Abstract
Development of colon adenocarcinoma (COAD) metastasis involves several mediators including fluid shear stress (FSS), intracellular ROS levels, and non-coding RNAs. In our present study, we identified and investigated the role of regulatory non-coding RNA molecules specifically involved in COAD metastasis and their association with FSS and ROS. Interactions between the mRNAs associated with FSS and ROS, the corresponding microRNAs (miRNAs), long noncoding RNAs (lncRNAs) and circular RNAs (circRNAs) in COAD metastasis were used to generate the mRNA-miRNA-lncRNA-circRNA network. Experimental validation of the identified RNA hubs using quantitative real-time PCR demonstrated a direct effect of the FSS on their expression levels in cancer cells. FSS resulted in the downregulation of HMGA1 and RAN, as well as the upregulation of HSP90AA1, PMAIP1 and BIRC5. Application of shear stress also led to downregulation of hsa-miR-26b-5p and hsa-miR-34a-5p levels in HCT116 cells. Further, functional enrichment and survival analysis of the significant miRNAs, as well as the OncoPrint and the survival analyses of the selected mRNAs were performed. Subsequently, their functional role was also corroborated with existing literature. Ten significant miRNA hubs were identified, out of which hsa-miR-17-5p and hsa-miR-20a-5p were found to interact with lncRNA (CCAT2) while hsa-miR-335 was found to interact with four circRNAs. Fifteen significant miRNAs were identified in 10 different modules suggesting their importance in FSS and ROS-mediated COAD metastasis. Finally, 10 miRNAs and 3 mRNAs associated with FSS and/or ROS were identified as significant overall survival markers; 33 mRNAs were also identified as metastasis-free survival markers whereas 15 mRNAs showed > 10% gene alterations in TCGA-COAD data and may serve as promising therapeutic biomarkers in the COAD metastasis.








Similar content being viewed by others
Abbreviations
- COAD:
-
Colon adenocarcinoma
- GO:
-
Gene Ontology
- FSS:
-
Fluid shear stress
- ROS:
-
Reactive oxygen species
- HCMDB:
-
Human cancer metastasis database
- GEPIA:
-
Gene expression profiling interactive analysis
- DE:
-
Differentially expressed
- dbDEMC:
-
Database of differentially expressed microRNAs
- TCGA:
-
The Cancer Genome Atlas
- miRNA:
-
MicroRNA
- lncRNA:
-
Long coding RNA
- circRNA:
-
Circular RNA
- miEAA:
-
MicroRNA Enrichment Analysis and Annotation
References
Agarwal E, Brattain MG, Chowdhury S (2013) Cell survival and metastasis regulation by Akt signaling in colorectal cancer. Cell Signal 25:1711–1719
Aguirre-Portoles C, Feliu J, Reglero G, Ramirez de Molina A (2018) ABCA1 overexpression worsens colorectal cancer prognosis by facilitating tumour growth and caveolin-1-dependent invasiveness, and these effects can be ameliorated using the BET inhibitor apabetalone. Mol Oncol 12:1735–1752
Backes C, Khaleeq QT, Meese E, Keller A (2016) miEAA: microRNA enrichment analysis and annotation. Nucleic Acids Res 44:W110-116
Banerjee S, Karunagaran D (2019) An integrated approach for mining precise RNA-based cervical cancer staging biomarkers. Gene 712:143961
Banerjee S, Kalyani Yabalooru SR, Karunagaran D (2020) Identification of mRNA and non-coding RNA hubs using network analysis in organ tropism regulated triple negative breast cancer metastasis. Comput Biol Med 127:104076
Barnes JM, Nauseef JT, Henry MD (2012) Resistance to fluid shear stress is a conserved biophysical property of malignant cells. PLoS ONE 7:e50973
Bolha L, Ravnik-Glavac M, Glavac D (2017) Long Noncoding RNAs as Biomarkers in Cancer. Dis Markers 2017:7243968
Cai Y, Yu X, Hu S, Yu J (2009) A brief review on the mechanisms of miRNA regulation. Genomics Proteomics Bioinformatics 7:147–154
