Abstract
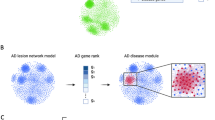
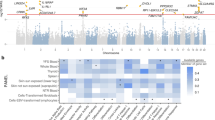
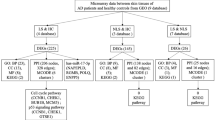
Atopic dermatitis (AD) is a condition driven by T cell-mediated immune response. Targeted therapy of AD is challenging due to its complex pathogenesis. In the current study, by analyzing multiple expression and network datasets, we aimed at: (1) identifying important transcriptomic signatures/profiles for AD to seek potential therapeutic targets and (2) discovering key regulators in the pathogenesis of AD. Our differentially expressed gene (DEG) analysis revealed multiple genes involved in immune response and dermal structural integrity. Functional enrichment analyses suggested that signaling pathways involved in epidermal barrier and inflammation and immunity are overrepresented in lesional AD. Protein–protein interaction (PPI) network and causal interactions analyses highlighted the roles of regulators of epidermal integrity and immune response in the pathogenesis of AD. Prominently, a negative regulator of the B-cell receptor-mediated immune response, PKCβ, has been suggested in the predicted pathogenesis model for AD, implying B cell-mediated immune response may play an equally important role as that of the T cell-mediated immune response in AD. A further search in a perturbagen database has identified small molecular drugs that may alter expression profiles of key regulators in the pathogenesis of AD. In this study, we propose a systemic multi-omics strategy incorporating multiple analyses on various datasets of transcriptomes, diseases, and pharmacology. Such integrative analyses will effectively advance our understanding on the pathogenesis and treatment of AD.





Similar content being viewed by others
Availability of data and material
All datasets are available in open public repositories, with information/links indicated in the "Materials and methods" section.
Code availability
Details of all software and codes used in the current study have been indicated in the "Materials and methods" section.
Change history
24 February 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00438-021-01770-5
References
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles–database and tools update. Nucleic Acids Res 35:D760-765
Bieber T (2008) Atopic dermatitis. N Engl J Med 358:1483–1494
Bolstad BM, Irizarry RA, Astrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19:185–193
Brunner PM, Guttman-Yassky E, Leung DY (2017) The immunology of atopic dermatitis and its reversibility with broad-spectrum and targeted therapies. J Allergy Clin Immunol 139:S65–S76
Cock PJ, Antao T, Chang JT, Chapman BA, Cox CJ, Dalke A, Friedberg I, Hamelryck T, Kauff F, Wilczynski B, de Hoon MJ (2009) Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25:1422–1423
Czarnowicki T, Krueger JG, Guttman-Yassky E (2014) Skin barrier and immune dysregulation in atopic dermatitis: an evolving story with important clinical implications. J Allergy Clin Immunol Pract 2:371–379 (quiz 380–371)
Czarnowicki T, Gonzalez J, Bonifacio KM, Shemer A, Xiangyu P, Kunjravia N, Malajian D, Fuentes-Duculan J, Esaki H, Noda S, Estrada Y, Xu H, Zheng X, Krueger JG, Guttman-Yassky E (2016) Diverse activation and differentiation of multiple B cell subsets in patients with atopic dermatitis but not in patients with psoriasis. J Allergy Clin Immunol 137(118–129):e115
Elias PM, Schmuth M (2009) Abnormal skin barrier in the etiopathogenesis of atopic dermatitis. Curr Opin Allergy Clin Immunol 9:437–446
Elias PM, Steinhoff M (2008) “Outside-to-inside” (and now back to “outside”) pathogenic mechanisms in atopic dermatitis. J Invest Dermatol 128:1067–1070
