2004) Syndecans in tumor cell adhesion and signaling. Reprod Biol Endocrinol 2:3. https://doi.org/10.1186/1477-7827-2-3
Buscail L, Bournet B, Cordelier P (2020) Role of oncogenic KRAS in the diagnosis, prognosis and treatment of pancreatic cancer. Nat Rev Gastroenterol Hepatol 17:153–168. https://doi.org/10.1038/s41575-019-0245-4
Cai X, Liu C, Zhang T et al (2018) Down-regulation of FN1 inhibits colorectal carcinogenesis by suppressing proliferation, migration, and invasion. J Cell Biochem 119:4717–4728. https://doi.org/10.1002/jcb.26651
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data: Figure 1. Cancer Discov 2:401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Chari ST, Zapiach M, Yadav D, Rizza RA (2005) Beta-cell function and insulin resistance evaluated by HOMA in pancreatic cancer subjects with varying degrees of glucose intolerance. Pancreatology 5:229–233. https://doi.org/10.1159/000085276
Chen J, Wu W, Chen L et al (2013a) Profiling the potential tumor markers of pancreatic ductal adenocarcinoma using 2D-DIGE and MALDI-TOF-MS: up-regulation of complement C3 and alpha-2-HS-glycoprotein. Pancreatology 13:290–297. https://doi.org/10.1016/j.pan.2013.03.010
Chen J, Wu W, Zhen C et al (2013b) Expression and clinical significance of complement C3, complement C4b1 and apolipoprotein E in pancreatic cancer. Oncol Lett 6:43–48. https://doi.org/10.3892/ol.2013.1326
Debroy R, Miryala SK, Naha A et al (2020) Gene interaction network studies to decipher the multi-drug resistance mechanism in Salmonella enterica serovar Typhi CT18 reveal potential drug targets. Microb Pathog 142:104096. https://doi.org/10.1016/j.micpath.2020.104096
Doi Y, Yashiro M, Yamada N et al (2012) VEGF-A/VEGFR-2 signaling plays an important role for the motility of pancreas cancer cells. Ann Surg Oncol 19:2733–2743. https://doi.org/10.1245/s10434-011-2181-6
Duan S, Gong B, Wang P et al (2018) Novel prognostic biomarkers of gastric cancer based on gene expression microarray: COL12A1, GSTA3, FGA and FGG. Mol Med Rep 18:3727–3736. https://doi.org/10.3892/mmr.2018.9368
Gao J, Mazor T, Ciftci E et al (2018) Abstract 923: the cBioPortal for cancer genomics: an intuitive open-source platform for exploration, analysis and visualization of cancer genomics data. Cancer Res 78:923–923. https://doi.org/10.1158/1538-7445.AM2018-923
Gouy M, Tannier E, Comte N, Parsons DP (2021) Seaview version 5: a multiplatform software for multiple sequence alignment, molecular phylogenetic analyses, and tree reconciliation, pp 241–260
Grzesiak JJ, Cao HST, Burton DW et al (2011) Knockdown of the β1 integrin subunit reduces primary tumor growth and inhibits pancreatic cancer metastasis. Int J Cancer 129:2905–2915. https://doi.org/10.1002/ijc.25942
Herreros-Villanueva M, Bujanda L (2016) Glypican-1 in exosomes as biomarker for early detection of pancreatic cancer. Ann Transl Med. https://doi.org/10.3978/J.ISSN.2305-5839.2015.10.39
Hu D, Ansari D, Zhou Q et al (2019) Stromal fibronectin expression in patients with resected pancreatic ductal adenocarcinoma. World J Surg Oncol 17:1–8. https://doi.org/10.1186/s12957-019-1574-z
Kaczmarek J, Castellani P, Nicolo G et al (1994) Distribution of oncofetal fibronectin isoforms in normal, hyperplastic and neoplastic human breast tissues. Int J Cancer 59:11–16. https://doi.org/10.1002/IJC.2910590104
Katchman BA, Antwi K, Hostetter G et al (2011) Quiescin sulfhydryl oxidase 1 promotes invasion of pancreatic tumor cells mediated by matrix metalloproteinases. Mol Cancer Res 9:1621–1631. https://doi.org/10.1158/1541-7786.MCR-11-0018
Korsse SE, Peppelenbosch MP, van Veelen W (2013) Targeting LKB1 signaling in cancer. Biochim Biophys Acta Rev Cancer 1835:194–210. https://doi.org/10.1016/j.bbcan.2012.12.006
Li C, Tang Z, Zhang W et al (2021) GEPIA2021: integrating multiple deconvolution-based analysis into GEPIA. Nucleic Acids Res 49:W242–W246. https://doi.org/10.1093/NAR/GKAB418
