Abstract
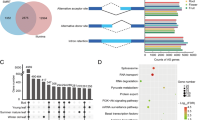
RNA alternative splicing (AS) is prevalent in higher organisms and plays a paramount role in biology; therefore, it is crucial to have comprehensive knowledge on AS to understand biology. However, knowledge is limited about how AS activates in a single plant and functions in a biological process. Ginseng is one of the most widely used medicinal herbs that is abundant in a number of medicinal bioactive components, especially ginsenosides. In this study, we sequenced the transcripts of 14 organs from a 4-year-old ginseng plant and quantified their ginsenoside contents. We identified AS genes by analyzing their transcripts with the ginseng genome and verified their AS events by PCR. The plant had a total of 13,863 AS genes subjected to 30,801 AS events with five mechanisms: skipped exon, retained intron, alternative 5′splice site, alternative 3′ splice site, and mutually exclusive exon. The genes that were more conserved, had more exons, and/or expressed across organs were more likely to be subjected to AS. AS genes were enriched in over 500 GO terms in the plant even though the number of AS gene-enriched GO terms varied across organs. At least 24 AS genes were found to be involved in ginsenoside biosynthesis. These AS genes were significantly up-enriched and more likely to form a co-expression network, thus suggesting the functions of AS and correlations of the AS genes in the process. This study provides comprehensive insights into the molecular characteristics and biological functions of AS in a single plant; thus, helping better understand biology.






Similar content being viewed by others
Data availability
The data analyzed for this study are available at SRA database of NCBI (SRP066368) and included within the article or supplementary files. The plant material is available through the corresponding authors, upon request.
References
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Breitbart RE, Andreadis A, Nadal-Ginard B (1987) Alternative splicing: a ubiquitous mechanism for the generation of multiple protein isoforms from single genes. Annu Rev Biochem 56:467–495
Bryant DM, Johnson K, DiTommaso T, Tickle T, Couger MB, Payzin-Dogru D, Lee TJ, Leigh ND, Kuo T-H, Davis FG, Bateman J, Bryant S, Guzikowski AR, Tsai SL, Coyne S, Ye WW, Freeman RM Jr, Peshkin L, Tabin CJ, Regev A, Haas BJ, Whited JL (2017) A Tissue-mapped axolotl De Novo transcriptome enables identification of limb regeneration factors. Cell Rep 18:762–776
Dong C, He F, Berkowitz O, Liu J, Cao P, Tang M, Shi H, Wang W, Li Q, Shen Z, Whelan J, Zheng L (2018) Alternative splicing plays a critical role in maintaining mineral nutrient homeostasis in rice (Oryza sativa). Plant Cell 30:2267–2285
Eckardt NA (2013) The plant cell reviews alternative splicing. Plant Cell 25:3639
Ewels P, Magnusson M, Lundin S, Kaller M (2016) MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32:3047–3048
Filichkin SA, Priest HD, Givan SA, Shen R, Bryant DW, Fox SE, Wong W-K, Mochler TC (2010) Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res 20:45–58
Gilbert WJN (1978) Why genes in pieces? Nature 271:501
Jin Y, Cui R, Zhao L, Fan J, Li B (2019) Mechanisms of Panax ginseng action as an antidepressant. Cell Prolif 52:e12696
Kiegle EA, Garden A, Lacchini E, Kater MM (2018) A genomic view of alternative splicing of long non-coding RNAs during rice seed development reveals extensive splicing and lncRNA gene families. Front Plant Sci 9:115
Kim N-H, Jayakodi M, Lee SC, Choi BS, Jang W, Lee J, Kim HH, Waminal NE, Lakshmanan M, van Nguyen B, Lee YS, Park HS, Koo HJ, Park JY, Perumal S, Joh HJ, Lee H, Kim J, Kim IS, Kim K, Koduru L, Kang KB, Sung SH, Yu Y, Park DS, Choi D, Seo E, Kim S, Kim YC, Hyun DY, Park YI, Kim C, Lee TH, Kim HU, Soh MS, Lee Y, In JG, Kim HS, Kim YM, Yang DC, Wing RA, Lee DY, Paterson AH, Yang TJ (2018) Genome and evolution of the shade-requiring medicinal herb Panax ginseng. Plant Biotech J 16:1904–1917
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL (2019) Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotech 37:907–915
Kolde R, Kolde MR (2019) Package ‘pheatmap’. R Package 1(7):790
Lee BY, Kim HS, Choi BS, Hwang DS, Choi AY, Han J, Won E-J, Choi I-Y, Lee S-H, Om A-S, Park HG, Lee J-S (2015) RNA-seq based whole transcriptome analysis of the cyclopoid copepod Paracyclopina nana focusing on xenobiotics metabolism. Comp Biochem Physiol Part D Genomics Proteomics 15:12–19
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Mao S, Zhang S, Zhou Z, Shi X, Huang T, Feng W, Yao C, Gu X, Yu B (2018) Alternative RNA splicing associated with axon regeneration after rat peripheral nerve injury. Exp Neurol 308:80–89
Marquez Y, Brown JW, Simpson C, Barta A, Kalyna M (2012) Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res 22:1184–1195