Calon A, Espinet E, Palomo-Ponce S, Tauriello DV, Iglesias M, Cespedes MV, Sevillano M, Nadal C, Jung P, Zhang XH, Byrom D, Riera A, Rossell D, Mangues R, Massague J, Sancho E, Batlle E (2012) Dependency of colorectal cancer on a TGF-beta-driven program in stromal cells for metastasis initiation. Cancer Cell 22:571–584
Chaffer CL, Weinberg RA (2011) A perspective on cancer cell metastasis. Science 331:1559–1564
Chakraborty C, Sharma AR, Sharma G, Doss CGP, Lee SS (2017) Therapeutic miRNA and siRNA: moving from bench to clinic as next generation medicine. Mol Ther Nucleic Acids 8:132–143
Chen LL, Yang L (2015) Regulation of circRNA biogenesis. RNA Biol 12:381–388
Chen J, Elfiky A, Han M, Chen C, Saif MW (2014) The role of Src in colon cancer and its therapeutic implications. Clin Colorectal Cancer 13:5–13
Chen W, Cai G, Liao Z, Lin K, Li G, Li Y (2019a) miRNA-766 induces apoptosis of human colon cancer cells through the p53/Bax signaling pathway by MDM4. Exp Ther Med 17:4100–4108
Chen Z, Ren R, Wan D, Wang Y, Xue X, Jiang M, Shen J, Han Y, Liu F, Shi J, Kuang Y, Li W, Zhi Q (2019b) Hsa_circ_101555 functions as a competing endogenous RNA of miR-597-5p to promote colorectal cancer progression. Oncogene 38:6017–6034
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11
Choi HY, Yang GM, Dayem AA, Saha SK, Kim K, Yoo Y, Hong K, Kim JH, Yee C, Lee KM, Cho SG (2019) Hydrodynamic shear stress promotes epithelial-mesenchymal transition by downregulating ERK and GSK3beta activities. Breast Cancer Res 21:6
Cima I, Kong SL, Sengupta D, Tan IB, Phyo WM, Lee D, Hu M, Iliescu C, Alexander I, Goh WL, Rahmani M, Suhaimi NA, Vo JH, Tai JA, Tan JH, Chua C, Ten R, Lim WJ, Chew MH, Hauser CA, van Dam RM, Lim WY, Prabhakar S, Lim B, Koh PK, Robson P, Ying JY, Hillmer AM, Tan MH (2016) Tumor-derived circulating endothelial cell clusters in colorectal cancer. Sci Transl Med 8:345ra389
Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M (2016) CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 13:34–42
Fang Y, Fullwood MJ (2016) Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genomics Proteomics Bioinformatics 14:42–54
Fosselteder J, Calin GA, Pichler M (2018) Long non-coding RNA CCAT2 as a therapeutic target in colorectal cancer. Expert Opin Ther Targets 22:973–976
Fu A, Ma S, Wei N, Tan BX, Tan EY, Luo KQ (2016) High expression of MnSOD promotes survival of circulating breast cancer cells and increases their resistance to doxorubicin. Oncotarget 7:50239–50257
Galadari S, Rahman A, Pallichankandy S, Thayyullathil F (2017) Reactive oxygen species and cancer paradox: to promote or to suppress? Free Radic Biol Med 104:144–164
Galperin MY, Fernández-Suárez XM, Rigden DJ (2017) The 24th annual Nucleic Acids Research database issue: a look back and upcoming changes. Nucleic Acids Res 45:5627–5627
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:l1
Gao Z, Fu P, Yu Z, Zhen F, Gu Y (2019) Comprehensive analysis of lncRNA-miRNA-mRNA network ascertains prognostic factors in patients with colon cancer. Technol Cancer Res Treat 18:1533033819853237
Goswami CP, Nakshatri H (2014) PROGgeneV2: enhancements on the existing database. BMC Cancer 14:970
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388
Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R (2015) InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics 16:169
Hentze MW, Preiss T (2013) Circular RNAs: splicing’s enigma variations. EMBO J 32:923–925
Huang CW, Chen YT, Tsai HL, Yeh YS, Su WC, Ma CJ, Tsai TN, Wang JY (2017) EGFR expression in patients with stage III colorectal cancer after adjuvant chemotherapy and on cancer cell function. Oncotarget 8:114663–114676
