Abstract
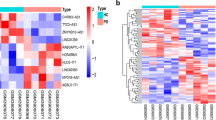
Multiple sclerosis (MS) is a chronic autoimmune disease of the central nervous system, and the pathogenesis is influenced by genetic susceptibility. Accumulating evidence has demonstrated that long non-coding RNAs (lncRNAs) play essential roles in complex diseases, including acting as competing endogenous RNAs (ceRNAs). However, the functional roles and regulatory mechanisms of lncRNAs acting as ceRNAs in MS are still unclear. In this study, we identified hub lncRNA ceRNAs in MS based on ceRNA mechanisms and annotated their functions. The lncRNA-associated ceRNA network (LACN) was constructed by integrating the expression profiles of lncRNA/mRNA and miRNA in MS and normal samples, and the experimentally validated interactions of lncRNA-miRNA and mRNA-miRNA. We found three hub lncRNA ceRNAs (XIST, OIP5-AS1, and CTB-89H12.4) using the network analysis and obtained 96 lncRNA-mediated competing triplets (LCTs, lncRNA-miRNA-mRNA) with the hub lncRNA ceRNAs, which constituted 3 hub ceRNA modules. The functional analysis identified 12 pathways enriched by the 3 hub lncRNA ceRNAs, of which 6 were confirmed to be related to MS. For example, XIST was enriched in the ‘spliceosome’ and ‘RNA transport’ related to the typing of MS, and CTB-89H12.4 was enriched in the ‘mTOR signaling pathway,’ a potential therapeutic target for MS. We dissected the expression patterns of the 96 LCTs in MS individually. LCT XIST-miR-326-HNRNPA1, for which the expression pattern in MS revealed that XIST and HNRNPA1 were up-regulated and miR-326 was down-regulated, consisted of risk RNAs for MS that were validated by other research. Therefore, XIST-miR-326-HNRNPA1 might play a central role in the pathogenesis of MS. These results will contribute to the discovery of novel biomarkers and the development of new therapeutic methods for MS.







Similar content being viewed by others
References
Alvarez JI, Dodelet-Devillers A, Kebir H, Ifergan I, Fabre PJ, Terouz S, Sabbagh M, Wosik K, Bourbonniere L, Bernard M, van Horssen J, de Vries HE, Charron F, Prat A (2011) The Hedgehog pathway promotes blood-brain barrier integrity and CNS immune quiescence. Science 334:1727–1731
Ascherio A, Munger KL (2007) Environmental risk factors for multiple sclerosis. Part I: the role of infection. Ann Neurol 61:288–299
Cai Y, Yu X, Hu S, Yu J (2009) A brief review on the mechanisms of miRNA regulation. Genom Proteom Bioinform 7:147–154
Cardamone G, Paraboschi EM, Solda G, Cantoni C, Supino D, Piccio L, Duga S, Asselta R (2019) Not only cancer: the long non-coding RNA MALAT1 affects the repertoire of alternatively spliced transcripts and circular RNAs in multiple sclerosis. Hum Mol Genet 28:1414–1428
Cheng Y, Chang Q, Zheng B, Xu J, Li H, Wang R (2019) LncRNA XIST promotes the epithelial to mesenchymal transition of retinoblastoma via sponging miR-101. Eur J Pharmacol 843:210–216
Compston A, Coles A (2008) Multiple sclerosis. Lancet 372:1502–1517
Das S, Ghosal S, Sen R, Chakrabarti J (2014) lnCeDB: database of human long noncoding RNA acting as competing endogenous RNA. PLoS ONE 9:e98965