Otasek D, Morris JH, Bouças J, Pico AR, Demchak B (2019) Cytoscape automation: empowering workflow-based network analysis. Genome Biol 20:185
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ (2008) Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet 40:1413–1415
Park JW, Tokheim C, Shen S, Xing Y (2013) Identifying differential alternative splicing events from RNA sequencing data using RNASeq-MATS. Methods Mol Biol 1038:171–179
Park YJ, Lee JH, Kim JY, Park CM (2019) Alternative RNA splicing expands the developmental plasticity of flowering transition. Front Plant Sci 10:606
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14:417–419
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotech 33:290–295
Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL (2016) Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc 11:1650–1667
Ruan J, Guo F, Wang Y, Li X, Wan S, Shan L, Peng Z (2018) Transcriptome analysis of alternative splicing in peanut (Arachis hypogaea L.). BMC Plant Biol 18:139
Shen S, Park JW, Huang J, Dittmar KA, Lu ZX, Zhou Q, Carstens RP, Xing Y (2012) MATS: a Bayesian framework for flexible detection of differential alternative splicing from RNA-Seq data. Nucleic Acids Res 40:e61
Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, Zhou Q, Xing Y (2014a) rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA 111:E5593-5601
Shen Y, Zhou Z, Wang Z, Li W, Fang C, Wu M, Ma Y, Liu T, Kong L-A, Peng D-L, Tian Z (2014b) Global dissection of alternative splicing in paleopolyploid soybean. Plant Cell 26:996–1008
Sun Y, Hou H, Song H, Lin K, Zhang Z, Hu J, Pang E (2018) The comparison of alternative splicing among the multiple tissues in cucumber. BMC Plant Biol 18:5
Wang L, Wang S, Li W (2012) RSeQC: quality control of RNA-seq experiments. Bioinformatics 28:2184–2185
Wang K, Jiang S, Sun C, Lin Y, Yin R, Wang Y, Zhang MP (2015) The spatial and temporal transcriptomic landscapes of ginseng, Panax ginseng C.A. Meyer. Sci Rep 5:18283
Xu L, Xu J, Shi G, Xiao S, Dai R, Wu S, Sun B, Zhang X, Zhao Y (2020) Optimization of flash extraction, separation of ginsenosides, identification by HPLC-FT-ICR-MS and determination of rare ginsenosides in mountain cultivated ginseng. RSC Adv 10:44050
Yang R, Li P, Mei H, Wang D, Sun J, Yang C, Hao L, Cao S, Chu C, Hu S, Song X, Cao X (2019a) Fine-tuning of MiR528 accumulation modulates flowering time in rice. Mol Plant 12:1103–1113
Yang Z, Yang Z, Yang C, Wang Z, Chen D, Xie Y, Wu Y (2019b) Identification and genetic analysis of alternative splicing of long non-coding RNAs in tomato initial flowering stage. Genomics 112:897–907
Zhang G, Guo G, Hu X, Zhang Y, Li Q, Li R, Zhuang R, Lu Z, He Z, Fang X, Chen L, Tian W, Tao Y, Kristiansen K, Zhang X, Li S, Yang H, Wang J, Wang J (2010) Deep RNA sequencing at single base-pair resolution reveals high complexity of the rice transcriptome. Genome Res 20:646–654
Zhang MP, Liu Y-H, Chang C-S, Zhi H, Wang S, Xu W, Smith CW, Zhang H-B (2019b) Quantification of gene expression while taking into account RNA alternative splicing. Genomics 111:1517–1528
Zhang H, Mao R, Wang Y, Zhang L, Wang C, Lv S, Liu X, Wang Y, Ji W (2019a) Transcriptome-wide alternative splicing modulation during plant-pathogen interactions in wheat. Plant Sci 288:110160
Zhang MP, Liu Y-H, Xu W, Smith CW, Murray SC, Zhang H-B (2020) Analysis of the genes controlling three quantitative traits in three diverse plant species reveals the molecular basis of quantitative traits. Sci Rep 10:10074
Zhao M, Lin Y, Wang YF, Li X, Han Y, Wang K, Sun C, Wang Y, Zhang MP (2019) Transcriptome analysis identifies strong candidate genes for ginsenoside biosynthesis and reveals its underlying molecular mechanism in Panax ginseng C.A. Meyer. Sci Rep 9:615
Funding
This research was supported by awards from China 863 Project (2013AA102604-3), the Bureau of Science and Technology of Jilin Province (20200801063GH, 20190201264JC, 20190103104JC, 20180414077GH, and 20180101027JC), and the Development and Reform Commission of Jilin Province (2018C047-3).
Author information
Authors and Affiliations
Contributions
MPZ and YW designed, conceived, and supervised this study. YLH performed the bioinformatics analysis, conducted PCR, and wrote the manuscript. WYF, LZ, LL ZMZ, LLY, PC, and JL extracted and measured the contents of ginsenosides. KW, CS, JC, and LL helped prepare the tables and figures. MPZ edited and revised the manuscript. All the authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study does not contain any studies with human participants performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, Y., Zhu, L., Li, L. et al. Characteristics of RNA alternative splicing and its potential roles in ginsenoside biosynthesis in a single plant of ginseng, Panax ginseng C.A. Meyer. Mol Genet Genomics 296, 971–983 (2021). https://doi.org/10.1007/s00438-021-01792-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01792-z