Abstract
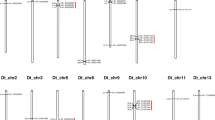
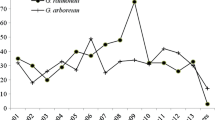
Cotton is the most important natural fiber used in textiles. Breeding for “three-lines”, i.e., cytoplasmic male sterility (CMS)-based sterile (A), maintainer (B), and restorer (R) line, is a promising approach to harness hybrid vigor in cotton. Pentatricopeptide repeat (PPR) protein-encoding genes play an important role in plant growth and development including restoration of CMS plants to male fertility. However, PPRs, especially those contributing to CMS and fiber development, remain largely unknown in cotton. In this study, a genome-wide identification and characterization of PPR gene family in four Gossypium species with genome sequences (G. arboreum, G. raimondii, G. hirsutum, and G. barbadense) were performed, and expressed PPR genes in developing floral buds, ovules, and fibers were compared to identify possible PPRs related to CMS restoration and fiber development. A total of 539, 558, 1032, and 1055 PPRs were predicted in the above four species, respectively, which were further mapped to chromosomes for a synteny analysis. Through an RNA-seq analysis, 86% (882) PPRs were expressed in flowering buds of upland cotton (G. hirsutum); however, only 11 and 6 were differentially expressed (DE) between restorer R and its near-isogenic (NI) B and between R and its NI A line, respectively. Another RNA-seq analysis identified the expression of only 54% (556) PPRs in 0 and 3 day(s) post-anthesis (DPA) ovules and 24% (247) PPRs in 10 DPA fibers; however, only 59, 6, and 27 PPRs were DE in 0 and 3 DPA ovules, and 10 DPA fibers between two backcross inbred lines (BILs) with differing fiber length, respectively. Only 2 PPRs were DE between Xuzhou 142 and its fiberless and fuzzless mutant. Quantitative RT-PCR analysis confirmed the validity of the RNA-seq results for the gene expression pattern. Therefore, only a very small number of PPRs may be associated with fertility restoration of CMS and genetic differences in fiber initiation and elongation. These results lay a foundation for understanding the roles of PPR genes in cotton, and will be useful in the prioritization of candidate PPR gene functional validation for cotton CMS restoration and fiber development.




Similar content being viewed by others
Abbreviations
- CMS:
-
Cytoplasmic male sterility
- PPR:
-
Pentatricopeptide repeat
- DPA:
-
Days post-anthesis
- BILs:
-
Backcross inbred lines
- Rf:
-
Restorer of fertility
- DEG:
-
Differentially expressed gene
References
Bentolila S, Alfonso AA, Hanson MR (2002) A pentatricopeptide repeat-containing gene restores fertility to cytoplasmic male-sterile plants. Proc Natl Acad Sci USA 99:10887–10892
Bohra A, Jha UC, Adhimoolam P, Bisht D, Singh NP (2016) Cytoplasmic male sterility (CMS) in hybrid breeding in field crops. Plant Cell Rep 35:967–993
Boli Y, Hu W, Li X, Cao J, Fan L (2015) Over-expression of GhUGP1 in Upland cotton improves fibre quality and reduces fibre sugar content. Plant Breed 134:197–202
Chase CD (2007) Cytoplasmic male sterility: a window to the world of plant mitochondrial-nuclear interactions. Trends Genet 23:81–90
Dahan J, Mireau H (2013) The Rf and Rf-like PPR in higher plants, a fast-evolving subclass of PPR genes. RNA Biol 10:1469–1476
Du X, Huang G, He S et al (2018) Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat Genet 50:796–802
Eckardt NA (2006) Cytoplasmic male sterility and fertility restoration. Plant Cell 18:515–517
Fan M, Jin L, Liu Q, Qu D (2008) Cloning of SoDIPPR gene of pentatricopeptide repeat (PPR) protein family in potato and analysis of expression characteristics under drought conditions. Sci Agric Sin 41:2249–2257
Fang DD (2018) Cotton fiber: physics, chemistry and biology. Springer, New Orleans, pp 2–5