Esaki H, Ewald DA, Ungar B, Rozenblit M, Zheng X, Xu H, Estrada YD, Peng X, Mitsui H, Litman T, Suarez-Farinas M, Krueger JG, Guttman-Yassky E (2015) Identification of novel immune and barrier genes in atopic dermatitis by means of laser capture microdissection. J Allergy Clin Immunol 135:153–163
Fukuda M, Ohashi H, Matsumoto C, Mishima S, Shimomura Y (2002) Methicillin-resistant Staphylococcus aureus and methicillin-resistant coagulase-negative Staphylococcus ocular surface infection efficacy of chloramphenicol eye drops. Cornea 21:S86-89
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20:307–315
Guttman-Yassky E, Suarez-Farinas M, Chiricozzi A, Nograles KE, Shemer A, Fuentes-Duculan J, Cardinale I, Lin P, Bergman R, Bowcock AM, Krueger JG (2009) Broad defects in epidermal cornification in atopic dermatitis identified through genomic analysis. J Allergy Clin Immunol 124(1235–1244):e1258
Hamid Q, Boguniewicz M, Leung DY (1994) Differential in situ cytokine gene expression in acute versus chronic atopic dermatitis. J Clin Invest 94:870–876
Harper EG, Simpson EL, Takiguchi RH, Boyd MD, Kurtz SE, Bakke AC, Blauvelt A (2008) Efalizumab therapy for atopic dermatitis causes marked increases in circulating effector memory CD4+ T cells that express cutaneous lymphocyte antigen. J Invest Dermatol 128:1173–1181
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Irvine AD, McLean WH, Leung DY (2011) Filaggrin mutations associated with skin and allergic diseases. N Engl J Med 365:1315–1327
Jensen JM, Folster-Holst R, Baranowsky A, Schunck M, Winoto-Morbach S, Neumann C, Schutze S, Proksch E (2004) Impaired sphingomyelinase activity and epidermal differentiation in atopic dermatitis. J Invest Dermatol 122:1423–1431
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kang SW, Wahl MI, Chu J, Kitaura J, Kawakami Y, Kato RM, Tabuchi R, Tarakhovsky A, Kawakami T, Turck CW, Witte ON, Rawlings DJ (2001) PKCbeta modulates antigen receptor signaling via regulation of Btk membrane localization. EMBO J 20:5692–5702
Kim BE, Leung DY, Boguniewicz M, Howell MD (2008) Loricrin and involucrin expression is down-regulated by Th2 cytokines through STAT-6. Clin Immunol 126:332–337
Kohl M, Wiese S, Warscheid B (2011) Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 696:291–303
Leung DY, Bieber T (2003) Atopic dermatitis. Lancet 361:151–160
Lo Surdo P, Calderone A, Iannuccelli M, Licata L, Peluso D, Castagnoli L, Cesareni G, Perfetto L (2018) DISNOR: a disease network open resource. Nucleic Acids Res 46:D527–D534
Lu J, Yin R, Fu Z, Lee VM, Nelson JS, Tan W (2018) Activation of RhoA, Smad2, c-Src, PKC-betaII/delta and JNK in atopic dermatitis. Australas J Dermatol 59:e258–e261
Mancini AJ, Kaulback K, Chamlin SL (2008) The socioeconomic impact of atopic dermatitis in the United States: a systematic review. Pediatr Dermatol 25:1–6
Oliva M, Renert-Yuval Y, Guttman-Yassky E (2016) The “omics” revolution: redefining the understanding and treatment of allergic skin diseases. Curr Opin Allergy Clin Immunol 16:469–476
Perfetto L, Briganti L, Calderone A, Cerquone Perpetuini A, Iannuccelli M, Langone F, Licata L, Marinkovic M, Mattioni A, Pavlidou T, Peluso D, Petrilli LL, Pirro S, Posca D, Santonico E, Silvestri A, Spada F, Castagnoli L, Cesareni G (2016) SIGNOR: a database of causal relationships between biological entities. Nucleic Acids Res 44:D548-554
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47
Saijo K, Mecklenbrauker I, Santana A, Leitger M, Schmedt C, Tarakhovsky A (2002) Protein kinase C beta controls nuclear factor kappaB activation in B cells through selective regulation of the IkappaB kinase alpha. J Exp Med 195:1647–1652
Simon D, Wittwer J, Kostylina G, Buettiker U, Simon HU, Yawalkar N (2008) Alefacept (lymphocyte function-associated molecule 3/IgG fusion protein) treatment for atopic eczema. J Allergy Clin Immunol 122:423–424
