Abstract
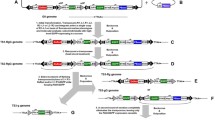
The piggyBac (PB) transposon is the most widely used vector for generating transgenic silkworms. The stability of the PB transposon in the receptor is a serious concern that requires attention because of biosafety concerns. In this study, we found that the transgene silkworm developed loss of reporter gene traits. To further investigate the regularity, we traced the genes and traits of this silkworm. After successful alteration of the silkworm genome with the MASP1 gene (named red-eyed silkworm; RES), silkworm individuals with lost reporter genes were found after long-term transgenerational breeding and were designated as the white-eyed silkworm (WES). PCR amplification indicated that exogenous genes had been lost in the WES. Testing was conducted on the PB transposons, and the left arm (L arm) did not exist; however, the right arm (R arm) was preserved. Amino acid analysis showed that the amino acid content of the WES changed versus the common silkworm and RES. These results indicate that the migration of PB transposons in Bombyx mori does occur and is unpredictable. This is because the silkworm genome contains multiple PB-like sequences that might influence the genetic stability of transgenic lines. When using PB transposons as a transgene vector, it is necessary to fully evaluate and take necessary measures to prevent its re-migration in the recipient organism. Further experiments are needed if we want to clarify the regularity of the retransposition phenomenon and the direct and clear association with similar sequences of transposons.



Similar content being viewed by others
References
Cao TT, Zhang YQ (2016) Processing and characterization of silk sericin from Bombyx mori and its application in biomaterials and biomedicines. Mater Sci Eng C 61:940–952. https://doi.org/10.1016/j.msec.2015.12.082
Dafa’alla TH et al (2006) Transposon-free insertions for insect genetic engineering. Nat Biotechnol 24:820–821. https://doi.org/10.1038/nbt1221
Daimon T, Mitsuhiro M, Katsuma S, Abe H, Mita K, Shimada T (2010) Recent transposition of yabusame, a novel piggyBac-like transposable element in the genome of the silkworm Bombyx mori. Genome 53:585–593. https://doi.org/10.1139/g10-035
Ding S, Wu X, Li G, Han M, Zhuang Y, Xu T (2005) Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 122:473–483. https://doi.org/10.1016/j.cell.2005.07.013
Goldsmith MR, Shimada T, Abe H (2005) The genetics and genomics of the silkworm Bombyx mori. Annu Rev Entomol 50:71–100. https://doi.org/10.1146/annurev.ento.50.071803.130456
Handler AM (2002) Use of the piggyBac transposon for germ-line transformation of insects. Insect Biochem Mol Biol 32(2002):1211–1220. https://doi.org/10.1016/S0965-1748(02)00084-X
Handler AM, Zimowska GJ, Horn C (2004) Post-integration stabilization of a transposon vector by terminal sequence deletion in Drosophila melanogaster. Nat Biotechnol 22:1150–1154. https://doi.org/10.1038/nbt1002
Handler AM, McCombs SD, Fraser MJ, Saul SH (1998) The lepidopteran transposon vector, piggyBac, mediates germ-line transformation in the Mediterranean fruit fly. Proc Natl Acad Sci USA 95:7520–7525
Horn C, Wimmer EA (2000) A versatile vector set for animal transgenesis. Dev Genes Evol 210:630–637
Howard WMRD (2002) Assessment of possible ecological risks and hazards of transgenic fish with implications for other sexually reproducing organisms. Transgenic Res 11:101–114
Kim HJ, Kim MK, Lee KH, Nho SK, Han MS, Um IC (2017) Effect of degumming methods on structural characteristics and properties of regenerated silk. Int J Biol Macromol 104:294–302. https://doi.org/10.1016/j.ijbiomac.2017.06.019
Lorenzen M, Kimzey T, Shippy T, Brown S, Denell R, Beeman R (2007) piggyBac-based insertional mutagenesis in Tribolium castaneum using donor/helper hybrids. Insect Mol Biol 16:265–275. https://doi.org/10.1111/j.1365-2583.2007.00727.x
Rubin GM, Spradling AC (1982) Genetic transformation of Drosophila with transposable element vectors. Science 218:348–353