Huang Q, Hu X, He W, Zhao Y, Hao S, Wu Q, Li S, Zhang S, Shi M (2018) Fluid shear stress and tumor metastasis. Am J Cancer Res 8:763–777
Jia B, Xia L, Cao F (2018) The role of miR-766-5p in cell migration and invasion in colorectal cancer. Exp Ther Med 15:2569–2574
Jing X, Liang H, Cui X, Han C, Hao C, Huo K (2017) Long noncoding RNA CCAT2 can predict metastasis and a poor prognosis: a meta-analysis. Clin Chim Acta 468:159–165
Kang T, Park C, Meghani N, Tran TTD, Tran PHL, Lee BJ (2020) Shear stress-dependent targeting efficiency using self-assembled gelatin-oleic nanoparticles in a biomimetic microfluidic system. Pharmaceutics 12:555
Kim NH, Kim HS, Kim NG, Lee I, Choi HS, Li XY, Kang SE, Cha SY, Ryu JK, Na JM, Park C, Kim K, Lee S, Gumbiner BM, Yook JI, Weiss SJ (2011) p53 and microRNA-34 are suppressors of canonical Wnt signaling. Sci Signal 4:ra71
Kumar S, Kim CW, Simmons RD, Jo H (2014) Role of flow-sensitive microRNAs in endothelial dysfunction and atherosclerosis: mechanosensitive athero-miRs. Arterioscler Thromb Vasc Biol 34:2206–2216
Lan J, Huang Z, Han J, Shao J, Huang C (2018) Redox regulation of microRNAs in cancer. Cancer Lett 418:250–259
Lee DY, Lin TE, Lee CI, Zhou J, Huang YH, Lee PL, Shih YT, Chien S, Chiu JJ (2017) MicroRNA-10a is crucial for endothelial response to different flow patterns via interaction of retinoid acid receptors and histone deacetylases. Proc Natl Acad Sci USA 114:2072–2077
Li J-H, Liu S, Zhou H, Qu L-H, Yang J-H (2013) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97
Li Y, Sun Z, Liu B, Shan Y, Zhao L, Jia L (2017) Tumor-suppressive miR-26a and miR-26b inhibit cell aggressiveness by regulating FUT4 in colorectal cancer. Cell Death Dis 8:e2892–e2892
Li J, Xia L, Zhou Z, Zuo Z, Xu C, Song H, Cai J (2018) MiR-186-5p upregulation inhibits proliferation, metastasis and epithelial-to-mesenchymal transition of colorectal cancer cell by targeting ZEB1. Arch Biochem Biophys 640:53–60
Lien SC, Chang SF, Lee PL, Wei SY, Chang MD, Chang JY, Chiu JJ (2013) Mechanical regulation of cancer cell apoptosis and autophagy: roles of bone morphogenetic protein receptor, Smad1/5, and p38 MAPK. Biochim Biophys Acta 1833:3124–3133
Ling H, Spizzo R, Atlasi Y, Nicoloso M, Shimizu M, Redis RS, Nishida N, Gafa R, Song J, Guo Z, Ivan C, Barbarotto E, De Vries I, Zhang X, Ferracin M, Churchman M, van Galen JF, Beverloo BH, Shariati M, Haderk F, Estecio MR, Garcia-Manero G, Patijn GA, Gotley DC, Bhardwaj V, Shureiqi I, Sen S, Multani AS, Welsh J, Yamamoto K, Taniguchi I, Song MA, Gallinger S, Casey G, Thibodeau SN, Le Marchand L, Tiirikainen M, Mani SA, Zhang W, Davuluri RV, Mimori K, Mori M, Sieuwerts AM, Martens JW, Tomlinson I, Negrini M, Berindan-Neagoe I, Foekens JA, Hamilton SR, Lanza G, Kopetz S, Fodde R, Calin GA (2013) CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res 23:1446–1461
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(− Delta Delta C(T)) method. Methods 25:402–408
Ma S, Fu A, Chiew GG, Luo KQ (2017) Hemodynamic shear stress stimulates migration and extravasation of tumor cells by elevating cellular oxidative level. Cancer Lett 388:239–248
Makondi PT, Wei PL, Huang CY, Chang YJ (2019) Development of novel predictive miRNA/target gene pathways for colorectal cancer distance metastasis to the liver using a bioinformatic approach. PLoS ONE 14:e0211968
Marin T, Gongol B, Chen Z, Woo B, Subramaniam S, Chien S, Shyy JY (2013) Mechanosensitive microRNAs-role in endothelial responses to shear stress and redox state. Free Radic Biol Med 64:61–68
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338