Davis AP, Murphy CG, Saraceni-Richards CA, Rosenstein MC, Wiegers TC, Mattingly CJ (2009) Comparative toxicogenomics database: a knowledgebase and discovery tool for chemical-gene-disease networks. Nucleic Acids Res 37:D786–D792
Dolati S, Aghebati-Maleki L, Ahmadi M, Marofi F, Babaloo Z, Ayramloo H, Jafarisavari Z, Oskouei H, Afkham A, Younesi V, Nouri M, Yousefi M (2018) Nanocurcumin restores aberrant miRNA expression profile in multiple sclerosis, randomized, double-blind, placebo-controlled trial. J Cell Physiol 233:5222–5230
Du Z, Sun T, Hacisuleyman E, Fei T, Wang X, Brown M, Rinn JL, Lee MG, Chen Y, Kantoff PW, Liu XS (2016) Integrative analyses reveal a long noncoding RNA-mediated sponge regulatory network in prostate cancer. Nat Commun 7:10982
Eaton WW, Rose NR, Kalaydjian A, Pedersen MG, Mortensen PB (2007) Epidemiology of autoimmune diseases in Denmark. J Autoimmun 29:1–9
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Farazi TA, Spitzer JI, Morozov P, Tuschl T (2011) miRNAs in human cancer. J Pathol 223:102–115
Feng C, Shen JM, Lv PP, Jin M, Wang LQ, Rao JP, Feng L (2018) Construction of implantation failure related lncRNA-mRNA network and identification of lncRNA biomarkers for predicting endometrial receptivity. Int J Biol Sci 14:1361–1377
Frankish A, Diekhans M, Ferreira AM, Johnson R, Jungreis I, Loveland J, Mudge JM, Sisu C, Wright J, Armstrong J, Barnes I, Berry A, Bignell A, Carbonell Sala S, Chrast J, Cunningham F, Di Domenico T, Donaldson S, Fiddes IT, Garcia Giron C, Gonzalez JM, Grego T, Hardy M, Hourlier T, Hunt T, Izuogu OG, Lagarde J, Martin FJ, Martinez L, Mohanan S, Muir P, Navarro FCP, Parker A, Pei B, Pozo F, Ruffier M, Schmitt BM, Stapleton E, Suner MM, Sycheva I, Uszczynska-Ratajczak B, Xu J, Yates A, Zerbino D, Zhang Y, Aken B, Choudhary JS, Gerstein M, Guigo R, Hubbard TJP, Kellis M, Paten B, Reymond A, Tress ML, Flicek P (2019) GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res 47:D766–D773
Giacoppo S, Pollastro F, Grassi G, Bramanti P, Mazzon E (2017) Target regulation of PI3K/Akt/mTOR pathway by cannabidiol in treatment of experimental multiple sclerosis. Fitoterapia 116:77–84
Guan H, Singh UP, Rao R, Mrelashvili D, Sen S, Hao H, Zumbrun EE, Singh NP, Nagarkatti PS, Nagarkatti M (2016) Inverse correlation of expression of microRNA-140-5p with progression of multiple sclerosis and differentiation of encephalitogenic T helper type 1 cells. Immunology 147:488–498
Honardoost MA, Kiani-Esfahani A, Ghaedi K, Etemadifar M, Salehi M (2014) miR-326 and miR-26a, two potential markers for diagnosis of relapse and remission phases in patient with relapsing-remitting multiple sclerosis. Gene 544:128–133
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Juan L, Wang G, Radovich M, Schneider BP, Clare SE, Wang Y, Liu Y (2013) Potential roles of microRNAs in regulating long intergenic noncoding RNAs. BMC Med Genomics 6(Suppl 1):S7
Kattimani Y, Veerappa AM (2018) Complex interaction between mutant HNRNPA1 and gE of varicella zoster virus in pathogenesis of multiple sclerosis. Autoimmunity 51:147–151
Koch MW, Ilnytskyy Y, Golubov A, Metz LM, Yong VW, Kovalchuk O (2018) Global transcriptome profiling of mild relapsing-remitting versus primary progressive multiple sclerosis. Eur J Neurol 25:651–658