Sato M, Inada E, Saitoh I, Watanabe S, Nakamura S (2020) piggyBac-based non-viral in vivo gene delivery useful for production of genetically modified animals and organs. Pharmaceutics. https://doi.org/10.3390/pharmaceutics12030277
Sethuraman N, Fraser MJ Jr, Eggleston P, O’Brochta DA (2007) Post-integration stability of piggyBac in Aedes aegypti. Insect Biochem Mol Biol 37:941–951. https://doi.org/10.1016/j.ibmb.2007.05.004
Tamura T et al (2000) Germline transformation of the silkworm Bombyx mori L. using a piggyBac transposon-derived vector. Nat Biotechnol 18:81–84. https://doi.org/10.1038/71978
Teule F et al (2012) Silkworms transformed with chimeric silkworm/spider silk genes spin composite silk fibers with improved mechanical properties. Proc Natl Acad Sci USA 109:923–928. https://doi.org/10.1073/pnas.1109420109
Thibault S et al (2004) P and piggyBac transposons display a complementary insertion spectrum in Drosophila: a multifunctional toolkit to manipulate an insect genome. Nat Genet 36:283–287
Wang YJ, Zhag YQ (2011) Three-layered sericins around the silk fibroin fiber from Bombyx mori cocoon and their amino acid composition. Adv Mater Res 175–176:158–163
Wang F et al (2014) Advanced silk material spun by a transgenic silkworm promotes cell proliferation for biomedical application. Acta Biomater 10:4947–4955. https://doi.org/10.1016/j.actbio.2014.06.031
Wang F et al (2015) Remobilizing deleted piggyBac vector post-integration for transgene stability in silkworm. Mol Genet Genomics 290:1181–1189. https://doi.org/10.1007/s00438-014-0982-6
Xia Q, Guo Y, Zhang Z, Li D, Xuan Z, Li Z et al (2009) Complete resequencing of 40 genomes reveals domestication events and genes in silkworm. Science 326:433–436. https://doi.org/10.1126/science
Xu HF et al (2006) Identification and characterization of piggyBac-like elements in the genome of domesticated silkworm, Bombyx mori. Mol Genet Genomics 276:31–40. https://doi.org/10.1007/s00438-006-0124-x
You Z et al (2018) Extraordinary mechanical properties of composite silk through hereditable transgenic silkworm expressing recombinant major ampullate spidroin. Sci Rep 8:15956. https://doi.org/10.1038/s41598-018-34150-y
Zhang M, Wang C, Wang H, Jian M, Hao X, Zhang Y (2017) Carbonized cotton fabric for high-performance wearable strain sensors. Adv Funct Mater. https://doi.org/10.1002/adfm.201604795
Zhang P, Liu S, Song HS, Zhang G, Jia Q, Li S (2017) Yorkie(CA) overexpression in the posterior silk gland improves silk yield in Bombyx mori. J Insect Physiol 100:93–99. https://doi.org/10.1016/j.jinsphys.2017.06.001
Zhao A et al (2010) New and highly efficient expression systems for expressing selectively foreign protein in the silk glands of transgenic silkworm. Transgenic Res 19:29–44. https://doi.org/10.1007/s11248-009-9295-7
Funding
This work was supported by the National Natural Science Foundation of China (No. 31830094), Chongqing postgraduate research innovation project (Grant No. CYS19110) and funds from the China Agriculture Research System (No. CARS-18-ZJ0102).
Author information
Authors and Affiliations
Contributions
XJ conducted molecular lab work, data analysis, sequence alignments, participated in the design of the study, and drafted the manuscript; XP, YY and QG conducted statistical analyses and critically revised the manuscript; TZ, ML, GZ, and FD developed the study design and helped draft the manuscript. All of the authors gave final approval for publication and agree to be held accountable for the work performed therein.
Corresponding author
Ethics declarations
Conflict of interest
We declare we have no competing interests.
Research involving animal rights
All animal studies were conducted in compliance with the protocols approved by the National Centre of Animal Science Experimental Teaching of Southwest University of China.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jia, X., Pang, X., Yuan, Y. et al. Unpredictable recombination of PB transposon in Silkworm: a potential risk. Mol Genet Genomics 296, 271–277 (2021). https://doi.org/10.1007/s00438-020-01743-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01743-0