Geddy R, Brown GG (2007) Genes encoding pentatricopeptide repeat (PPR) proteins are not conserved in location in plant genomes and may be subject to diversifying selection. BMC Genom 8:130
Hajrah NH, Obaid AY, Atef A et al (2017) Transcriptomic analysis of salt stress responsive genes in Rhazya stricta. PLoS ONE 12:1–18
Horna R, Guptab KJ, Colomboc N (2014) Mitochondrion role in molecular basis of cytoplasmic male sterility. Mitochondrion 19:198–205
Jo YD, Ha Y, Lee JH, Park M, Bergsma AC, Choi H, Goritschnig S, Kloosterman B, Dijk PJ, Choi D, Kang BC (2016) Fine mapping of restorer-of-fertility in pepper (Capsicum annuum L.) identified a candidate gene encoding a pentatricopeptide repeat (PPR)-containing protein. Theor Appl Genet 129:2003–2017
Krzywinski M, Schein J, Birol I et al (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Laluk K, AbuQamar S, Mengiste T (2011) The Arabidopsis mitochondria-localized pentatricopeptide repeat protein PGN functions in defense against necrotrophic fungi and abiotic stress tolerance. Plant Physiol 156:2053–2068
Letunic I, Bork P (2019) Interactive tree of life (iTOL): an online tool for phylogenetic tree display and annotation. FEBS Lett 232:78–82
Li S, Sun Q, Hu M, Li S, Zhu Y, Zhu Y (2012) Phylogenetic genome-wide comparisons of the pentatricopeptide repeat gene family in indica and japonica rice. Biochem Genet 50:978–989
Li F, Fan G, Wang K, Sun F et al (2014a) Genome sequence of the cultivated cotton Gossypium arboreum. Nat Genet 46:567–573
Li X, Zhang Y, Hou M, Sun F, Shen Y, Xiu Z, Wang X, Chen Z, Sun SS, Ian Small, Tan BC (2014b) Small kernel 1 encodes a pentatricopeptide repeat protein required for mitochondrial nad7 transcript editing and seed development in maize (Zea mays) and rice (Oryza sativa). Plant J 79:797–809
Li F, Fan G, Lu C et al (2015) Genome sequence of cultivated upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotech 33:524–530
Li X, Wu M, Liu G, Pei W, Zhai H, Yu J, Zhang J, Yu S (2017) Identification of candidate genes for fiber length quantitative trait loci through RNA-Seq and linkage and physical mapping in cotton. BMC Genom 18:427
Liu Y, Xiu Z, Meeley R, Tan B (2013) Empty pericarp5 encodes a pentatricopeptide repeat protein that is required for mitochondrial RNA editing and seed development in maize. Plant Cell 25:868–883
Liu J, Xu Z, Lu P, Li W, Chen M, Guo C, Ma Y (2016) Genome-wide investigation and expression analyses of the pentatricopeptide repeat protein gene family in foxtail millet. BMC Genom 17:840
Liu Z, Dong F, Wang X, Wang T, Su R, Hong D, Yang G (2017) A pentatricopeptide repeat protein restores nap cytoplasmic male sterility in Brassica napus. J Exp Bot 68:4115–4123
Lurin C, Andrés C, Aubourg S et al (2004) Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16:2089–2103
Lv H, Huang C, Guo G, Yang Z (2014) Roles of the nuclear-encoded chloroplast SMR domain-containing PPR protein SVR7 in photosynthesis and oxidative stress tolerance in Arabidopsis. J Plant Biol 57:291–301
Ma Q, Wu M, Pei W, Wang X, Zhai H, Wang W, Li X, Zhang J, Yu J, Yu S (2016) RNA-seq-mediated transcriptome analysis of a fiberless mutant cotton and its possible origin based on SNP markers. PLoS One 11:e0151994
Manna S (2015) An overview of pentatricopeptide repeat proteins and their applications. Biochim 113:93e99
Meyer VG (1975) Male sterility from Gossypium harknessii. J Hered 66:23–27
Paterson AH, Wendel JF, Gundlach H et al (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–428
Qin Y, Zhu Y (2011) How cotton fibers elongate: a tale of linear cell-growth mode. Curr Opin Plant Biol 14:106–111
Saha D, Prasad AM, Srinivasan R (2007) Pentatricopeptide repeat proteins and their emerging roles in plants. Plant Physiol Biochem 45:521e534
Schmitz-Linneweber C, Small I (2008) Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends Plant Sci 13:663–670
Stewart JM (1992) A new cytoplasmic male sterility and restorer for cotton. Proc Beltwide Cotton Res Conf p 610