Mitchell MJ, King MR (2013) Fluid shear stress sensitizes cancer cells to receptor-mediated apoptosis via trimeric death receptors. New J Phys 15:015008
Nakayama M, Oshima M (2018) Mutant p53 in colon cancer. J Mol Cell Biol 11:267–276
Neth P, Nazari-Jahantigh M, Schober A, Weber C (2013) MicroRNAs in flow-dependent vascular remodelling. Cardiovasc Res 99:294–303
Northcott JM, Dean IS, Mouw JK, Weaver VM (2018) Feeling stress: the mechanics of cancer progression and aggression. Front Cell Dev Biol 6:17
Okumura K, Huang S, Sinicrope FA (2008) Induction of Noxa sensitizes human colorectal cancer cells expressing Mcl-1 to the small-molecule Bcl-2/Bcl-xL inhibitor, ABT-737. Clin Cancer Res 14:8132–8142
Olson MF, Sahai E (2009) The actin cytoskeleton in cancer cell motility. Clin Exp Metastasis 26:273–287
Paraskevopoulou MD, Vlachos IS, Karagkouni D, Georgakilas G, Kanellos I, Vergoulis T, Zagganas K, Tsanakas P, Floros E, Dalamagas T, Hatzigeorgiou AG (2016) DIANA-LncBase v2: indexing microRNA targets on non-coding transcripts. Nucleic Acids Res 44:D231-238
Pavlopoulos GA, Secrier M, Moschopoulos CN, Soldatos TG, Kossida S, Aerts J, Schneider R, Bagos PG (2011) Using graph theory to analyze biological networks. BioData Min 4:10
Qu S, Zhong Y, Shang R, Zhang X, Song W, Kjems J, Li H (2017) The emerging landscape of circular RNA in life processes. RNA Biol 14:992–999
Regmi S, Fung TS, Lim S, Luo KQ (2018) Fluidic shear stress increases the anti-cancer effects of ROS-generating drugs in circulating tumor cells. Breast Cancer Res Treat 172:297–312
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16:203–222
Saei H, Govahi A, Abiri A, Eghbali M, Abiri M (2021) Comprehensive transcriptome mining identified the gene expression signature and differentially regulated pathways of the late-onset preeclampsia. Pregnancy Hypertens 25:91–102
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7:e30733
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO (2013) Cell-type specific features of circular RNA expression. PLoS Genet 9:e1003777
Schumacker PT (2006) Reactive oxygen species in cancer cells: live by the sword, die by the sword. Cancer Cell 10:175–176
Shi L, Jackstadt R, Siemens H, Li H, Kirchner T, Hermeking H (2014) p53-induced miR-15a/16-1 and AP4 form a double-negative feedback loop to regulate epithelial-mesenchymal transition and metastasis in colorectal cancer. Cancer Res 74:532–542
Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin 69:7–34
Staudacher JJ, Bauer J, Jana A, Tian J, Carroll T, Mancinelli G, Ozden O, Krett N, Guzman G, Kerr D, Grippo P, Jung B (2017) Activin signaling is an essential component of the TGF-beta induced pro-metastatic phenotype in colorectal cancer. Sci Rep 7:5569
Su G, Morris JH, Demchak B, Bader GD (2014) Biological network exploration with Cytoscape 3. Curr Protoc Bioinformatics 47:11–24
Sun P, Zhu X, Shrubsole MJ, Ness RM, Hibler EA, Cai Q, Long J, Chen Z, Li G, Hou L, Smalley WE, Edwards TL, Giovannucci E, Zheng W, Dai Q (2017) Genetic variation in SLC7A2 interacts with calcium and magnesium intakes in modulating the risk of colorectal polyps. J Nutr Biochem 47:35–40
Sungwan P, Lert-Itthiporn W, Silsirivanit A, Klinhom-On N, Okada S, Wongkham S, Seubwai W (2021) Bioinformatics analysis identified CDC20 as a potential drug target for cholangiocarcinoma. PeerJ 9:e11067
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–W102
Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP (2007) Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 3:12
Vlachos IS, Paraskevopoulou MD, Karagkouni D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos I-L, Maniou S, Karathanou K, Kalfakakou D, Fevgas A, Dalamagas T, Hatzigeorgiou AG (2014) DIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res 43:D153–D159