Su TT, Guo B, Kawakami Y, Sommer K, Chae K, Humphries LA, Kato RM, Kang S, Patrone L, Wall R, Teitell M, Leitges M, Kawakami T, Rawlings DJ (2002) PKC-beta controls I kappa B kinase lipid raft recruitment and activation in response to BCR signaling. Nat Immunol 3:780–786
Suarez-Farinas M, Tintle SJ, Shemer A, Chiricozzi A, Nograles K, Cardinale I, Duan S, Bowcock AM, Krueger JG, Guttman-Yassky E (2011) Nonlesional atopic dermatitis skin is characterized by broad terminal differentiation defects and variable immune abnormalities. J Allergy Clin Immunol 127(954–964):e951-954
Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, Gould J, Davis JF, Tubelli AA, Asiedu JK, Lahr DL, Hirschman JE, Liu Z, Donahue M, Julian B, Khan M, Wadden D, Smith IC, Lam D, Liberzon A, Toder C, Bagul M, Orzechowski M, Enache OM, Piccioni F, Johnson SA, Lyons NJ, Berger AH, Shamji AF, Brooks AN, Vrcic A, Flynn C, Rosains J, Takeda DY, Hu R, Davison D, Lamb J, Ardlie K, Hogstrom L, Greenside P, Gray NS, Clemons PA, Silver S, Wu X, Zhao WN, Read-Button W, Wu X, Haggarty SJ, Ronco LV, Boehm JS, Schreiber SL, Doench JG, Bittker JA, Root DE, Wong B, Golub TR (2017) A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171(1437–1452):e1417
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613
Thaci D, Simpson EL, Beck LA, Bieber T, Blauvelt A, Papp K, Soong W, Worm M, Szepietowski JC, Sofen H, Kawashima M, Wu R, Weinstein SP, Graham NM, Pirozzi G, Teper A, Sutherland ER, Mastey V, Stahl N, Yancopoulos GD, Ardeleanu M (2016) Efficacy and safety of dupilumab in adults with moderate-to-severe atopic dermatitis inadequately controlled by topical treatments: a randomised, placebo-controlled, dose-ranging phase 2b trial. Lancet 387:40–52
The Gene Ontology C (2019) The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res 47:D330–D338
Tsui C, Martinez-Martin N, Gaya M, Maldonado P, Llorian M, Legrave NM, Rossi M, MacRae JI, Cameron AJ, Parker PJ, Leitges M, Bruckbauer A, Batista FD (2018) Protein kinase C-beta dictates B cell fate by regulating mitochondrial remodeling, metabolic reprogramming, and heme biosynthesis. Immunity 48(1144–1159):e1145
Werner Y, Lindberg M (1985) Transepidermal water loss in dry and clinically normal skin in patients with atopic dermatitis. Acta Derm Venereol 65:102–105
Wolk K, Witte E, Wallace E, Docke WD, Kunz S, Asadullah K, Volk HD, Sterry W, Sabat R (2006) IL-22 regulates the expression of genes responsible for antimicrobial defense, cellular differentiation, and mobility in keratinocytes: a potential role in psoriasis. Eur J Immunol 36:1309–1323
Wollenberg A, Howell MD, Guttman-Yassky E, Silverberg JI, Kell C, Ranade K, Moate R, van der Merwe R (2019) Treatment of atopic dermatitis with tralokinumab, an anti-IL-13 mAb. J Allergy Clin Immunol 143:135–141
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287
Funding
N/A.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All animal protocols have been approved by the Animal Welfare and Care Guidelines in accordance with the National Institute of Health’s guidelines (2004).
Consent to participate
This article does not contain any studies with human participants performed by any of the authors.
Consent for publication
All authors have expressed consents for publishing the current study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, Y., Chen, S. & Chen, J. Profiling transcriptomic changes and signaling pathways in atopic dermatitis by integrative analyses on multiple databases. Mol Genet Genomics 296, 341–353 (2021). https://doi.org/10.1007/s00438-020-01754-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01754-x