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, Lerner J, Brunet JP, Subramanian A, Ross KN, Reich M, Hieronymus H, Wei G, Armstrong SA, Haggarty SJ, Clemons PA, Wei R, Carr SA, Lander ES, Golub TR (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313:1929–1935
Levin MC, Lee S, Gardner LA, Shin Y, Douglas JN, Salapa H (2017) Autoantibodies to heterogeneous nuclear ribonuclear protein A1 (hnRNPA1) cause altered “ribostasis” and neurodegeneration; the legacy of HAM/TSP as a model of progressive multiple sclerosis. J Neuroimmunol 304:56–62
Li JH, Liu S, Zhou H, Qu LH, Yang JH (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97
Li W, Wang Q, Feng Q, Wang F, Yan Q, Gao SJ, Lu C (2019) Oncogenic KSHV-encoded interferon regulatory factor upregulates HMGB2 and CMPK1 expression to promote cell invasion by disrupting a complex lncRNA-OIP5-AS1/miR-218-5p network. PLoS Pathog 15:e1007578
Li Z, Li X, Jiang C, Qian W, Tse G, Chan MTV, Wu WKK (2018) Long non-coding RNAs in rheumatoid arthritis. Cell Prolif 51
Liu K, Kurien BT, Zimmerman SL, Kaufman KM, Taft DH, Kottyan LC, Lazaro S, Weaver CA, Ice JA, Adler AJ, Chodosh J, Radfar L, Rasmussen A, Stone DU, Lewis DM, Li S, Koelsch KA, Igoe A, Talsania M, Kumar J, Maier-Moore JS, Harris VM, Gopalakrishnan R, Jonsson R, Lessard JA, Lu X, Gottenberg JE, Anaya JM, Cunninghame-Graham DS, Huang AJW, Brennan MT, Hughes P, Illei GG, Miceli-Richard C, Keystone EC, Bykerk VP, Hirschfield G, Xie G, Ng WF, Nordmark G, Eriksson P, Omdal R, Rhodus NL, Rischmueller M, Rohrer M, Segal BM, Vyse TJ, Wahren-Herlenius M, Witte T, Pons-Estel B, Alarcon-Riquelme ME, Guthridge JM, James JA, Lessard CJ, Kelly JA, Thompson SD, Gaffney PM, Montgomery CG, Edberg JC, Kimberly RP, Alarcon GS, Langefeld CL, Gilkeson GS, Kamen DL, Tsao BP, McCune WJ, Salmon JE, Merrill JT, Weisman MH, Wallace DJ, Utset TO, Bottinger EP, Amos CI, Siminovitch KA, Mariette X, Sivils KL, Harley JB, Scofield RH (2016) X Chromosome dose and sex bias in autoimmune diseases: increased prevalence of 47, XXX in systemic Lupus Erythematosus and Sjogren’s syndrome. Arthritis Rheumatol 68:1290–1300
Matsui M, Corey DR (2017) Non-coding RNAs as drug targets. Nat Rev Drug Discov 16:167–179
Mendibe Bilbao M, Boyero Duran S, Barcena Llona J, Rodriguez-Antiguedad A (2019) Multiple sclerosis: Pregnancy and women’s health issues. Neurologia 34:259–269
Mougeot JL, Noll BD, Bahrani Mougeot FK (2019) Sjogren’s syndrome X-chromosome dose effect: an epigenetic perspective. Oral Dis 25:372–384
Paralkar VR, Weiss MJ (2013) Long noncoding RNAs in biology and hematopoiesis. Blood 121:4842–4846
Qu XA, Rajpal DK (2012) Applications of connectivity map in drug discovery and development. Drug Discovery Today 17:1289–1298
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47