Liang B, Ding H, Huang L et al (2020) GWAS in cancer: progress and challenges. Mol Genet Genom 295:537–561. https://doi.org/10.1007/S00438-020-01647-Z
Lieverse RIY, Marcus D, Wiel AMA et al (2020) Human fibronectin extra domain B as a biomarker for targeted therapy in cancer. Mol Oncol 14:1555–1568. https://doi.org/10.1002/1878-0261.12705
Lopes CT, Franz M, Kazi F et al (2010) Cytoscape web: an interactive web-based network browser. Bioinformatics 26:2347–2348. https://doi.org/10.1093/bioinformatics/btq430
Luo H, Xu X, Yang J et al (2020) Genome-wide somatic copy number alteration analysis and database construction for cervical cancer. Mol Genet Genom 2953(295):765–773. https://doi.org/10.1007/S00438-019-01636-X
Melo SA, Luecke LB, Kahlert C et al (2015) Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523:177–182. https://doi.org/10.1038/nature14581
Mihaljevic AL, Michalski CW, Friess H, Kleeff J (2010) Molecular mechanism of pancreatic cancer—understanding proliferation, invasion, and metastasis. Langenbeck’s Arch Surg 395:295–308
Miryala SK, Anbarasu A, Ramaiah S (2018) Discerning molecular interactions: a comprehensive review on biomolecular interaction databases and network analysis tools. Gene 642:84–94. https://doi.org/10.1016/j.gene.2017.11.028
Miryala SK, Ramaiah S (2021) Cellular and molecular level host-pathogen interactions in Francisella tularensis: a microbial gene network study. Comput Biol Chem 96:107601. https://doi.org/10.1016/j.compbiolchem.2021.107601
Mori T, Doi R, Toyoda E et al (2005) Regulation of the resistance to TRAIL-induced apoptosis as a new strategy for pancreatic cancer. Surgery 138:71–77. https://doi.org/10.1016/j.surg.2005.03.001
Morton JP, Jamieson NB, Karim SA et al (2010) LKB1 haploinsufficiency cooperates with Kras to promote pancreatic cancer through suppression of p21-dependent growth arrest. Gastroenterology 139:586-597.e6. https://doi.org/10.1053/j.gastro.2010.04.055
Naha A, Kumar Miryala S, Debroy R et al (2020) Elucidating the multi-drug resistance mechanism of Enterococcus faecalis V583: a gene interaction network analysis. Gene 748:144704. https://doi.org/10.1016/J.GENE.2020.144704
Nelakurti DD, Pappula AL, Rajasekaran S et al (2020) Comprehensive analysis of MEN1 mutations and their role in cancer. Cancers (basel) 12:2616. https://doi.org/10.3390/cancers12092616
Pagano MT, Ortona E, Dupuis ML (2020) A role for estrogen receptor alpha36 in cancer progression. Front Endocrinol (lausanne). https://doi.org/10.3389/fendo.2020.00506
Pan X, Hu XH, Zhang YH et al (2018) Identification of the copy number variant biomarkers for breast cancer subtypes. Mol Genet Genom 2941(294):95–110. https://doi.org/10.1007/S00438-018-1488-4
Pathan M, Keerthikumar S, Ang C-S et al (2015) FunRich: an open access standalone functional enrichment and interaction network analysis tool. Proteomics 15:2597–2601. https://doi.org/10.1002/pmic.201400515
Priyamvada P, Debroy R, Anbarasu A, Ramaiah S (2022) A comprehensive review on genomics, systems biology and structural biology approaches for combating antimicrobial resistance in ESKAPE pathogens: computational tools and recent advancements. World J Microbiol Biotechnol 38:153. https://doi.org/10.1007/s11274-022-03343-z
Quan B, Qi X, Yu Z et al (2014) Pathway analysis of genome-wide association study and transcriptome data highlights new biological pathways in colorectal cancer. Mol Genet Genom 2902(290):603–610. https://doi.org/10.1007/S00438-014-0945-Y
Saito R, Smoot ME, Ono K et al (2012) A travel guide to Cytoscape plugins. Nat Methods 911(9):1069–1076. https://doi.org/10.1038/nmeth.2212
Santana-Codina N, Roeth AA, Zhang Y et al (2018) Oncogenic KRAS supports pancreatic cancer through regulation of nucleotide synthesis. Nat Commun 91(9):1–13. https://doi.org/10.1038/s41467-018-07472-8