Wang J, Wang X, Liu F, Fu Y (2017) microRNA-335 inhibits colorectal cancer HCT116 cells growth and epithelial-mesenchymal transition (EMT) process by targeting Twist1. Pharmazie 72:475–481
Wirtz D, Konstantopoulos K, Searson PC (2011) The physics of cancer: the role of physical interactions and mechanical forces in metastasis. Nat Rev Cancer 11:512–522
Wong WT, Ma S, Tian XY, Gonzalez AB, Ebong EE, Shen H (2016) Targeted delivery of shear stress-inducible micrornas by nanoparticles to prevent vulnerable atherosclerotic lesions. Methodist Debakey Cardiovasc J 12:152–156
Wu L, Hui H, Wang LJ, Wang H, Liu QF, Han SX (2015) MicroRNA-326 functions as a tumor suppressor in colorectal cancer by targeting the nin one binding protein. Oncol Rep 33:2309–2318
Xiao Y, Li ZH, Bi YH (2019) MicroRNA-889 promotes cell proliferation in colorectal cancer by targeting DAB2IP. Eur Rev Med Pharmacol Sci 23:3326–3334
Xu W, Zhang Z, Zou K, Cheng Y, Yang M, Chen H, Wang H, Zhao J, Chen P, He L, Chen X, Geng L, Gong S (2017) MiR-1 suppresses tumor cell proliferation in colorectal cancer by inhibition of Smad3-mediated tumor glycolysis. Cell Death Dis 8:e2761–e2761
Yang Z, Wu L, Wang A, Tang W, Zhao Y, Zhao H, Teschendorff AE (2017) dbDEMC 2.0: updated database of differentially expressed miRNAs in human cancers. Nucleic Acids Res 45:D812–D818
Yin J, Zeng X, Ai Z, Yu M, Wu Y, Li S (2020) Construction and analysis of a lncRNA-miRNA-mRNA network based on competitive endogenous RNA reveal functional lncRNAs in oral cancer. BMC Med Genomics 13:84
Yoo HI, Kim BK, Yoon SK (2016) MicroRNA-330-5p negatively regulates ITGA5 expression in human colorectal cancer. Oncol Rep 36:3023–3029
Zaki N, Efimov D, Berengueres J (2013) Protein complex detection using interaction reliability assessment and weighted clustering coefficient. BMC Bioinformatics 14:163
Zeilstra J, Joosten SP, Wensveen FM, Dessing MC, Schutze DM, Eldering E, Spaargaren M, Pals ST (2011) WNT signaling controls expression of pro-apoptotic BOK and BAX in intestinal cancer. Biochem Biophys Res Commun 406:1–6
Zhang Y, Wang J (2017) MicroRNAs are important regulators of drug resistance in colorectal cancer. Biol Chem 398:929–938
Zhang C, Tong J, Huang G (2013) Nicotinamide phosphoribosyl transferase (Nampt) is a target of microRNA-26b in colorectal cancer cells. PLoS ONE 8:e69963
Zhang Z, Kim K, Li X, Moreno M, Sharp T, Goodheart MJ, Safe S, Dupuy AJ, Amendt BA (2014) MicroRNA-26b represses colon cancer cell proliferation by inhibiting lymphoid enhancer factor 1 expression. Mol Cancer Ther 13:1942–1951
Zhang Z, Xie H, Liang D, Huang L, Liang F, Qi Q, Yang X (2018) Long non-coding RNA CCAT1 as a diagnostic and prognostic molecular marker in various cancers: a meta-analysis. Oncotarget 9:23695–23703
Zhang H, Wang R, Wang M (2019) miR-331-3p suppresses cell invasion and migration in colorectal carcinoma by directly targeting NRP2. Oncol Lett 18:6501–6508
Zheng G, Ma Y, Zou Y, Yin A, Li W, Dong D (2017) HCMDB: the human cancer metastasis database. Nucleic Acids Res 46:D950–D955
Acknowledgements
The authors would like to thank Dr. Swagatika Sahoo, Department of Chemical Engineering, IIT Madras, India and Dr. Karthik Raman, Department of Biotechnology, Bhupat and Jyoti Mehta School of Biosciences, IIT Madras, India for providing invaluable inputs to fine-tune and strengthen the analysis. We would also like to thank Priyanshu Sharma and Anand Kumar Patel, Department of Biotechnology, Bhupat and Jyoti Mehta School of Biosciences, IIT Madras, India for providing useful suggestions in the analysis of real-time PCR results.