Ross MT, Grafham DV, Coffey AJ, Scherer S, McLay K, Muzny D, Platzer M, Howell GR, Burrows C, Bird CP, Frankish A, Lovell FL, Howe KL, Ashurst JL, Fulton RS, Sudbrak R, Wen G, Jones MC, Hurles ME, Andrews TD, Scott CE, Searle S, Ramser J, Whittaker A, Deadman R, Carter NP, Hunt SE, Chen R, Cree A, Gunaratne P, Havlak P, Hodgson A, Metzker ML, Richards S, Scott G, Steffen D, Sodergren E, Wheeler DA, Worley KC, Ainscough R, Ambrose KD, Ansari-Lari MA, Aradhya S, Ashwell RI, Babbage AK, Bagguley CL, Ballabio A, Banerjee R, Barker GE, Barlow KF, Barrett IP, Bates KN, Beare DM, Beasley H, Beasley O, Beck A, Bethel G, Blechschmidt K, Brady N, Bray-Allen S, Bridgeman AM, Brown AJ, Brown MJ, Bonnin D, Bruford EA, Buhay C, Burch P, Burford D, Burgess J, Burrill W, Burton J, Bye JM, Carder C, Carrel L, Chako J, Chapman JC, Chavez D, Chen E, Chen G, Chen Y, Chen Z, Chinault C, Ciccodicola A, Clark SY, Clarke G, Clee CM, Clegg S, Clerc-Blankenburg K, Clifford K, Cobley V, Cole CG, Conquer JS, Corby N, Connor RE, David R, Davies J, Davis C, Davis J, Delgado O, Deshazo D et al (2005) The DNA sequence of the human X chromosome. Nature 434:325–337
Sawcer S, Hellenthal G, Pirinen M, Spencer CC, Patsopoulos NA, Moutsianas L, Dilthey A, Su Z, Freeman C, Hunt SE, Edkins S, Gray E, Booth DR, Potter SC, Goris A, Band G, Oturai AB, Strange A, Saarela J, Bellenguez C, Fontaine B, Gillman M, Hemmer B, Gwilliam R, Zipp F, Jayakumar A, Martin R, Leslie S, Hawkins S, Giannoulatou E, D’Alfonso S, Blackburn H, Martinelli Boneschi F, Liddle J, Harbo HF, Perez ML, Spurkland A, Waller MJ, Mycko MP, Ricketts M, Comabella M, Hammond N, Kockum I, McCann OT, Ban M, Whittaker P, Kemppinen A, Weston P, Hawkins C, Widaa S, Zajicek J, Dronov S, Robertson N, Bumpstead SJ, Barcellos LF, Ravindrarajah R, Abraham R, Alfredsson L, Ardlie K, Aubin C, Baker A, Baker K, Baranzini SE, Bergamaschi L, Bergamaschi R, Bernstein A, Berthele A, Boggild M, Bradfield JP, Brassat D, Broadley SA, Buck D, Butzkueven H, Capra R, Carroll WM, Cavalla P, Celius EG, Cepok S, Chiavacci R, Clerget-Darpoux F, Clysters K, Comi G, Cossburn M, Cournu-Rebeix I, Cox MB, Cozen W, Cree BA, Cross AH, Cusi D, Daly MJ, Davis E, de Bakker PI, Debouverie M, D’Hooghe MB, Dixon K, Dobosi R, Dubois B, Ellinghaus D, Elovaara I, Esposito F et al (2011) Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476:214–219
Shaker OG, Mahmoud RH, Abdelaleem OO, Ahmed TI, Fouad NA, Hussein HA, Nassr MH, Zaki OM, Abdelghaffar NK, Hefzy EM (2019) Expression profile of long noncoding RNAs, lnc-Cox2, and HOTAIR in rheumatoid arthritis patients. J Interferon Cytokine Res 39:174–180
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Stadelmann C (2011) Multiple sclerosis as a neurodegenerative disease: pathology, mechanisms and therapeutic implications. Curr Opin Neurol 24:224–229
Syrett CM, Sindhava V, Sierra I, Dubin AH, Atchison M, Anguera MC (2018) Diversity of epigenetic features of the inactive X-chromosome in NK cells, dendritic cells, and macrophages. Front Immunol 9:3087
Tan JY, Sirey T, Honti F, Graham B, Piovesan A, Merkenschlager M, Webber C, Ponting CP, Marques AC (2015) Extensive microRNA-mediated crosstalk between lncRNAs and mRNAs in mouse embryonic stem cells. Genome Res 25:655–666
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505:344–352
Vawter MP, Evans S, Choudary P, Tomita H, Meador-Woodruff J, Molnar M, Li J, Lopez JF, Myers R, Cox D, Watson SJ, Akil H, Jones EG, Bunney WE (2004) Gender-specific gene expression in post-mortem human brain: localization to sex chromosomes. Neuropsychopharmacology 29:373–384