Wang F, Yue B, Hu J, Stewart JM, Zhang J (2009) A Target region amplified polymorphism marker for fertility restorer gene Rf1 and chromosomal localization of Rf1 and Rf2 in cotton. Crop Sci 49:1602–1608
Wang K, Wang Z, Li F et al (2012) The draft genome of a diploid cotton Gossypium raimondii. Nat Genet 44:1098–1103
Wang K, Peng X, Ji Y, Yang P, Zhu Y, Li S (2013) Gene, protein and network of male sterility in rice. Front Plant Sci 4:92
Wang M, Tu L, Yuan D et al (2019) Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nature Genet 51:224
Wendel JF, Grover CE (2015) Taxonomy and evolution of the cotton genus, Gossypium. In: Fang DD, Percy RG (eds) cotton, 2nd edn. ASA-SSSA-CSSA, Madison, pp 25–44
Whitford R, Fleury D, Reif JC, Garcia M, Okada T, Korzun V, Langridge P (2013) Hybrid breeding in wheat: technologies to improve hybrid wheat seed production. J Exp Bot 64:5411–5428
Wu J, Cao X, Guo L, Qi T, Wang H, Tang H, Zhang J, Xing C (2014) Development of a candidate gene marker for Rf1 based on a PPR gene in cytoplasmic male sterile CMS-D2 upland cotton. Mol Breed 34:231–240
Wu J, Zhang M, Zhang B, Zhang X, Guo L, Qi T, Wang H, Zhang J, Xing C (2017) Genome-wide comparative transcriptome analysis of CMS-D2 and its maintainer and restorer lines in upland cotton. BMC Genom 18:454
Zhang J, Stewart JM (2001a) CMS-D8 restoration in cotton is conditioned by one dominant gene. Crop Sci 41:283–288
Zhang J, Stewart JM (2001b) Inheritance and genetic relationships of the D8 and D2-2 restorer genes for cotton cytoplasmic male sterility. Crop Sci 41:289–294
Zhang J, Percy RG, McCartyJr JC (2014) Introgression genetics and breeding between upland and pima cotton: a review. Euphytica 198:1–12
Zhang B, Liu G, Li X et al (2017) A genome-wide identification and analysis of the DYW-deaminase genes in the pentatricopeptide repeat gene family in cotton (Gossypium spp.). PLoS One 12:e0174201
Zhao N, Wang YM, Hua JP (2018) Genomewide identification of PPR gene family and prediction analysis on restorer gene in Gossypium. J Genet 97:1083–1095
Zhu Q, Dugardeyn J, Zhang C et al (2014) The Arabidopsis thaliana RNA editing factor SLO2, which affects the mitochondrial electron transport chain, participates in multiple stress and hormone responses. Mol Plant 7:290–310
Funding
This work was financially supported by National Natural Science Foundation of China (31240011), the Youth Scientific Research Foundation of Shandong Academy of Agricultural Sciences (2014QNM05), and Natural Science Foundation of Shandong Province (BS2011NY09).
Author information
Authors and Affiliations
Contributions
Zongfu Han performed the study, analyzed pentatricopeptide repeat (PPR) protein-encoding genes in Gossypium species, and drafted the manuscript. Yuxiang Qin helped in synteny analysis of PPR genes and edited the manuscript. Xihua Li collected (ovules and fibers) samples and extracted RNA, and analyzed RNA-Seq data. Jiwen Yu collected flowering buds, extracted RNA and analyzed RNA-Seq data. Ruzhong Li assisted Xinhua Li in collecting samples; Chaozhu Xing assisted Jiwen Yu in sample collection and RNA extraction. Mingzhou Song assisted Li and Yu in finding of DEGs from the RNA-seq raw data, Jianyong Wu helped to add detailed information on RNA-seq experiment. Jinfa Zhang conceived the project and revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Informed consent
Not applicable.
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, Z., Qin, Y., Li, X. et al. A genome-wide analysis of pentatricopeptide repeat (PPR) protein-encoding genes in four Gossypium species with an emphasis on their expression in floral buds, ovules, and fibers in upland cotton. Mol Genet Genomics 295, 55–66 (2020). https://doi.org/10.1007/s00438-019-01604-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01604-5