Author information
Authors and Affiliations
Contributions
SK and SB conceptualized the work, processed the data and performed the network analysis. SK and SO performed the real-time PCR experiments and analyzed the data. DK and GK oversaw the work and provided directions to improve it. All the authors worked on the manuscript together.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by Martine Collart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2022_1924_MOESM1_ESM.docx
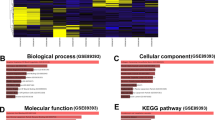
Supplementary file1 Supplementary Fig. 1a: Top 5 most significant Reactome pathways enriched with common miRNA interacting partners of FSS, ROS and HCMDB-COAD (p-value ≤ 0.05). Supplementary Fig. 1b: Top 5 most significant Gene Ontology terms enriched with common miRNA interacting partners of FSS, ROS and HCMDB-COAD (p-value ≤ 0.05). Supplementary Fig. 2a: ROS mRNA-mRNA-miRNA network. Supplementary Fig. 2b: FSS mRNA-mRNA-miRNA network. Supplementary Fig. 2c: HCMDB-COAD mRNA-mRNA-miRNA network. Supplementary Fig. 2d: Merged FRM mRNA-mRNA-miRNA network. Supplementary Fig. 3a: Degree based top 20 RNA hubs of ROS mRNA-mRNA-miRNA network. Supplementary Fig. 3b: Degree based top 20 RNA hubs of FSS mRNA-mRNA-miRNA network. Supplementary Fig. 3c: Degree based top 20 RNA hubs of HCMDB-COAD mRNA-mRNA-miRNA network. Supplementary Fig. 4: InteractiVenn diagram representing mRNA from degree-based top 20 hubs of individual FSS, ROS, HCMDB-COAD, merged FRM mRNA-mRNA-miRNA networks and the DE genes in COAD (GEPIA-TCGA-COAD). Supplementary Fig. 5: FRM COAD mRNA-miRNA-lncRNA-circRNA network. Supplementary Fig. 6: FRM COAD DE mRNA-DE miRNA-DE lncRNA-DE circRNA expression network. Supplementary Fig. 7: Module analysis using FRM mRNA-mRNA-miRNA network as the input: Extraction of modules showing direct association of identified miRNAs with either ROS and HCMDB-COAD or FSS and HCMDB-COAD. Supplementary Fig. 8: OncoPrint analysis of nine significant mRNAs associated with FSS, ROS, and HCMDB-COAD obtained using MSKCC metastasis data. (DOCX 2458 KB)
438_2022_1924_MOESM2_ESM.xlsx
Supplementary file2 Supplementary Table 1: Common and unique genes obtained from the InteractiVenn diagram representing genes associated with FSS, ROS, HCMDB-COAD and the DE genes in COAD (GEPIA-TCGA-COAD). Supplementary Table 2: Role of FSS- and ROS-associated genes in COAD metastasis based on a literature survey. Supplementary Table 3: List of 134 DE miRNAs (DE-miRs) – common among [DE miR COAD-FSS] and [DE miR COAD-ROS] and [DE miR COAD-HCMDB COAD]. Supplementary Table 4: Common miRNA-enriched pathways and their role in COAD metastasis based on a literature survey. Supplementary Table 5: List of miRNAs directly associated with FSS and ROS based on miEAA result data. Supplementary Table 6: Degree-based top 20 RNA hubs of ROS mRNA-mRNA-miRNA network. Supplementary Table 7: Degree-based top 20 RNA hubs of FSS mRNA-mRNA-miRNA network. Supplementary Table 8: Degree-based top 20 RNA hubs of HCMDB-COAD mRNA-mRNA-miRNA network. Supplementary Table 9: Degree-based top 20 RNA hubs of merged FRM mRNA-mRNA-miRNA network. Supplementary Table 10: Common mRNAs among top 20 degree-based hubs of individual FSS, ROS, HCMDB-COAD, FRM mRNA-mRNA-miRNA networks and the DE genes in COAD (GEPIA-TCGA-COAD). Supplementary Table 11: Degree-based top 20 RNA hubs of FRM COAD mRNA-miRNA- lncRNA-circRNA network. Supplementary Table 12: Degree-based top 20 RNA hubs of FRM COAD DE mRNA-DE miRNA-DE lncRNA-DE circRNA network. Supplementary Table 13: Variation of metastasis-free survival status of COAD patients with the expression levels of significant mRNAs associated with FSS and ROS. Supplementary Table 14: Variation of survival status of COAD patients with the expression levels of significant miRNAs and mRNAs associated with FSS and ROS using TCGA data. Supplementary Table 15: List of primers used in reverse transcription (RT) and quantitative real-time PCR (qRT-PCR) (XLSX 119 KB)
Rights and permissions
About this article
Cite this article
KrishnaPriya, S., Omer, S., Banerjee, S. et al. An integrated approach to understand fluid shear stress-driven and reactive oxygen species-mediated metastasis of colon adenocarcinoma through mRNA-miRNA-lncRNA-circRNA networks. Mol Genet Genomics 297, 1353–1370 (2022). https://doi.org/10.1007/s00438-022-01924-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01924-z