Walavalkar K, Saravanan B, Singh AK, Jayani RS, Nair A, Farooq U, Islam Z, Soota D, Mann R, Shivaprasad PV, Freedman ML, Sabarinathan R, Haiman CA, Notani D (2020) A rare variant of African ancestry activates 8q24 lncRNA hub by modulating cancer associated enhancer. Nature Communications 11:3598
Wang H, Lu X, Chen F, Ding Y, Zheng H, Wang L, Zhang G, Yang J, Bai Y, Li J, Wu J, Zhou M, Xu L (2018) Landscape of SNPs-mediated lncRNA structural variations and their implication in human complex diseases. Briefings in Bioinformatics
Wang J, Syrett CM, Kramer MC, Basu A, Atchison ML, Anguera MC (2016) Unusual maintenance of X chromosome inactivation predisposes female lymphocytes for increased expression from the inactive X. Proc Natl Acad Sci USA 113:E2029–E2038
Wang L, Zheng Z, Feng X, Zang X, Ding W, Wu F, Zhao Q (2019) circRNA/lncRNA-miRNA-mRNA network in oxidized, low-density, lipoprotein-induced foam cells. DNA Cell Biol 38:1499–1511
Waschbisch A, Atiya M, Linker RA, Potapov S, Schwab S, Derfuss T (2011) Glatiramer acetate treatment normalizes deregulated microRNA expression in relapsing remitting multiple sclerosis. PLoS ONE 6:e24604
Ye N, Rao S, Du T, Hu H, Liu Z, Shen Y, Xu Q (2017) Intergenic variants may predispose to major depression disorder through regulation of long non-coding RNA expression. Gene 601:21–26
Zhang R, Xia T (2017) Long non-coding RNA XIST regulates PDCD4 expression by interacting with miR-21-5p and inhibits osteosarcoma cell growth and metastasis. Int J Oncol 51:1460–1470
Zhang Z, Liu F, Yang F, Liu Y (2018) Kockdown of OIP5-AS1 expression inhibits proliferation, metastasis and EMT progress in hepatoblastoma cells through up-regulating miR-186a-5p and down-regulating ZEB1. Biomed Pharmacother 101:14–23
Zhou M, Diao Z, Yue X, Chen Y, Zhao H, Cheng L, Sun J (2016a) Construction and analysis of dysregulated lncRNA-associated ceRNA network identified novel lncRNA biomarkers for early diagnosis of human pancreatic cancer. Oncotarget 7:56383–56394
Zhou M, Wang X, Shi H, Cheng L, Wang Z, Zhao H, Yang L, Sun J (2016b) Characterization of long non-coding RNA-associated ceRNA network to reveal potential prognostic lncRNA biomarkers in human ovarian cancer. Oncotarget 7:12598–12611
Acknowledgements
This work was supported by the Natural Science Foundation of Heilongjiang Province (Grant No. LH2019C043). We would like to express our gratitude to EditSprings (https://www.editsprings.com/) for the expert linguistic services provided.
Author information
Authors and Affiliations
Contributions
YD: data curation, formal analysis, methodology, writing—original draft. TL: investigation, resources. XY: investigation, resources. MC: validation, visualization. CW: validation, visualization. SW: software. FZ: project administration, supervision, writing—review and editing. RZ: conceptualization, funding acquisition, project administration, supervision, writing—review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, Y., Li, T., Yan, X. et al. Identification of hub lncRNA ceRNAs in multiple sclerosis based on ceRNA mechanisms. Mol Genet Genomics 296, 423–435 (2021). https://doi.org/10.1007/s00438-020-01750-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01750-1