Shao DD, Xue W, Krall EB et al (2014) KRAS and YAP1 converge to regulate EMT and tumor survival. Cell 158:171–184. https://doi.org/10.1016/j.cell.2014.06.004
Sheng W, Chen C, Dong M et al (2017) Calreticulin promotes EGF-induced EMT in pancreatic cancer cells via Integrin/EGFR-ERK/MAPK signaling pathway. Cell Death Dis 8:e3147. https://doi.org/10.1038/cddis.2017.547
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. CA Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
Singla S, Pippin JA, Drebin JA (2012) Dual ErbB1 and ErbB2 receptor tyrosine kinase inhibition exerts synergistic effect with conventional chemotherapy in pancreatic cancer. Oncol Rep 28:2211–2216. https://doi.org/10.3892/or.2012.2053
Staton CA, Brown NJ, Lewis CE (2003) The role of fibrinogen and related fragments in tumour angiogenesis and metastasis. Expert Opin Biol Ther 3:1105–1120. https://doi.org/10.1517/14712598.3.7.1105
Szklarczyk D, Gable AL, Nastou KC et al (2021) The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49:605–612. https://doi.org/10.1093/nar/gkaa1074
Szklarczyk D, Morris JH, Cook H et al (2017) The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res 45:362–368. https://doi.org/10.1093/nar/gkw937
Tang Z, Li C, Kang B et al (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Web Serv Issue Publ Online. https://doi.org/10.1093/nar/gkx247
Thomas JK, Kim MS, Balakrishnan L et al (2014) Pancreatic cancer database: an integrative resource for pancreatic cancer. Cancer Biol Ther 15:963–967. https://doi.org/10.4161/cbt.29188
von Mering C, Huynen M, Jaeggi D et al (2003) STRING: a database of predicted functional associations between proteins. Nucleic Acids Res 31:258–261. https://doi.org/10.1093/nar/gkg034
Wang N, Wang D (2021) Genome-wide transcriptome and translatome analyses reveal the role of protein extension and domestication in liver cancer oncogenesis. Mol Genet Genom 296:561–569. https://doi.org/10.1007/S00438-021-01766-1/FIGURES/5
Wang M, Zhang G, Zhang Y et al (2020) Fibrinogen alpha chain knockout promotes tumor growth and metastasis through integrin-akt signaling pathway in lung cancer. Mol Cancer Res 18:943–954. https://doi.org/10.1158/1541-7786.MCR-19-1033
Weinel RJ, Rosendahl A, Pinschmidt E et al (1995) The α6-integrin receptor in pancreatic carcinoma. Gastroenterology 108:523–532. https://doi.org/10.1016/0016-5085(95)90082-9
Zhang H, Sun Z, Li Y et al (2017) MicroRNA-200c binding to FN1 suppresses the proliferation, migration and invasion of gastric cancer cells. Biomed Pharmacother 88:285–292. https://doi.org/10.1016/j.biopha.2017.01.023
Acknowledgements
The authors would like to thank VIT management for providing the necessary facilities to carry out this research work. The authors gratefully acknowledge the Indian Council of Medical Research (ICMR), the Government of India agency, for the research grant. Gayathri Ashok and Megha Treesa Saju would also like to extend sincere thanks to Mr. Aniket Naha, ICMR-RA for his intellectual inputs and guidance while writing the manuscript.
Funding
The authors gratefully acknowledge the Indian Council of Medical Research (ICMR), the Government of India agency, for the research grant (IRIS ID:2020-0690).
Author information
Authors and Affiliations
Contributions
SR: Conceptualization, funding acquisition, and supervision. AA: Conceptualization, methodology, and supervision. GA: Data curation, formal analysis, visualization, writing-original draft preparation SKM: Data analysis, network validation, manuscript-editing and reviewing. MTS: Data curation, data analysis of gene networks, writing-original draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ashok, G., Miryala, S.K., Saju, M.T. et al. FN1 encoding fibronectin as a pivotal signaling gene for therapeutic intervention against pancreatic cancer. Mol Genet Genomics 297, 1565–1580 (2022). https://doi.org/10.1007/s00438-022-01943-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01943-w